* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download second of three for Chapter 8
Comparative genomic hybridization wikipedia , lookup
Transposable element wikipedia , lookup
Human genome wikipedia , lookup
Oncogenomics wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Minimal genome wikipedia , lookup
Gene desert wikipedia , lookup
Ridge (biology) wikipedia , lookup
Gene expression profiling wikipedia , lookup
Genomic library wikipedia , lookup
Point mutation wikipedia , lookup
Copy-number variation wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Hybrid (biology) wikipedia , lookup
Genomic imprinting wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Genome evolution wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Designer baby wikipedia , lookup
Genome (book) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microevolution wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Gene expression programming wikipedia , lookup
Y chromosome wikipedia , lookup
X-inactivation wikipedia , lookup
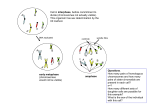
Chapter 8 Human Karyotypes and Chromosome Behavior Jones and Bartlett Publishers © 2005 Changes in chromosome structure - deletions - duplications - inversions - translocations Deletions/Deficiencies Interstitial deficiency Terminal deficiency Terminal Deficiencies unstable chromosomes Haplo-insufficient genes • Heterozygotes for a deficiency may have a mutant phenotype. • If two working copies of a gene are required for normal gene function, only havinog one is a problem. • ‘Cri du Chat’ syndrome in humans: – Deficiency of a part of chromosome 5 Mapping the deletion of part of a chromosome segment by testcrosses and uncovering of recessive genes Mapping of genes in Drosophila using overlapping deletions and polytene chromosomes Duplications • Mutations can produce an extra copy of a part of a chromosome called a duplication. • Tandem duplication – same sequence, adjacent to the original copy. • Reverse tandem duplication – opposite orientation. • Displaced duplication • Small free chromosome Unequal crossing over of misaligned repeat sequences leads to gain or loss of repeats Unequal crossing over involving eye pigment genes Inversions Mechanism of creation of a chromosomes with an inverted segment Pairing of homologous chromosomes in an inversion heterozygote An inversion which does not involve the centromere is called a paracentric inversion Synapsis between chromosomes bearing inversions requires the formation of an Inversion Loop If there is no crossing over inside the loop chromosomes disjoin normally Crossing over in an inversion loop for a pericentric inversion duplications and deletions Crossing over in an inversion loop for a paracentric inversion messed-up chromosomes Absence of recombination within an inversion loop does not create deletions or duplications A crossover within an inversion loop of a paracentric inversion creates dicentric and acentric chromosomes A crossover within an inversion loop of a pericentric inversion creates chromosomes with deletion and duplication When an inversion involves the centromere, it is called a pericentric inversion. Crossing over in a pericentric inversion does not create dicentric and acentric chromosomes Translocations • Broken chromosomes can reattach to different chromosomes that also were fragmented. • Exchanges between nonhomologous chromosomes are called translocations. • Reciprocal translocations – distal portions of two chromosomes are exchanged. • Transposition – insertion into another chromosome. Structure of chromosomes with a reciprocal translocation Pairing and segregation of chromosomes with a reciprocal translocation during meiosis I Robertsonian translocation • Exchange of entire arms between chromosomes. • This can result in change of chromosome number in a monoploid set. • Can occur by fusion of two acrocentric chromosomes, or a split in a metacentric into two. • Robertsonian translocations can result in barriers, speciation. Mechanism of creation of a Robertsonian translocation Pairing and segregation with a Robertsonian translocation involving human chromosomes 14 and 21 Such a translocation results in a high probability of having a child with Down syndrome. Variegation (mottling) of eye color due to positioning of the eye color gene near centromeric heterochromatin When the expression of a gene is affected by its location on a chromosome (even though the gene itself is not changed), such a variation is called “position effect” Two kinds of polyploidy Multiplication of the entire chromosome complement is called polyploidy. When all the genomes are the same, it is called autopolyploidy. When two (or more) different genomes are duplicated, it is called allopolyploidy. Formation of a tetraploid organism Creation of a totally homozygous diploid cell by doubling of chromosome number in a monoploid cell by colchicine Monoploid cells can only be grown in plants. In humans, the only viable monoploid cells are the egg and the sperm. Monoploidy in somatic cells is lethal.