* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lab 8: Atomic force microscopy imaging of cells PI: Lab Instructor: Summary
Signal transduction wikipedia , lookup
Cytokinesis wikipedia , lookup
Cell growth wikipedia , lookup
Programmed cell death wikipedia , lookup
Extracellular matrix wikipedia , lookup
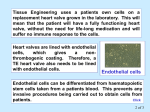
Tissue engineering wikipedia , lookup
Cell culture wikipedia , lookup
Cell encapsulation wikipedia , lookup
Cellular differentiation wikipedia , lookup
List of types of proteins wikipedia , lookup
Lab 8: Atomic force microscopy imaging of cells PI: Krystyn Van Vliet Lab Instructor: Sunyoung Lee Summary In this laboratory, you will use the atomic force microscope to image the structure and stiffness of living and chemically fixed human microvascular endothelial cells. The pN- to nN-scale mechanical force used to create these images allows you to observe both the micrometer-scale height of these cells, as well as the nanometer-scale cytoskeletal network beneath the cell surface. Because the cells are living and imaged under near in vitro conditions, it is possible to observe cell processes in real time, including migration, response to drugs added to the imaging media, and of course apoptosis. It is also possible to compare the nearsurface structure of living and diseased cells. If time allows, you will also observe the near-field optical / fluorescent image of these cell surfaces. Recommended Reading D. Pesen and J. H. Hoh, "Micromechanical Architecture of the Endothelial Cell Cortex," Biophys. J. 88. N. Almqvist et al., "Elasticity and Adhesion Force Mapping Reveals Real-Time Clustering of Growth Factor Receptors and Associated Changes in Local Cellular Rheological Properties," Biophys. J. 86. GEM4 2006 Summer School http://www.openwetware.org/wiki/GEM4labs