* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Integration of heterogeneous informations sources for
RNA silencing wikipedia , lookup
Non-coding RNA wikipedia , lookup
History of genetic engineering wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Public health genomics wikipedia , lookup
Pathogenomics wikipedia , lookup
Genome evolution wikipedia , lookup
Genomic imprinting wikipedia , lookup
Minimal genome wikipedia , lookup
Genome (book) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Ridge (biology) wikipedia , lookup
Microevolution wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Designer baby wikipedia , lookup
Gene expression programming wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Metagenomics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
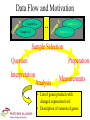
Integration of Heterogeneous Informations Sources for Proteomics and Transcriptomics Steffen Möller University of Rostock Proteome Center Data Flow and Motivation Sample A.a Sample A.1 ... Sample Z.z Sample Z.1 Sample Selection Question Preparation Interpretation Analysis Measurements • List of genes products with changed expression level • Description of variants of genes Data available online • Grouping of samples in homogeneous groups Lab-internal information • Portioning and preparation of samples • Data derived from a preparation – – – – DNA/RNA sequencing Affymetrix Microarrays 2DE Gels (Tandem) mass spectrometry • External bioinformatics databases • Internal extensions to the above – Communication of ideas between researchers Measurements Aids for Interpretation of Data Organisation of Samples Access of MS Spectra • MASCOT peptide identification • MS/MS fragment sequencing Addition to external data sources • Genes discussed among researchers Overview on Identified Spots on Gel • Integration of Protein expression levels – Spot Volume – Spot Area – Spot Peak intensity • with RNA expression levels – from Affymetrix chips Application of Agent Technology • Automated retrieval and integration of presumed relevant in-house data • Assistance in interpretation – Heuristics to extend/shrink list of genes presumed relevant • Integration with external online data – Pathways – Known relevance of genes in other diseases Data Flow Seed of Genes Heuristic Modified List of Genes Adapted for Agents: • Input: List of Gene IDs • Output: List of ( Gene ID Agent ID Evaluation Explanation History ) Examples for Heuristics • Towards extension/shrinking of list of genes under investigation – Gene lies within chromosomal locus linked to disease – Chromosomal neighbourhood to other genes of investigation – Gene is of presumed low abundance • Guidance of further wet-lab analysis – Comparison of ration RNA/protein levels • Search for pre- or post-transcriptional control Example: Interaction with EnsEMBL • Visualisation of QTLs with expression data (G. Fischer et al. 2002, submitted) Transfer from Automated Sequence Annotation • EDITtoTrEMBL (Möller et al. 1998) – Introduction of intermediate level for data integration – Hierarchical organisation of agents Program Integration Program TrEMBL EDITtoTrEMBL: Self-introducing Agents • Sequence-Analysing agents described their input and their output to dispatching agents •SWISS-PROT syntax and controlled vocabulary •Regular expressions as constraints • Dispatchers provide automated planning of annotation path of entries Application in sequence annotation of transmembrane proteins • A variety of programs exist to predict – membrane spanning regions – direction of insertion into the membrane Out In Conflict resolution • Implemented with REVISE (C. V. Damasio; 1997) application described in (S. Möller, M. Schroeder; 2000) Problems with the transfer of these techniques to the wet-lab • Analysers cannot describe themselves or their results – No ontology for methods of expression data analysis has been defined (yet) – The motivation of an analyser to include a gene cannot be formally expressed • No rules for conflict resolution applicable – Conflicts point the unexpected, not to artefacts Discussion • Should I implement the best possible agent system or rather ASAP hunt for the causing agents of autoimmune diseases? • New agents are recruited from Perl scripts that are implemented to provide a quick answer to requests of biological researchers. • Integration on a pragmatical level • The system is accepted by wet-lab researchers. • The system has a PHP-based web-frontend, – communication between agents is implemented via SOAP – adaptations and extensions to the system are easily implemented. Acknowledgements University of Rostock Michael Kreutzer, Gertrud Fischer, Bernd Scheidt, Ines Weber, Angelika Allenberg, Björn Damm, Michael Glocker, Hans-Jürgen Thiesen City University, London Michael Schroeder EMBL-EBI, Cambridge Rolf Apweiler Funded by the BMBF Leitprojekt „Proteom-Analyse des Menschen“ and the Landesforschungsschwerpunkt „Genomorientierte Biotechnologie“