* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download presentation_courese_wed_3
Medical genetics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Human genetic variation wikipedia , lookup
Heritability of IQ wikipedia , lookup
Behavioural genetics wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Designer baby wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene expression programming wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Genome (book) wikipedia , lookup
Public health genomics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Expanded genetic code wikipedia , lookup
Point mutation wikipedia , lookup
Genetic code wikipedia , lookup
Identification and evaluation of
causative genetic variants
corresponding to a certain
phenotype
Xidan Li
Outline
• SIT - identify and evaluate the causative genetic
variants within a QTL/GWAS defined region.
• PASE - evaluate the effect of amino acid
substitution to the hosting protein function
• DIPT - to identify causative genes underlying an
expression phenotype
• Parallelizing computing
Genetic variances identification
Possible solutions?
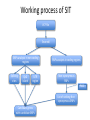
Working process of SIT
VCF file
Ensembl
SNPs analysis in non-coding
regions
Splicing
sites
CpG
island
UTR
region
SNPs analysis in coding regions
Non-synonymous
SNPs
PASE
List of ranking Nonsynonymous SNPs
Candidate genes
with candidate SNPs
Sample results
Non-synonymous SNPs are ranked
The life is easy!
Amino acid substitutions effects
prediction
Effect of amino acid substitutions
Selected seven physico-chemical
properties of Amino acids
Seven Physiochemical properties of
Amino acid
Transfer free energy from octanol to
water
Normalized van der Waals volume
Isoelectric point
Polarity
Normalized frequency of alpha-helix
Free energy of solution in water
Normalized frequency of turn
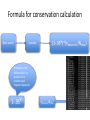
Formula for conservation calculation
Blast search
clustalw
(1-.95N)*(nobserved /Ntotal)
Probability of 20
different AAs in a
position for N
random equal
frequent sequences.
1-.95N
nobserved /Ntotal
Protein kinase AMP-activated gamma
3 (PRKAG3) gene
• (R200Q) in AMPK3 in purebred Hampshire pigs – RN
• (V199I) in AMPK3 Co-participate in the effective
process with R200Q
• RN that causes excess glycogen content in pig skeletal
muscle
•
Milan D, et. al. (2000). A mutation in PRKAG3 associated with excess glycogen content in pig skeletal muscle.
Science 288 (5469): 1248–51.
•
Ciobanu,D, et. al. (2001). Evidence for New Alleles in the Protein Kinase Adenosine Monophosphate-Activated 3-Subunit
Gene Associated With Low Glycogen Content in Pig Skeletal Muscle and Improved Meat Quality. Genetics, 159, 1151-1162.
(R200Q) Cause major increase in the muscle glycogen
content
(V199I) Contribute with smaller effect
Genes ID
Coordinate
REF ALT Conservations score
(MSAC)
PRKAG_3
200
R
Q
PRKAG_3
199
V
I
PASE
score
PASEC
(combined)
score
0.93
0.54
0.50
0.85
0.14
0.12
Ciobanu,D, et. al. (2001). Evidence for New Alleles in the Protein Kinase Adenosine Monophosphate-Activated 3-Subunit Gene
Associated With Low Glycogen Content in Pig Skeletal Muscle and Improved Meat Quality. Genetics, 159, 1151-1162.
Testing with SIFT and POLYPHEN
SIFT
PolyPhen
Conservation
scores
(MSAC)
PASE scores
(Physicochemical
properties
changings)
PASEC score
(combined)
Tolerated (1987)
0.47
0.39
0.18
Deleterious (1351)
0.60
0.51
0.30
Benign (1637)
0.44
0.37
0.16
Possibly damaging
(539)
0.56
0.43
0.24
Probably damaging
(1162)
0.63
0.53
0.33
Features
• Other tool
SIFT, PolyPhen
MAINLY rely on calculating sequence conservation scores (finding homologous
sequences).
• PASE
not only uses the physico-chemical property changing score, but also combine
with sequence conservation score
Potentially being able to analyze the evolutionary-distant protein sequence
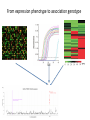
From expression phenotype to association genotype
Sample result of DIPT
www.computationalgenetics.se/DIPT/
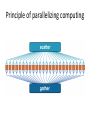
Parallelizing computing
Principle of parallelizing computing
Multiple threads – efficient work
Single thread - tough job!
• Usually in the loop
• Data must be independent
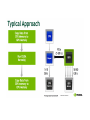
GPU vs. CPU
Cuda Vs. C
#include <cuda.h>
#include <stdio.h>
// Prototypes
__global__ void helloWorld(char*);
// Host function
int main(int argc, char** argv)
{
int i;
// desired output
char str[] = "Hello World!";
// mangle contents of output ; the null character is left intact for simplicity
for(i = 0; i < 12; i++) str[i] -= i;
// allocate memory on the device
char *d_str;
size_t size = sizeof(str);
cudaMalloc((void**)&d_str, size);
// copy the string to the device
cudaMemcpy(d_str, str, size, cudaMemcpyHostToDevice);
// set the grid and block sizes
dim3 dimGrid(2); // one block per word
dim3 dimBlock(6); // one thread per character
// invoke the kernel
helloWorld<<< dimGrid, dimBlock >>>(d_str);
// retrieve the results from the device
cudaMemcpy(str, d_str, size, cudaMemcpyDeviceToHost);
// free up the allocated memory on the device
cudaFree(d_str);
// everyone's favorite part
printf("%s\n", str);
return 0;
}
// Device kernel
__global__ void helloWorld(char* str)
{
// determine where in the thread grid we are
int idx = blockIdx.x * blockDim.x + threadIdx.x;
// unmangle output
str[idx] += idx;
}
#include <stdio.h>
int main(void)
{
printf("Hello World\n");
return 0;
}
Thank You!