* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Barcode - Statistical Center for HIV/AIDS Research and Prevention
Human genome wikipedia , lookup
Essential gene wikipedia , lookup
Gene expression programming wikipedia , lookup
Genetic engineering wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Genomic imprinting wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Pathogenomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genomic library wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Ridge (biology) wikipedia , lookup
Public health genomics wikipedia , lookup
Designer baby wikipedia , lookup
History of genetic engineering wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Helitron (biology) wikipedia , lookup
Gene expression profiling wikipedia , lookup
Microevolution wikipedia , lookup
Minimal genome wikipedia , lookup
Genome (book) wikipedia , lookup
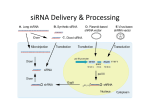
Finding Host Proteins Required for HIV Replication Abe Brass Partners AIDS Research Center and GI Unit Mass. General Hospital Harvard Medical School How do we find what HIV needs to replicate? Rationale • Employ new methods in mammalian genetics to find host factors that HIV depends upon (HDFs). • HDFs provide targets for anti-retroviral therapy and chemical prophylaxis. Rationale • HIV may be hard pressed to evade HDF-directed therapies. • Comprehensive information about the lifecycle of the virus will benefit the HIV research community. Overview • • • • • RNAi Mechanism RNAi Tools Finding HIV-dependency factors Three HIV siRNA Screens TNPO3 RNAi Mechanism RNAi Tools Genetic Screens • Deplete protein expression with shRNAs or siRNAs. • Test how depletion impacts phenotype with simple in vitro functional assay. • Unbiased whole genome screens bring new targets into the “pipeline”. Genetic Screens • The way a genetic screen is designed can profoundly influence which genes are uncovered • Different screen platforms yield different results (i.e. libraries, viruses, cell lines, transfection conditions and efficiencies, readouts) • Some weak hits may be the most important unlike small molecule screens (knockdown efficiency unknown). Caveats • False positives (OTEs). Present, but minimized through bioinformatic functional clustering, expression studies and reagent redundancy. • False negatives. Why didn’t this host factor score? Saturation is the goal, but hard to obtain by generating hypomorphs with the current siRNA technology. Optimized validated reagents will help. shRNA Libraries •Whole genome, 3 shRNAs/gene •Packaged into Retroviral Pools, Stable knockdown •Focused Libraries: Kinases, Ubiquitin pathway •shRNAs have unique barcodes •Formats : MSCV, lentivirus, Inducible. shRNA Libraries mir30-shRNA Retrovirus LTR mir30-5’ mir30-3’ barcode LTR Phenotype Integration mir30-shRNA Processing of shRNA Target mRNAsingle copy knockdown shRNA Libraries Control Experimental PCR recovery of and color label of barcode Competitive hybridization to barcode microarray shRNA dropped out following selection shRNA enriched following selection Barcode: unique 60 nt sequences that allow the abundance of any particular retroviral shRNA in a complex population to be followed using microarray hybridization. siRNA Libraries • Arrayed format-”one gene per well” • High throughput whole genome screens done with liquid handling robotics. • Transiently transfect siRNAs, RISC active for 6 days post transfection. • Image based=scanning microscope. • Reporter gene=plate reader. Finding HIV-dependency factors Luciferase Percent infected 120 90 Part one 60 30 0 CD4 Tat Beta-gal (RLU) 400000 300000 200000 100000 0 Part two siRNA Library • Dharmacon: SMARTpool library, 4 siRNAs per pool, whole human genome (21,121 genes). • Initial screen done with pools. • Validation round done with the four individual siRNAs. Screen Results • 16 genes scored with • Out of 21,121 4/4 individual siRNAs genes, 386 scored 2 SD below control • 44 genes 3/4 • 99 genes 2/4 • 1.8% hit rate • 115 genes 1/4 • 273 of 386 siRNA pools were confirmed • Each of the four by at least 1 siRNA individual siRNA (71%) were retested • Reagent Redundancy tries to minimize OTEs, but some of the ¼ are “knowns”. Known Host Factors Found in the Screen A4GALT (2/4 siRNAs) DDX3X (2) PSME2 (1) AKT1 (2) ERCC3 (3) PURA (2) AP2M1 (1 ) FBXW11 (4) Rab9p40 (3) Arf1 (2) GCN5L2 (1) RANBP1 (1) CD4 (2) H3F3A (1) RelA (4) CD147 (3) HRS (SP) SIP1 (1) CRTC2 (1 ) HTATSF1 (1) ST3GAL5 (1) CRTC3 (3 ) IKBG (2) TFAP4 (3) CTDP1 (1) La Autoantigen (2) TFE3 (2) CXCR-4 (2) FAPP1 (1) VPRBP (1) CyclinT1 (1) NMT1 (3 ) ZNRD1 (2) Biologic Processes Enzymes Found in the Screen ADAM10 (3/4 siRNAs) WNK1 (3) Jak1 (1) DDX55 (3) PSPHL (2) USP26 (2) DDX49 (2) DPM1 (2) OTUD3 (2) ATPV0A1 (2) OST48 (3) LPL (2) GAPVD1(2) PRKX1 (1) HUWE1 (2) PIGH (1) STT3A (2) HERC3 (3) PIGY (2) RNF170 (3) EXOD1(2) ABGL5 (2) FNTA(4) HERC6 (2) FLJ32569 (2) ALKBH8 (2) DDX33 (2) ITPKA (2) NMT1 (3) SET7 (3) MOS (2) DIMT1L(4) ARF1 (2) PIP5K1C (3) C14orf125 (3) CENTG1 (3) Three HIV siRNA Screens Comparison of Three HIV siRNA Screens Cells Zhou HeLa-Bgal KD Virus Infection Readout Hits 24 h HXB2 48 hr; 96 hr β-gal reporter activation 48 h NL 4.3 luc vector, VSV-G 24 hr Luc reporter 295 72 h HIV-1- 48 hr; 48 hr in IIIB new cells p24 ; reporter activation 280 et al Konig 293T et al Brass et al TZM-bl, HeLa Stephen Goff Cell 135 2008 207 Konig et al Zhou et al 295 207 9 13 18 280 Brass et al 783 total RelA Med6 Med7 HDFs Found by Two Independent Screens ADRBK1 AKT1 ANAPC2 ANKRD30A (1) Cav2 CD4 CHST1 CTDP1 (1) CXCR4 CyclinT1 (1) WNK1 DDX3X DMXL1 IDH1 (1) Jak1 (1) MAP4 Med14 Med19 Med28 Med4 Med6 TRIM55 Med7 MID1IP1 Mre11A Nup153 Nup155 Rab28 (1) RanBP2 RelA RNF26 (1) TCEB3 TNPO3 Why so little overlap? • Different screen platforms yield different results. • Technology caveats: False positives (OTEs) and false negatives (reagents not validated). • High throughput methodologies • Secondary filters of the primary data TNPO3 SR 1.2 HIV LTR Late RT 1.2 0.8 0.8 0.4 0.4 0 0 Virion RTC PIC PIC Provirus 1.4 1.2 2-LTR 1 0.8 0.6 0.4 0.2 0 Conclusions • Functional enrichment, finding known factors, and confirmation studies suggests the majority of the genes found in the screens impact HIV replication. • Clearly conventional validation work is required, but host factor discovery has been accelerated (33+ functionally confirmed host factors in 10 months). • Two screens yielded TNPO3, which is very likely the factor that permits HIV-1, HIV-2, EIAV, and SIV access to their host’s genomes. Thank You • Stephen J. Elledge, HHMI, BWH, HMS • Judy Lieberman, Nan Yan, Derek Dykxhoorn, IDI, Childrens Hospital • Ramnik Xavier, Yair Benita, CCIB, MGH • Caroline Shamu and staff, ICCB-L HMSFelipe Diaz-Griffero, DFCI, HMS • Bill and Melinda Gates Foundation • Harvard CFAR