* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Sunday, 28 October 2007
Quantitative trait locus wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Gene therapy wikipedia , lookup
Oncogenomics wikipedia , lookup
Genetic engineering wikipedia , lookup
Genome evolution wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Public health genomics wikipedia , lookup
Genomic imprinting wikipedia , lookup
Genome (book) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
Microevolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene expression programming wikipedia , lookup
Nutriepigenomics wikipedia , lookup
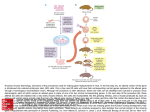
Microarray Analysis of Genes Orchestrating Craniofacial Development Leo R. Otake, MD, PhD and John A. Persing, MD. Specific Aims The objective of this project is to identify candidate interacting genes which are temporally differentially expressed during craniofacial development using the mouse animal model. The Affymetrix GeneChip Mouse Genome 430 2.0 Array has been utilized in this investigation. As the molecular underpinnings of craniofacial development are not well understood at the present time, modalities such as molecular diagnostics and gene therapy lie at early stages of development. A greater understanding of the genetic network involved in craniofacial development would be an asset towards elucidating the molecular mechanisms underlying craniofacial development in general as well as craniosynostosis in particular. Significance Although genetic lesions associated with various congenital craniofacial syndromes have been identified, the interacting genetic networks/pathways that result in the syndromic craniosynostoses have not been clearly elucidated. The candidate human genes and mutations which have been associated with syndromes have been reported, including the homeobox protein MSX2 with Boston-type craniosyostosis, the family of fibroblast growth factor receptors (FGFR) type 1 (FGFR1) with Pfeifer type I, FGFR2 with Apert, Crouzon, and Pfeifer types I/II/III, FGFR3 with Apert/Acanthosis Nigricans and the transcription factor TWIST with Saethre-Chotzen. A comprehensive examination allows for evaluation of the complex interplay of pathways which lead to the clinical phenotype. A compelling question lies in the observation that although Apert and Crouzon Syndromes are both associated with mutations in FGFR2, the clinical presentation on physical examination and temporal progression of craniosynostoses, as well as, mental functional outcome with Apert patients being more severely affected, differ so greatly; this suggests that alternative or differential dysregulation of similar pathways occurs downstream of the identified genetic lesion. Experimental Design and Methods Dr. Hongbo Chi and Dr. Richard Flavell of the Department of Immunobiology, Yale School of Medicine; have developed a targeted, single locus gene knockout of the mouse MAP kinase kinase kinase 4 (MEKK4) gene. Mice which are homozygous for the gene knockout are viable to birth, but die shortly thereafter. These mice demonstrate severe derangements in craniofacial development, notably, the cranial vault fails to develop with attendant exencephaly and spina bifida. Additionally, the cortex fails to develop normally in the knockout mice. The development of such a striking craniofacial defect in the setting of a targeted, single gene knockout presents a unique opportunity to examine, using microarray technology, specific genetic pathways which have been disrupted. Heads from E18 homozygous MEKK4 knockout and homozygous normal fetuses from the same mating were collected. Total cellular RNA was extracted and matched pairs of homozygous knockout and normal samples were applied to the Affymetrix mouse 430 v 2.0 total genomic microarray and quantitated using GeneSpring GX 7.3 (Agilent Technologies). Synopsis of Results Results from 2 pairs of Mekk4 wild-type and gene knockout E18 fetuses (total of 4 animals) are shown in table form (Table1). Microarray data were normalized and fold-changes in expression were determined and segregated into functional groups. Representative genes are presented if an alteration of expression in the given locus resulted in a reported phenotype in the literature relevant to craniofacial development. Cell Differentiation The locus wnt5a was found to be down regulated 1.85 fold in KO vs wild type littermates. Wnt5a has been shown to be required for cranial neural crest cell migration in zebra fish (Barallo-Gimeno et al., 2003); in mice the midline interfrontal suture is neural crest derived in contrast to the sagittal suture which has neural crest cells interposed between the parietal bones which are derived from mesoderm (Wang et al. 2005). Locus Forkhead Box G1 (FoxG1) was down regulated 1.50 fold in KO littermates. This locus has been implicated in the development of the telencephalon in mouse (Storm et al. 2006). The locus endothelin receptor B which was upregulated 1.54 fold in KO mice had been implicated in congenital anomalies associated with Hirschprung’s Disease; of note these anomalies include craniofacial, skeletal, and limb anomalies (Moore, SW, 2006). Extracellular Space Expression levels of biglycan (bgn) and decorin (dcn), are decreased1.74 and 1.87 fold, respectively, in KO mice. As loss of bgn and dcn have been shown to lead to osteogenic cell apoptosis as well as to decreased sequestration of tgf-1 in the ECM (Bi et al., 2005), the loss of these two mediators of bone development in the KO mice would be consistent with the lack of calvarial development which is observed. Interestingly, a 1.67 fold decrease in the fibrillin locus was noted in KO mice; this locus was shown to be part of a genomic deletion resulting in craniofacial abnormalities and mental delay in a human patient (Garcia-Minaur et al., 2005). In contrast, overexpression of the transthyretin gene in mice was found to result in fronto-nasal hypoplasia (Noguchi et al., 2002); this locus is upregulated 1.60 fold in the KO mouse and would be consistent with its hypoplastic phenotype. Extracellular Matrix The expression level of procollagen XIa1 was 2.55 fold decreased in the KO mouse. Interestingly, a defect in collagen XIa1 has been associated with human Marshall Syndrome, the presentation of which includes craniofacial abnormalities (Griffith et al., 1998). Receptor Activity The expression of EphA4 was decreased 1.89 fold in KO mice. The expression of EphA4 has been shown to be associated with patterning of cranial neural crest cells (Ishii et al, 2005). Given that in mice the midline interfrontal suture is neural crest derived in contrast to the sagittal suture which has neural crest cells interposed between the parietal bones which are derived from mesoderm (Wang et al. 2005), alterations in the expression of this gene would be consistent with the calvarial defect noted in the KO mouse phenotype. Table 1: Functional Groupings of Genes with Altered Expression Levels Cell Differentiation Wnt5a -1.85 Forkhead BoxG1 -1.50 Endothelin Recept B +1.54 Extracellular Space Biglycan -1.74 Dermatopontin -1.87 Decorin -1.55 Fibrillin -1.67 Endothelin Recept-B +1.54 Transthyretun +1.60 Extracellular Matrix Procollagen XIa1 -2.55 Fibrillin2 -2.34 Receptor Activity Eph Receptor A4 -1.89 Future Investigations Subsequent to the described examination of the gene expression profiles in the heads of the MEKK4 knockout mouse at the E18 stage of development, we propose to extend our analyses to compare expression profiles of other mutant mice with craniofacial defects. Starting with the initial state of absent calvarial vault development in the MEKK4 mouse, our objective is to gain a greater understanding of the repertoire of genes needed to orchestrate calvarial development in both a normal and abnormal synostotic state. References Barrallo-Gimeno1,A., Jochen Holzschuh,, Wolfgang Driever and Ela W. Knapik. (2003). Neural crest survival and differentiation in zebrafish depends on mont blanc/tfap2a gene function. Development 131, 1463-1477 Bi, Y, Christina H. Stuelten, Tina Kilts, Sunil Wadhwa, Renato V. Iozzo, Pamela G. Robey, Xiao-Dong Chen, and Marian F. Young. (2005). Extracellular Matrix Proteoglycans Control the Fate of Bone Marrow Stromal Cells. Journal of Biological Chemistry.Vol. 280, No. 34, Issue of August 26, pp. 30481–30489, Garcia-Minaur, S., Jacqueline Ramsay, Elizabeth Grace, Robert A. Minns,Lynn M. Myles, and David R. FitzPatrick (2005). Interstitial Deletion of the Long Arm of Chromosome 5 in a Boy With Multiple Congenital Anomalies and Mental Retardation: Molecular Characterization of the Deleted Region to 5q22.3q23.3. American Journal of Medical Genetics 132A:402–410. Griffith, A, Leslie K. Sprunger,1 D. Alexa Sirko-Osadsa,3 George E. Tiller,5 Miriam H. Meisler,1 and Matthew L. Warman (1998). Marshall Syndrome Associated with a Splicing Defect at the COL11A1 Locus. Am. J. Hum. Genet. 62:816–823. Ishii1, M, Jun Han, Hai-Yun Yen, Henry M. Sucov, Yang Chai and Robert E. Maxson, Jr. (2005). Combined deficiencies of Msx1and Msx2cause impaired patterning and survival of the cranial neural crest. Development 132, 4937-4950. Moore, SW (2006). The contribution of associated congenital anomalies in understanding Hirschsprung’s disease. Pediatr Surg Int (22): 305–315. Noguchi, H, Tadashi Kaname, Tomohisa Sekimoto, Kei Senba, Yasushi Nagata, Masatake Araki, Makoto Abe ,Naomi Nakagata, Tomomichi Ono, Ken-ichi Yamamura, and Kimi Araki (2002). Naso-maxillary deformity due to frontonasal expression of human transthyretin gene in transgenic mice. Genes to Cells (2002) 7, 1087–1098. Storm, Elaine E, Sonia Garel, Ugo Borello, Jean M. Hebert, Salvador Martinez, Susan K. McConnell, Gail R. Martin and John L. R. Rubenstein. (2006). Dose-dependent functions of Fgf8 in regulating telencephalic patterning centers. Development 133, 1831-1844. Wang, Y, Ran Xiao1, Fan Yang, Baktiar O. Karim, Anthony J. Iacovelli, Juanliang Cai1,Charles P. Lerner, Joan T. Richtsmeier, Jen M. Leszl, Cheryl A. Hill, Kai Yu, David M. Ornitz, Jennifer Elisseeff, David L. Huso and Ethylin Wang Jabs (2005). Abnormalities in cartilage and bone development in the Apert syndrome FGFR2+/S252W mouse. Development 132, 3537-3548.