* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Third HANDOUT
Gene desert wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene therapy wikipedia , lookup
Point mutation wikipedia , lookup
Gene expression programming wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome (book) wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genetically modified crops wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Protein moonlighting wikipedia , lookup
Gene expression profiling wikipedia , lookup
Microevolution wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genetic engineering wikipedia , lookup
Gene nomenclature wikipedia , lookup
Helitron (biology) wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Pathogenomics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Designer baby wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
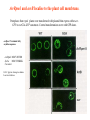
Recent advances in understanding gene –for – gene interactions Avirulence Gene Function – what is currently known Avr Pathogen Protein Homology with and/or possible virulence function * Location in the plant cell NK ‡ Common domain for avirulence and virulence function NK Location of the matching R gene * NK AvrPphC Pseudomonas syringae pv. phaseolicola Interferes with the plant defense response AvrPphF Pseudomonas syringae pv. phaseolicola Interferes with the plant defense response NK NK NK AvrRpt2 Pseudomonas syringae pv. tomato Interferes with the plant defense response NK NK NK AvrRpm1 Pseudomonas syringae pv. maculicola Enhances growth Plasma membrane [ NK Plasma membrane AvrPto Pseudomonas syringae pv. tomato Enhances growth Degradation of resistance gene Plasma membrane No NK AvrBst Xanthomonas campestris pv. campestris Ubiquitin-like protease NK Yes NK AvrXa7 Xanthomonas oryzae pv. oryzae Transcriptional activation domain required for virulence Nucleus Yes NK AvrBs2 Xanthomonas campestris pv. vesicatoria Agrocinopine synthase (from Agrobacterium tumefaciens) NK Recognition region localized in the portion NK with highest homology to synthase AvrBs3 Xanthomonas campestris pv. glycinea Transcriptional activation domain required for virulence Nucleus Yes NK PthA Xanthomonas citri Transcriptional activation domain required for virulence/causes cell hyperplasia [53] Nucleus NK NK Avr9 Cladosporium fulvum NK Exoplasmic NK External plasma membrane Avirulence genes promote virulence Chen et al., (2000) The Pseudomonas syringae avrRpt2 Gene Product Promotes Pathogen Virulence from Inside Plant Cells. Molecular Plant-Microbe Interactions Vol. 13: 1312–1321 Disease symptoms and growth of Pseudomonas syringae pv. tomato strain DC3000 (PstDC3000) on Arabidopsis thaliana leaf tissue. A, Diseasesymptoms caused by PstDC3000 on A. thaliana No-0 rps2 (top) and Col-0 rps2 (bottom) plants. Leaves are shown 4 days after inoculation with PstDC3000 (left) and PstDC3000(avrRpt2) (right). Plants were inoculated by dipping them into bacterial suspensions (3 to 5 × 10 8 CFU/cm 2 ) containing the surfactant Silwet L77. B, Growth of PstDC3000 and PstDC3000(avrRpt2) in leaf tissue of Col-0 rps2, No-0 rps2, and No-0 RPS2 plants.. In B, A. thaliana plants were inoculated by vacuum infiltration with the indicated PstDC3000 strains at an initial density of 1 × 10 5 CFU/cm 2 . The concentration of bacteria in the plant leaves was assayed after 0, 2, and 4 days. Datapoints represent means of three independent determinations ± standard error of the mean. Experiments presented in A, B, were carried out a minimum of three times with similar results Virulence genes suppress avr/R gene interactions Chen et al., (2000) The Pseudomonas syringae avrRpt2 Gene Product Promotes Pathogen Virulence from Inside Plant Cells. Molecular Plant-Microbe Interactions Vol. 13: 1312–1321 RPM1::MYC is a peripheral PM protein. Anti-myc antibody can be used to detect myclabelled RPM1 C-Myc epitope has been “artificially ” added to RPM1 Plasma Membrane Cytoplasm and ER Fractionation of RPM1::MYC on sucrose gradients. A 12-55% (wt/vol) linear sucrose gradient was used to fractionate total extract from transgenic plants. Aliquots of each fraction were blotted to nitrocellulose and were analyzed with either anti-c-Myc or the subcellular compartment marker antibodies listed at the right of each panel. Boyes et al., (1998) The Arabidopsis thaliana RPM1 disease resistance gene product is a peripheral plasma membrane protein that is degraded coincident with the hypersensitive response. 95, PNAS USA 1584915854 AvrRpm1 and avrB localise to the plant cell membrane. Protoplasts from rpm1 plants were transformed with plasmid that express either avrGFP or avrG2A-GFP constructs. Control transformations were with GFP alone. avrRpm1 N-terminal fatty acylation sequence avrRpm1 MGCVSSTSR Soc3a MGCSVSKKK Ca sensor G2A = glycine change to alanine Loss in avirulence RPM1::MYC is degraded after inoculation with avirulent P. syringae isolates, which trigger LZ-NBS-LRR resistance genes Hours post inoculation Boyes et al., (1998) The Arabidopsis thaliana RPM1 disease resistance gene product is a peripheral plasma membrane protein that is degraded coincident with the hypersensitive response. 95, PNAS USA 15849-15854 AvrXa7 : a nuclear targeted protein PX086mx53 = NLS- version Resistant Susceptible NORMAL avrXa7 avrXa7 avrXa7 over express NLSexpress NORMAL avrXa7 avrXa7 avrXa7 over express NLSexpress Fig. 1. AvrXa7 is a DNA-binding protein. Double-stranded oligonucleotides were end labeled with 32P, mixed with AvrXa7 or GST (glutathione S-transferase), and subjected to polyacrylamide electrophoresis. Arrow indicates position of well. Yang et al., (2000) The virulence factor AvrXa7 of Xanthomonas oryzae pv. oryzae is a type III secretion pathwaydependent nuclear-localized doublestranded DNA-binding protein.Proc Natl Acad Sci U S A 15;:9807 “Guard” Hypotheses