* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Site-specific mutagenesis of M13 clones
Gel electrophoresis of nucleic acids wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Community fingerprinting wikipedia , lookup
Gene expression wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Point mutation wikipedia , lookup
Molecular cloning wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Genome evolution wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Non-coding DNA wikipedia , lookup
Molecular evolution wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Silencer (genetics) wikipedia , lookup
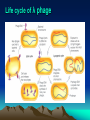
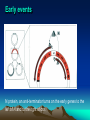
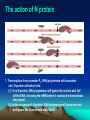
Site-specific mutagenesis of M13 clones (5) Site-specific mutagenesis of M13 clones i. Site-specific mutagenesis involves making a predetermined change in the DNA sequence, unlike random mutagenesis, where the change is made by chance. (Fig. 7.15 and Fig. 7.16) ii. The major difficulty for the method of Fig.7.15 is how to know which plaque contains the mutated DNA. iii. In Fig. 7.16, thymines in the M13 cloning vector are replaced with uracils by propagating the phage in a dUTPase- and uracil-N-glycosylase-deficient (Dur-Ung-) host. (i) When the double-stranded RF molecules are synthesized from such template, the newly synthesized strand contains thymines while the template strand still contains uracil. Site-specific mutagenesis of M13 clones (ii) When these RFs are transfected into Ung+ E. coli strains, the uracilcontaining template strands are preferentially degraded by uracilN-glycosylase, so that most of the phage that survive will be descended from the mutated complementary strands. Site-specific mutagenesis of M13 clones III. Phage DNA replication - phage T4 2. Phage T4 – a complex phage with an icosahedral head and a filamentous tail, and a linear, double-stranded DNA. (1) Terminally redundant DNA – A DNA, usually a phage genome, that has repeats at both ends, that is the sequences at both ends are the same in the direct orientation. (2) Cyclically permuted genome – In a Cyclically permuted genome, there are no unique ends. If a genome of such phage is drawn as a circle, each genome starts from somewhere on the circle and extends around the circle until it return to the same place, so that individual genome have different endpoints, but contain all of the genes. III. Phage DNA replication - phage T4 III. Phage DNA replication - phage T4 III. Phage DNA replication - phage T4 III. Phage DNA replication – phage T4 (3) T4 replication occurs in two stages illustrated in Fig. 7.18: i. In stage 1, replication initiates at specific origins, using RNA primers as usual. ii. In stage 2, recombinational intermediates furnish the primers for initiation, that is the leading strand of replication is primed by recombinational intermediates rather than by RNAs. (i) Repeated rounds of strand invasion and replication lead to very long branched concatemers which can then packaged into phage heads. (ii) Packaging Of DNA into head is initiated by cutting the DNA by a terminase complex which remains attached to the end. This complex then binds to the head at an opening called the portal, and DNA begins to be sucked into the head. (iii) Each packaged genome is a different cyclic permutation with different terminal redundancies. III. Phage DNA replication – phage T4 IV. Lysogeny • Lytic life cycle is not the life style of phages, some phages are able to maintain a stable relationship with host cell in which they neither multiply nor are lost from the cell. Such a phage is called lysogen-forming or temperate phage. • Phage lambda (phage λ) is a kind of lysogen-forming or temperate phage. Temperate Bacteriophages and Lysogeny • lysogeny – nonlytic relationship between a phage and its host – usually involves integration of phage genome into host DNA • prophage – integrated phage genome – lysogens (lysogenic bacteria) • infected bacterial host – temperate phages • phages able to establish lysogeny Life cycle of λ phage λ phage • doublestranded DNA phage • linear genome with cohesive ends – circularizes upon entry into host Genetic map of λ phage Pattern of gene expression Very early events Very early after infection of E. coli, RNA polymerase transcrubes genes N and cro from different strands of the DNA. Early events N protein, an anti-terminator turns on the early genes to the left of N and to the right of cro. The action of N protein 1. Transcription from promoter PL, RNA polymerase will encounter nut ( N protein utilization) site: (1) If no N protein, RNA polymerase will ignore the nut site and fall off the DNA, releasing the mRNA when it reaches the downstream stop signal. (2) In the presence of N protein, RNA polymerase will pass over nut and ignore the downstream stop signal. Late lytic events 1. Protein Cro represses the transcription from PL and PR when it binds to the operators between these two promoters. 2. Protein Q, also an anti-terminator recognizes Qut, lies very near the beginning of the long transcript that initiates at P’R. 3. The Q-modified RNA polymerase transcribes the late genes into a single long transcript. A short segment of the DNA molecule 1. This segment locates between cI and cro genes shown in previous slide. 2. PRM: promoter of maintenance; PR: right promoter; cl: repressor gene; cro; cro gene; O; operator Late events in establishing lysogeny 1. CII protein directs transcription of the two genes needed for finally establising lysogeny 2. PRE: promoter for repression establishment; Pint: promoter for the genes leftward the int gene. The lysogeny decision 1. Host proteases (Hf1A ) regulate the level of CII protein. Although CIII protein is not shown here, it protects CII. 2. Other host proteins could regulate translation of the CII mRNA. 3. Growth medium conditions of infected bacteria influence the activities of bacterial proteases: (1) Rich medium activates the proteases; (2) Starvation has opposite effect. Establishing lysogeny CII-stimulated transcription on int whose promoter lies within the xis gene Establishing lysogeny Retroregulation involved in hairpin structure, RNAaseIII and other RNAase Establishing lysogeny Integration (recombination at att) has separated sib from int. Induction • process by which phage reproduction is initiated • results in switch to lytic cycle • triggered by drop in levels of lambda repressor (even only 5 %) – caused by exposure to UV light and chemicals that cause DNA damage • excisionase – binds integrase – enables integrase to reverse integration process Induction Repressor is cleaved between the amino acids alanine and glycine located in the linker between repressor’s domains. Lysogenic conversion • change in host phenotype induced by lysogeny – e.g., modification of Salmonella lipopolysaccharide structure – e.g., production of diphtheria toxin by Corynebacterium diphtheriae V. Generalized transduction vs specialized transduction • Transduction is the transfer of the bacterial genes by phages. • Bacterial genes are incorporated into a phage capsid because of errors made during the phage life cycle. • The phage containing these genes then injects them into another bacterium, completing the transfer. • There are two different kinds of transductions: generalized and specialized. Generalized transduction Specialized transduction Using phage DNA molecule as a cloning vector Lambda ZAMP® II vector (of STRATAGENE)