* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download The Aerobic Fate of Pyruvate
Basal metabolic rate wikipedia , lookup
Catalytic triad wikipedia , lookup
Mitochondrion wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Photosynthesis wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Butyric acid wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Metalloprotein wikipedia , lookup
Glyceroneogenesis wikipedia , lookup
Lactate dehydrogenase wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Microbial metabolism wikipedia , lookup
Biosynthesis wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Electron transport chain wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Biochemistry wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
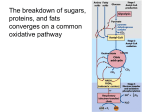
The Aerobic Fate of Pyruvate February 12, 2003 Bryant Miles I could tell that some of you were not impressed by the mere 2 ATPs produced per glucose by glycolysis. The 2 ATP’s produced are only a small fraction of the potential energy available from glucose. Under anaerobic conditions, animals convert glucose into 2 molecules of lactate. Much of the potential energy of the glucose molecule remains untapped. Under Aerobic conditions a much more dynamic pyruvate metabolism occurs. The 2 moles of NADH produced by glyceraldehyde-3-phosphate dehydrogenase are oxidized in the electron transport chain back to NAD+. The electron transport chain generates a proton gradient that drives the synthesis of 5 ATP molecules from ADP and Pi. Further more, the pyruvate formed by glycolysis is converted to acetyl-CoA by pyruvate dehydrogenase (generating another 2 moles of NADH per glucose and another 5 ATPs by oxidative phosphorylation). The acetyl-CoA formed enters into the citric acid cycle where it is completely oxidized into CO2. The electrons liberated by oxidation are captured by NAD+ or FAD which are then transferred via the electron transport chain ultimately to O2, the final electron acceptor. The electron transport chain is coupled to generating a proton gradient which produces a proton motive force that drives the synthesis of ATP. This allows the net production of 32 molecules of ATP to be formed per glucose molecule. The function of the citric acid cycle is to harvest high energy electrons from carbon fuels. The citric acid cycle is the central metabolic hub of the cell, the gateway of aerobic metabolism. The citric acid cycle produces intermediates which are precursors for fatty acids, amino acids, nucleotide bases, cholesterol and porphoryins. The citric acid cycle is shown on the next page. The citric acid cycle occurs in the mitochondria of eukaryotes. New carbons enter the citric acid cycle through acetyl-CoA. The acetyl group may come from pyruvate, fatty acids, ketobodies, ethanol or alanine. The two carbons of acetyl-CoA are transferred to oxaloacetate to yield the first tricarboxylic acid citrate in a reaction catalyzed by citrate synthase. A dehydration followed by a rehydration rearranges citrate into isocitrate. Two successive decarboxylations coupled to the generation of 2 NADH produce succinyl-CoA. Four steps later oxaloacetate is regenerated along with a GTP, FADH2 and NADH. O C Cα Cβ Cleavage site The citric acid cycle may seem like an elaborate way to oxidize acetate into carbon dioxide, but there is chemical logic to the cycle. In order to directly oxidize acetate into two molecules of CO2 a C—C bond must be broken. Under the mild conditions found in cells, there is insufficient energy to break the bond. Biological systems often break C—C bonds between carbon atoms α and β to a carbonyl group. Examples are aldolase and transaldolase. O OH C Cα Cβ Cleavage site Another common type of C—C cleavage is α-cleavage of an α-hydroxy-ketone which we have seen before in transketolase and pyruvate decarboxylase, two TPP dependent enzymes. Neither of these common strategies for cleavage of C—C bonds is available to acetate. Instead the acetate is activated in the form acetyl-CoA, condensed with oxaloacetate to form citrate and then carrying out a β-cleavage in subsequent steps. The Net reaction of the citric acid cycle is: 3NAD+ + FAD + GDP + Pi + Acetyl-CoA + 2H2O 3NADH + FADH2 + GTP + CoA + 2CO2 + 3H+ The Aerobic Fate of Pyruvate In eukaryotes, the reactions of the citric acid cycle occur in the mitochondria. The mitochondrian is enclosed by a double membrane. All of the glycolyitic enzymes are found in the cytosol of the cell. Charged molecules such as pyruvate must be transported in and out of the mitochondria. Small charged molecules with molecular weights of less than 10,000 can freely diffuse through the outer mitochondrial membrane through aqueous channels called porins. A transport protein called pyruvate translocase specifically transports pyruvate through the inner mitochondrial membrane into the matrix of the mitochondrian. Once in the mitochondria, pyruvate dehydrogenase converts pyruvate into acetyl-CoA and CO2. - O O O C C NAD= CH3 NADH + CO2 O + S S CH2 Pyruvate Dehydrogenase NH CH2 C NH O CH2 O CH2 CH2 NH CH2 NH O HO C H H3C C CH3 O N O P O- N O O P N N NH2 C O HO C H H3C C CH3 NH 2 C H2C CH3 CH2 H CH2 C C H2C O N O P O- O N O O P O H O OH H O H P H H O OH H H O- N O O- O O- N O H P O- O- O- Coenzyme A Acetyl Coenzyme A This reaction may look straight forward but it is complex. Pyruvate dehydrogenase is a humongous enzyme that can be visualized by electron micrographs. See Figure 16-3 of text book. Pyruvate dehydrogenase is a multienzyme complex held together by noncovalent interactions. There are three different enzymes and five coenzymes. The three individual enzymes are E1, pyruvate dehydrogenase, E2, dihydrolipoamide transacetylase, and E3, dihyrolipoamide dehydrogenase. The multienzyme complex consists of 24 subunits of E1, 24 subunits of E2 and 12 subunits of E3. In E1 we find a thiamine pyrophosphate coenzyme. In E2 the is a lipoic acid coenzyme. In E3 there is a FAD coenzyme. The molecular weight of pyruvate dehydrogenase isolated from E. coli is 4.6 million Daltons, slightly larger than a ribosome. In mammals, pyruvate dehydrogenase is twice that size and has two additional regulatory subunits. One is a protein kinase which phosphorylates three serine residues in E1, the other is a phosphatase which hydrolytically removes the same phosphoryl groups from the serines of E1. The structure of the pyruvate dehydrogenase complex is shown above. The E2, dihydrolipoamide transacetylase are shown as red balls. These 24 subunits create the core upon which the pyruvate dehydrogenase complex is built. The E1, pyruvate dehydrogenase enzyme is an α2β2 dimer shown as the orange and yellow balls. There are 24 of the αβ dimers in the complex. There are 12 subunits of the E3, dihyrolipoamide dehydrogenase complex shown in purple. The E3 subunits associate into dimers, so there are six E3 dimers in the complex. Lets look at the chemistry going on in this complex. R' R' + R In E1, pyruvate dehydrogenase enzyme, we have a tightly bound thiamine pyrophosphate coenzyme or prosthetic group. We have encountered this cofactor before, pyruvate decarboxylase and transketolase. The reaction catalyzed in this subunit is similar to that of pyruvate decarboxylase. H3 C H3 C + R N N S C S : -C H - O C C O O CH3 The next step in the enzyme catalyzed reaction involves a new cofactor we haven’t encountered yet. It is lipoic acid. H B H3C H3 C R' R' R + R C S C O C C O O S S H H S C - S N CO2 N S CH3 CH3 O O O - O H H HN Lipoic Acid :B H3C H3C R' + R N R C H C R ' + S CH3 C B H H N - :C O O H H S N H O CH3 Lipoamide Lipoic acid is not found free in nature. Instead lipoic acid is covalently attached to a lysine residue through an amide linkage. The enzyme that catalyzes the formation of the lipoamide linkage uses ATP to generate AMP and pyrophosphate. Lipoic acid is an important cofactor because it couples acyl-group transfers with electron transfers during the oxidation and decarboxylation of α-keto acids. Because the ring strain inherent in the disulfide cyclic structure is relieved upon reduction, lipoic acid has a strong negative reduction potential, Eo’ = −0.30 V. 2e- + 2H+ R S R H S H SH HS Eo' = -0.30 V H3C H3C R' + R N R C H N S C B: R' S C CH3 C O O H H CH3 H3C H3C R' R' R R N C S C SH O S C CH3 - S H R B: B H3C R' R C SH O CH3 + R N - C : S S :C O S H R N H S CH3 To summarize the chemistry up to this point, This enzyme has taken pyruvate, used TPP to catalyze the decarboxylation of an α-ketoacid to form the hydroxyethyl-TPP intermediate which is stable enough to isolate. The hydroxyethyl-TPP was then oxidized by and concomitantly transferred to lipoamide to form an acetyllipoamide. The acetyllipoamide is the final product of E1. The lipoamide prosthetic group is part of the E2 Dihydrolipoyl transacetylase enzyme. The next step in the course of the reaction mechanism occurs in the active site of the E2, Dihydrolipoyl transacetylase enzyme, which catalyzes the transfer of the acetyl group from lipoamide to coenzyme A. H S CoA O S C SH O SH CH3 SH + H3C C S CoA R R If you look back to the structure of lipoamide, you will see it has a long flexible tether linking the amide to the E2 enzyme. The length of this tether is about 45 Å. This is long enough to reach the active site of E1 to form the acetyllipoamide and then carry this intermediate to the active site of the E2 enzyme to form acetyl-CoA and the dihydrolipoamide which is then carried to yet the third active site, the E3, dihyrolipoamide dehydrogenase enzyme. R NH2 N N FMN O H2C P O H C OH H C OH H C OH N O O P O O O- O- H H OH OH H N FAD Quinone H3C N H3C N N H O H3C N H N O NH CH2 H3C N . O NH R O FAD H3C N H3C N FADH Semiquinone The E3, dihyrolipoamide dehydrogenase enzyme regenerates lipoamide for another round of catalysis. This enzyme is a flavoprotein, meaning, it has a tightly bound FAD prosthetic group. Flavins undergo 2 electron reductions, but can accept electrons one at a time,unlike NAD+ which can only accept two electrons at a time in the form of a hydride. This gives flavoproteins more catalytic diversity than NAD coenzymes. FADH2 Hydroquinone N . H H3C N H3C N NH O H R O . H N O NH H O Dihydrolipoyl dehydrogenase SH SH R S + FAD + FADH2 S R The final step of the reaction catalyzed by this E3 enzyme is the transfer of the 2 electrons from the tightly bound FADH2 to the transiently bound NAD+ to generate NADH + H+. FADH2 + NAD+ FAD + NADH + H+ This is a usual electron transfer. The common role of FAD is to accept electrons from NADH. One thing about flavoproteins is that the protein bound flavins have a great variety of reduction potentials. In this enzyme the reduction potential is shifted such that FADH2 is the electron donor and NAD+ is the acceptor. Typical flavoprotein FAD + 2e- + 2H+ FADH2 Eo’≈ 0 V NAD + 2e- + H+ NADH Eo’ = -0.315 V