* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Explanation of Scaffold`s Display Options - Proteome Software
Genetic code wikipedia , lookup
Immunoprecipitation wikipedia , lookup
Biochemistry wikipedia , lookup
Index of biochemistry articles wikipedia , lookup
List of types of proteins wikipedia , lookup
Gene expression wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Magnesium transporter wikipedia , lookup
Peptide synthesis wikipedia , lookup
Protein design wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Homology modeling wikipedia , lookup
Protein moonlighting wikipedia , lookup
Protein folding wikipedia , lookup
Interactome wikipedia , lookup
Bottromycin wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Self-assembling peptide wikipedia , lookup
Protein (nutrient) wikipedia , lookup
Protein structure prediction wikipedia , lookup
Western blot wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Protein adsorption wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
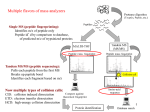
Explanation of Scaffold’s Display Options % of total spectra: This is the percentage of the total number of spectra which is assigned to the protein in question. This number is the number of assigned spectra for this protein divided by the total spectra in the sample (as seen in the Load Data View). Assigned spectra: This is the number of spectra which Protein Prophet assigns to the protein in question.The peptides represented by these spectra may also be present in the sample from a different protein, but the program assigns each spectrum to only the protein (or protein group) for which there is the most evidence, so in this display option, spectra are counted only once. Unique Peptides: This is the number of distinct amino acid sequences represented in the peptides that match to a protein. Peptides with the same a.a. sequence but different charges, or with different modifications are grouped together and counted only once in this count. Unique Spectra: Two spectra are unique if they match different peptides (even if the peptides overlap) or match two different charge states of the same peptide or match both a peptide and a modified form of the peptide. The number of unique spectra is a count of these unique spectra. Only peptides that are "good" or "valid" and are assigned to a given protein count. Percentage coverage: The percentage of all amino acids in the protein sequence that were detected in the sample. Unweighted spectrum count: This is all of the spectral counts which could possibly be assigned to a protein regardless of whether or not Protein Prophet actually assigned the spectra to these proteins. Spectra matching a shared peptide are counted in each protein containing that peptide. Quantitative Values (=Normalized spectral counts): Our protein normalizing entails averaging the unweighted spectral counts for all of the MS-samples and then multiplying the spectrum counts in each sample by the average divided by the individual sample's sum. In all cases, the proteins listed depend on the filter values selected at the top of the window. For the displayed proteins, the actual values of all options except Protein ID Probability depend on the setting for the minimum peptide probability filter.