{PDOC00000} {BEGIN
... asparagine residue is glycosylated, due to the fact that the folding of the protein plays an important role in the regulation of N-glycosylation [2]. It has been shown [3] that the presence of proline between Asn and Ser/Thr will inhibit N-glycosylation; this has been confirmed by a recent [4] stati ...
... asparagine residue is glycosylated, due to the fact that the folding of the protein plays an important role in the regulation of N-glycosylation [2]. It has been shown [3] that the presence of proline between Asn and Ser/Thr will inhibit N-glycosylation; this has been confirmed by a recent [4] stati ...
New Generation Collagens
... Colway’s Natural Collagen is totally different. It is extracted from fish skin in the form of triple helix (spiral) molecules which are precursors of the collagen fibers. So, in simple words, it would have become collagen fibre but it was captured and extracted from fish skin at earlier stages of de ...
... Colway’s Natural Collagen is totally different. It is extracted from fish skin in the form of triple helix (spiral) molecules which are precursors of the collagen fibers. So, in simple words, it would have become collagen fibre but it was captured and extracted from fish skin at earlier stages of de ...
STANDALONE
...The peaks read in are normalized so that the most
intense peak
is set to the dynamic range value. All peaks with values of
less that
1, using this normalization, are not used. This
normalization has the
overall effect of setting a threshold value for peak
intensities.
...
ldentification of Surface-Exposed Domains on the Reducing Side of
... complexes with 20 ng Glu-C/mg Chl (Fig. IB). To determine their relative association with the PSI core, the proteasetreated PSI complexes were filtered using the Centricon-100, and retentates were analyzed by Tricine-urea-SDS-PAGE. The El and EII cleavage products of PsaE were more tightly associate ...
... complexes with 20 ng Glu-C/mg Chl (Fig. IB). To determine their relative association with the PSI core, the proteasetreated PSI complexes were filtered using the Centricon-100, and retentates were analyzed by Tricine-urea-SDS-PAGE. The El and EII cleavage products of PsaE were more tightly associate ...
Forced Expression of Dystrophin Deletion Constructs Reveals
... component of the extracellular matrix in muscle tissue, binds to a-dystroglycan, which binds directly to p-dystroglycan (14). This link to the extracellular matrix has been named the dystroglycan complex. The sarcoglycan complex consisting of the integral membrane proteins c~-, p-, and 7-, sarcoglyc ...
... component of the extracellular matrix in muscle tissue, binds to a-dystroglycan, which binds directly to p-dystroglycan (14). This link to the extracellular matrix has been named the dystroglycan complex. The sarcoglycan complex consisting of the integral membrane proteins c~-, p-, and 7-, sarcoglyc ...
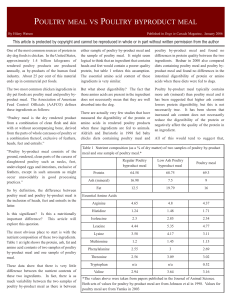
poultry meal vs poultry byproduct meal
... poultry ingredients are of equal quality. There are substantial differences between different grades within each of these ingredients. These differences may be due to differences in processing or to differences in the quality of the parts used to make the ingredient. The process used to produce both ...
... poultry ingredients are of equal quality. There are substantial differences between different grades within each of these ingredients. These differences may be due to differences in processing or to differences in the quality of the parts used to make the ingredient. The process used to produce both ...
Reconstitution of an Allophycocyanin Trimer Complex Containing
... plied to the reconstitution o f (a APß A P ) 3 and of (a APß A P ) 3 • L c 8 9 (G o ttsch alk et al., 1993). D uring this w ork, a n o th e r reconstituted com plex, (a APßAP)3-2 1 -2 3 k D a , had been prepared (G ottschalk et al., 1993), b ut could n ot be further c h a r acterized then because t ...
... plied to the reconstitution o f (a APß A P ) 3 and of (a APß A P ) 3 • L c 8 9 (G o ttsch alk et al., 1993). D uring this w ork, a n o th e r reconstituted com plex, (a APßAP)3-2 1 -2 3 k D a , had been prepared (G ottschalk et al., 1993), b ut could n ot be further c h a r acterized then because t ...
Protein Sequencing Problems
... sequence. Lys has an NH2 on its side chain. That is why FDNB attacked it. Chain = Gly ~ Carb A – Carb A attacks the AA on the other end of the protein (the carboxylic acid side). Since Ser was released, it must have been the last AA in the chain. Chain = Gly ~~~~~~~~~~~~~~~~~~~~~~~~~~Ser Cyanobromid ...
... sequence. Lys has an NH2 on its side chain. That is why FDNB attacked it. Chain = Gly ~ Carb A – Carb A attacks the AA on the other end of the protein (the carboxylic acid side). Since Ser was released, it must have been the last AA in the chain. Chain = Gly ~~~~~~~~~~~~~~~~~~~~~~~~~~Ser Cyanobromid ...
Bacterial protein toxins targeting Rho GTPases
... grated slower, suggesting a di¡erent type of modi¢cation than that induced by CNF [41]. The puzzle was solved when it was recognised that DNT possesses transglutaminase activity. DNT attaches primary amines onto Rho at position Gln63. Further comparison of the enzyme activities of DNT and CNFs revea ...
... grated slower, suggesting a di¡erent type of modi¢cation than that induced by CNF [41]. The puzzle was solved when it was recognised that DNT possesses transglutaminase activity. DNT attaches primary amines onto Rho at position Gln63. Further comparison of the enzyme activities of DNT and CNFs revea ...
Identification and Structural Characterization of the ATP/ADP
... Although the structure of the intact Hsp90 molecule has not yet been determined, crystal structures of an amino-terminal domain identified by limited proteolysis have been determined for yeast (Prodromou et al., 1997) and human (Stebbins et al., 1997) proteins. Consistent with the high homology amon ...
... Although the structure of the intact Hsp90 molecule has not yet been determined, crystal structures of an amino-terminal domain identified by limited proteolysis have been determined for yeast (Prodromou et al., 1997) and human (Stebbins et al., 1997) proteins. Consistent with the high homology amon ...
Investigations of non-enzymatic glycation with new bioanalytical
... elutions. The results clearly demonstrate that the affinity tips bind the highest amount of glucose when the pH of the binding buffer varies between 7.7 and 8.2. Further enhancement of the pH of the binding buffer results in a decrease of the bound glucose. The maximal binding capacity of the tips w ...
... elutions. The results clearly demonstrate that the affinity tips bind the highest amount of glucose when the pH of the binding buffer varies between 7.7 and 8.2. Further enhancement of the pH of the binding buffer results in a decrease of the bound glucose. The maximal binding capacity of the tips w ...
Lesson 24. lmmuno- chemical techniques
... All vertebrates have advanced immune system. The more complex the organism the more advanced the immune system. The immune system of mammals has evolved over a million years; it provides an incredible protection system capable of responding to infective challenges that arise in the body. Immunity in ...
... All vertebrates have advanced immune system. The more complex the organism the more advanced the immune system. The immune system of mammals has evolved over a million years; it provides an incredible protection system capable of responding to infective challenges that arise in the body. Immunity in ...
Detergents and Surfactants
... Since the negative charges repel each other, the positive cationic detergent neutralizes this charge. It may be surprising that it even works because the ammonium (+1) nitrogen is buried under the methyl groups as can be seen in the space filling model Neutral or non-ionic detergents: As the name im ...
... Since the negative charges repel each other, the positive cationic detergent neutralizes this charge. It may be surprising that it even works because the ammonium (+1) nitrogen is buried under the methyl groups as can be seen in the space filling model Neutral or non-ionic detergents: As the name im ...
Soy Allergy Doc - Amherst College
... protein (HVP) in sauces. Multi-grain breads, doughnuts, doughnut mix and pancake mix commonly contain soy flour. Nearly all bread products available in the US now contain soy. Soy can now be found in nearly all types of foods such as meat, ice cream, cheese and french fries. Many foods are contamina ...
... protein (HVP) in sauces. Multi-grain breads, doughnuts, doughnut mix and pancake mix commonly contain soy flour. Nearly all bread products available in the US now contain soy. Soy can now be found in nearly all types of foods such as meat, ice cream, cheese and french fries. Many foods are contamina ...
00:40, 26 August 2010
... BLASTing the protein sequence derived from the nmd gene sequence revealed high levels of similarity between nmd and members of the AAA+ STPase super family. Other members of this family are Spastin and Katanin. These proteins are involved in microtubule depolymerization from the minus and plus ends ...
... BLASTing the protein sequence derived from the nmd gene sequence revealed high levels of similarity between nmd and members of the AAA+ STPase super family. Other members of this family are Spastin and Katanin. These proteins are involved in microtubule depolymerization from the minus and plus ends ...
LS1a Fall 2014 Lab 6: Ribosomal Protein Translation (PyMOL lab #3)
... particles of the microsomal fraction” (try saying that three times fast). It was originally described in this way because just about all that was known about it was that it consisted of RNA (hence the “ribonucleo-”) and protein (hence the, uh, “protein” part of the title). In both bacterial and euka ...
... particles of the microsomal fraction” (try saying that three times fast). It was originally described in this way because just about all that was known about it was that it consisted of RNA (hence the “ribonucleo-”) and protein (hence the, uh, “protein” part of the title). In both bacterial and euka ...
Cell Division: The Place and Time of Cytokinesis Dispatch
... seems to be essential for specifying the site of furrowing. What about the timing? Furrow initiation is known to follow degradation of cyclins B and B3 [16], suggesting that activity of either or both of these two cyclins opposes furrow initiation and that their degradation might contribute to the t ...
... seems to be essential for specifying the site of furrowing. What about the timing? Furrow initiation is known to follow degradation of cyclins B and B3 [16], suggesting that activity of either or both of these two cyclins opposes furrow initiation and that their degradation might contribute to the t ...
Urine
... breakdown of creatine phosphate in muscle tissue. • It is usually produced by the body at a fairly constant rate (which depends on the muscle mass of the body). ...
... breakdown of creatine phosphate in muscle tissue. • It is usually produced by the body at a fairly constant rate (which depends on the muscle mass of the body). ...
Identification and analysis of new phloem proteins from
... (Oparka and Turgeon, 1999; van Bel et al., 2002) and allows the selective import not only of small substances but also of macromolecules such as proteins and nucleic acids from CC into SE (Ayre et al., 2003). The mature PD between SE and CC are branched on the side of the companion cell that turn in ...
... (Oparka and Turgeon, 1999; van Bel et al., 2002) and allows the selective import not only of small substances but also of macromolecules such as proteins and nucleic acids from CC into SE (Ayre et al., 2003). The mature PD between SE and CC are branched on the side of the companion cell that turn in ...
Diversity of Amyloid Motifs in NLR Signaling in Fungi
... transduction was first identified in the NWD2/HET-S system and further generalized to other NLR and effector protein pairs [8–11] as will be described herein (Figure 1B). Description of this mode of amyloid signaling stems from the study of a fungal non-self recognition phenomenon known as heterokar ...
... transduction was first identified in the NWD2/HET-S system and further generalized to other NLR and effector protein pairs [8–11] as will be described herein (Figure 1B). Description of this mode of amyloid signaling stems from the study of a fungal non-self recognition phenomenon known as heterokar ...
Isolation of Casein, Lactose and Albumin from Milk
... lecithins (phospholipids conjugated with choline). The structures of phospholipids and lecithins are shown. The phospholipids help to stabilize the whole milk emulsion; the phosphate groups help to achieve partial water solubility for the fat globules. All the fat can be removed from milk by extract ...
... lecithins (phospholipids conjugated with choline). The structures of phospholipids and lecithins are shown. The phospholipids help to stabilize the whole milk emulsion; the phosphate groups help to achieve partial water solubility for the fat globules. All the fat can be removed from milk by extract ...
Protein Cleavage Due to Pro-oxidative Activity in Some Spices
... Fragmentation of BSA The fragmentation of BSA in three different systems were observed on SDSPAGE (Fig. 1, 2 and 3). BSA was fragmented markedly by oxidation reaction in the Cu2+/H2O2 system since the oxidizing power of the Fe2+/ascorbate and Fe3+/ EDTA/H2O2 on the fragmentation of BSA were much wea ...
... Fragmentation of BSA The fragmentation of BSA in three different systems were observed on SDSPAGE (Fig. 1, 2 and 3). BSA was fragmented markedly by oxidation reaction in the Cu2+/H2O2 system since the oxidizing power of the Fe2+/ascorbate and Fe3+/ EDTA/H2O2 on the fragmentation of BSA were much wea ...
PDF - Beilstein
... in a variety of RiPPs that also feature conventional lanthionine rings, such as epidermin [63] (Figure 5A), mersacidin [64] and cypemycin [65]. In epidermin, a S-[(Z)-2-aminovinyl]-Dcysteine (AviCys) residue is formed by the 1,4-nucleophilic addition of an oxidatively decarboxylated cysteine residue ...
... in a variety of RiPPs that also feature conventional lanthionine rings, such as epidermin [63] (Figure 5A), mersacidin [64] and cypemycin [65]. In epidermin, a S-[(Z)-2-aminovinyl]-Dcysteine (AviCys) residue is formed by the 1,4-nucleophilic addition of an oxidatively decarboxylated cysteine residue ...
Identification of the Factors Responsible for the Interaction of
... Identification of Hydrophobic patches by Surface Hydrophobicity plot: Kyte-Doolittle is an online software widely applied scale for determining hydrophobic character of a protein [18]. Regions with values above ‘0’ are hydrophobic in character. Setting window size to 5-7 is suggested to be a good va ...
... Identification of Hydrophobic patches by Surface Hydrophobicity plot: Kyte-Doolittle is an online software widely applied scale for determining hydrophobic character of a protein [18]. Regions with values above ‘0’ are hydrophobic in character. Setting window size to 5-7 is suggested to be a good va ...
Protein mass spectrometry

Protein mass spectrometry refers to the application of mass spectrometry to the study of proteins. Mass spectrometry is an important emerging method for the characterization of proteins. The two primary methods for ionization of whole proteins are electrospray ionization (ESI) and matrix-assisted laser desorption/ionization (MALDI). In keeping with the performance and mass range of available mass spectrometers, two approaches are used for characterizing proteins. In the first, intact proteins are ionized by either of the two techniques described above, and then introduced to a mass analyzer. This approach is referred to as ""top-down"" strategy of protein analysis. In the second, proteins are enzymatically digested into smaller peptides using a protease such as trypsin. Subsequently these peptides are introduced into the mass spectrometer and identified by peptide mass fingerprinting or tandem mass spectrometry. Hence, this latter approach (also called ""bottom-up"" proteomics) uses identification at the peptide level to infer the existence of proteins.Whole protein mass analysis is primarily conducted using either time-of-flight (TOF) MS, or Fourier transform ion cyclotron resonance (FT-ICR). These two types of instrument are preferable here because of their wide mass range, and in the case of FT-ICR, its high mass accuracy. Mass analysis of proteolytic peptides is a much more popular method of protein characterization, as cheaper instrument designs can be used for characterization. Additionally, sample preparation is easier once whole proteins have been digested into smaller peptide fragments. The most widely used instrument for peptide mass analysis are the MALDI time-of-flight instruments as they permit the acquisition of peptide mass fingerprints (PMFs) at high pace (1 PMF can be analyzed in approx. 10 sec). Multiple stage quadrupole-time-of-flight and the quadrupole ion trap also find use in this application.