* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download PowerPoint 프레젠테이션
Gene desert wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Gene therapy wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Cancer epigenetics wikipedia , lookup
DNA vaccination wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Molecular cloning wikipedia , lookup
Genomic library wikipedia , lookup
Epigenomics wikipedia , lookup
DNA supercoil wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Non-coding DNA wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Population genetics wikipedia , lookup
Point mutation wikipedia , lookup
Genome evolution wikipedia , lookup
Genome (book) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gene expression programming wikipedia , lookup
Designer baby wikipedia , lookup
Genetic engineering wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genome editing wikipedia , lookup
History of genetic engineering wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Helitron (biology) wikipedia , lookup
Microevolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Homologous recombination wikipedia , lookup
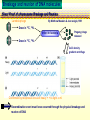
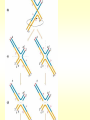
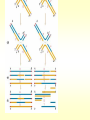
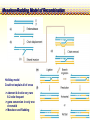
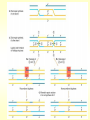
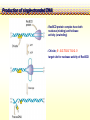
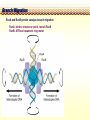
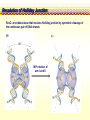
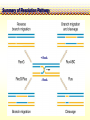
CHAPTER 19 MECHANISMS OF RECOMBINATION Recombination occurs at regions of homology between chromosomes through the breakage and reunion of DNA molecules. Models for recombination, such as the Holliday model, involve the creation of a heteroduplex branch, or cross bridge, that can migrate and the subsequent splicing of the intermediate structure to yield different types of recombinant DNA molecules. Recombination models can be applied to explain genetic crosses. Many of the enzymes participating in recombination in bacteria have been identified. Basic Crossover Event Linkage analysis: recombination of genes by cross-over -> Molecular mechanism of recombination by cross-over Break and rejoin Benzer’s work; Recombination within the gene -> should be precise -> base-pair complementarity Breakage and reunion of DNA molecules Direct Proof of chromosome Breakage and Reunion Lambda phage ++ By Matthew Meselen & Jean weigle, 1961 Grow in 12C, 14N Infect to bacteria c mi Progeny phage released Grow in 13C, 15N CsCl density gradient centrifuge of phage DNA Confirmed by reciprocal cross of heavy + + to light c mi Recombination event must have occurred through the physical breakage and reunion of DNA Chiasmata: the crossover points Chiasmata Are Actual Site of Crossover Direct Evidence; Harlequin chromosome - by C. Tease & G. H. Jones, 1978 (see Ch. 5, 8) Centromeres are pulled apart Indirect Evidence; recombination mapping average of one crossover per meiosis produces 50 m.u. = mean number of chiasmata Genetic results leading to recombination models Tetrad analyses in filamentous fungi; Neurospora crassa (see Ch.6) Gene conversion Polarity of conversion frequency Conversion and crossing-over Co-conversion These crucial findings provided the impetus for the models of intragenic recombination. Genetic results leading to recombination models Gene Conversion during Meiosis Departures from predicted Mendelian 4:4 segregation 0.1-1.0% in filamentous fungi, up to 4% in yeast Gene conversion 5:3 or 3:5 ratios ; two different strand of double helix carrying information for two different alleles at the conclusion of meiosis Mutation The allele that is converted always changes into the other specific allele taking part in the cross Genetic results leading to recombination models Polarity, Conversion and Crossing-over Accurate allele maps are available, there is a gradient, or polarity, of conversion frequencies along the gene Polarity (gradient): the site closer to one end show higher conversion frequency than do the sites farther away from that end Meiosis, crossover and gene conversion Genetic results leading to recombination models Co-conversion Co-conversion: a single conversion event including several sites at once - Frequency of co-conversion increases as the distance between alleles decreases. Holliday Model Formation of heteroduplex DNA Branch migration (along the two heteroduplex strands) Meselson-Radding Model Heterodplex DNA occurred primarily in only one chromatid Double-Strand Break-Repair Model Double strand break, rather than a nick, is the start point Holliday Model of Recombination Formation of heteroduplex DNA -> cross bridge -> branch migration -> mismatch repair -> resolution Holliday Structure: partially heteroduplex double helix Holliday Model of Recombination Branch Migration; the movement of the crossover point between DNA complexes Cross bridge Holliday Model of Recombination Resolution of the Holliday structure Holliday Model of Recombination Application of the Holliday model to genetic crosses Gene conversion & Aberrant ratio ; a consequence of mismatch repair Polarity of gene conversion ; in heteroduplex region Coconversion ; both sites within heteroduplex same excision-repair act Meselson-Radding Model of Recombination (a) (b) (d) (c) Holliday model Could not explain all of cross -> aberrant 4:4 ratio very rare 6:2 ratio frequent -> gene conversion in only one chromatid -> Meselson and Radding Double-Strand Break-Repair Model of Recombination In yeast, induction of double strand break in plasmid stimulates 1000-fold of transformation. -> J. Szostak, T. Orr-Weaver, and R. Rothstein Visualization of recombination intermediates H. Potter and D. Dressler Several Genes involved in general recombination in E.coli recA, recB, recC, recD, SsB(single strand binding protein) RecBCD pathway RecBCD pathway RecA RecF pathway RecE pathway Minor pathway Production of single-stranded DNA - RecBCD protein complex have both nuclease (nicking) and helicase activity (unwinding) - Chi site; 5’- G C TG G T G G -3’ target site for nuclease activity of RecBCD RecA-protein-mediated Single-Strand Exchange - RecA protein can bind to single strand forming a nucleoprotein complex, and catalyze single strand invasion of a duplex forming a D loop Branch Migration RuvA and RuvB protein catalyze branch migration RuvA: bind to crossover point, recruit RuvB RuvB: ATPase hexameric ring motor Resolution of Holliday Junction RuvC: an endonuclease that resolves Holliday junction by symmetric cleavage of the continuous pair of DNA strands (b) 180° rotation of arm I and II Summary of Resolution Pathway + RecA - RecA Recombination produces new gene combinations by exchanging homologous chromosomes. Both genetic and physical evidence has led to several models of recombination Common features of recombination models heteroduplex DNA formation mismatch repair resolution (splicing) The process of recombination itself is under genetic control by numerous genes RecA, B, C, D, E, F and G RuvA, B and C Rus AND ………..