* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Young Innovators 2009
5-HT2C receptor agonist wikipedia , lookup
Discovery and development of non-nucleoside reverse-transcriptase inhibitors wikipedia , lookup
Discovery and development of beta-blockers wikipedia , lookup
5-HT3 antagonist wikipedia , lookup
NMDA receptor wikipedia , lookup
Drug interaction wikipedia , lookup
Pharmacogenomics wikipedia , lookup
DNA-encoded chemical library wikipedia , lookup
Prescription costs wikipedia , lookup
Pharmacokinetics wikipedia , lookup
Cannabinoid receptor antagonist wikipedia , lookup
Discovery and development of antiandrogens wikipedia , lookup
Pharmaceutical industry wikipedia , lookup
Pharmacognosy wikipedia , lookup
Discovery and development of angiotensin receptor blockers wikipedia , lookup
Psychopharmacology wikipedia , lookup
Nicotinic agonist wikipedia , lookup
Toxicodynamics wikipedia , lookup
Drug design wikipedia , lookup
NK1 receptor antagonist wikipedia , lookup
Drug discovery wikipedia , lookup
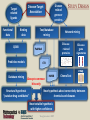
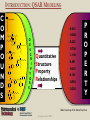
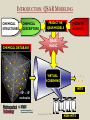
YOUNG INNOVATORS 2009 A novel chemogenomic approach to predict receptor-mediated clinical effects of chemicals: Applications to anti-Alzheimer’s agents Rima Hajjo╫, Bryan L. Roth† and Alexander Tropsha╫ ╫Laboratory for Molecular Modeling, Division of Medicinal Chemistry and Natural Products, Eshelman School of Pharmacy, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina 27599-7360 †National Institute of Mental Health Psychoactive Drug Screening Program and Departments of Pharmacology, Medicinal Chemistry, and Psychiatry, School of Medicine, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina 27599 ABSTRACT Modern paradigms in drug discovery are shifting from traditional receptor-specific studies to multi-receptor profiling to identify complex mechanisms underlying clinical effects of drugs. These approaches relate to chemogenomics, an emerging interdisciplinary field that exploits chemical profiling of receptor panels to identify specific ligands for all targets. We have developed a novel chemogenomic approach combining QSAR modeling, model-based virtual screening (VS), and mining of both biological literature and online databases of gene signatures and biological profiles. Young Innovators 2009 ABSTRACT This approach was applied for the analysis of data pertinent to the Alzheimer’s disease. Gene signatures for Alzheimer’s disease were used to query the Connectivity Map (cmap: http://www.broad.mit.edu/cmap/) database to formulate testable hypotheses about potential treatments. Concurrently, QSAR models have been developed for the ligands of 5HT receptors (i.e. 5-HT6 receptors) known to be involved in cognition enhancement. Externally validated models were used for VS of the World Drug Index (WDI) database to identify putative ligands. Common chemical hits from QSAR/VS studies and the cmap were subjected to radioligand binding assays against 5-HT receptors. Young Innovators 2009 INTRODUCTION: ALZHEIMER’S DISEASE Alzheimer's Disease (AD), is the most common form of dementia. It is incurable and degenerative. As of September 2009, there are reported 35 million-plus cases worldwide. This number is expected to reach 107 million by 2050. AD is characterized by the accumulation of amyloid beta peptide (Abeta), hyperphosphorylation of tau protein, and increased inflammatory activity in the hippocampus and cerebral cortex. The pharmacotherapy of AD consists of symptomatic and disease-modifying therapies. No Treatments to Prevent, Delay or Halt Disease Progression are available today Young Innovators 2009 Artwork by William Utermohlen (died in 2007 of Alzheimer's Disease): Represents the most complete and coherent view of a patient's experience with dementia. ALZHEIMER’S DISEASE AND THE NEED FOR A NOVEL APPROACH TO INNOVATE DRUG DISCOVERY AD has a complex multi-factorial nature and conventional drug discovery approaches may not be suitable for finding novel therapeutics. As a computational scientist, I am interested in developing and exploiting, by the means of computational data analysis, chemogenomic approaches to drug discovery. In conventional QSAR studies we typically explore data related to one receptor; this may be insufficient to understand complex diseases and suggest new therapeutic options. OUR HYPOTHESIS : A Novel Integrated Chemogenomic Approach Could Provide More Comprehensive Answers About Alzheimer’s Disease Etiology and Innovate Drug Discovery Young Innovators 2009 INTRODUCTION: CHEMOGENOMICS Chemogenomics is a new interdisciplinary field, which exploits multi-receptor chemical profiling to identify specific ligands for all targets. The term chemogenomics is applied to a multiplicity of approaches that use chemical compound libraries to probe biological systems. Chemogenomic Space: A complex multidimensional space, where diseases, molecules, genes, proteins, and pathways constitute some of its dimensions. However, few studies attempted to tie together the data on ligand-protein-disease associations in a format allowing new discoveries of both drugs and their targets implicated in the underlying diseases. Most of the available studies, including QSAR studies are limited to pharmacological data (i.e., proteins-molecules subspace)!! Chemogenomic space Young Innovators 2009 Functional data Disease related genes or proteins Disease-Target Association Target related ligands Binding data Text/database mining QSAR Network mining Disease related proteins PubMed Predictive models STUDY DESIGN Disease gene signatures CTD cmap Database mining Accept common hits only Structural hypothesis “putative drug candidates” HMDB ChemoText New hypothesis about connectivity between chemicals and diseases New testable hypothesis with higher confidence Young Innovators 2009 INTRODUCTION: STUDY DESIGN We have developed some elements of the integrated workflow focused on the discovery of new drug candidates by merging predictions from independent lines of evidence. Currently, The overall workflow combines QSAR modeling and model-based virtual screening (VS), gene expression profiling, and text mining of biomedical literature for drug discovery. Predictive QSAR/VS workflow: Rigorous models correlating chemical features of compounds and their biological effects are developed and employed for virtual screening of commercially available chemical compounds to identify hits; CHEMOTEXT, an in-house repository of chemical entities and terms that annotate compounds in terms of their biological activities and, if available, diseases they could treat; cmap (www.broadinstitute.org/cmap/) developed at the Broad Institute, Cambridge, MA, links the effects of different drugs and diseases using gene expression signatures. Only common hits from QSAR/VS and cmap/ChemoText/etc, are considered as high confidence hits to be confirmed in further experimental studies. Young Innovators 2009 INTRODUCTION: QSAR MODELING C O M P O U N D S O N 0.613 O 0.380 N O N O N O N O N O N O O N N O D E S C R I P T O R S -0.222 0.708 Quantitative Structure Property Relationships 1.146 0.491 0.301 0.141 0.956 0.256 N P R O P E R T Y Slide: Courtesy of Dr. Denis Fourches Young Innovators 2009 INTRODUCTION: QSAR MODELING C O M P O U N D S O 0.613 N 0.380 -0.222 O N O N O N O N O N O N O N O D E S C R I P T O R S 0.708 1.146 Quantitative Structure Property Relationships 0.491 0.301 0.141 0.956 0.256 0.799 N 1.195 P R O P E R T Y Slide: Courtesy of Dr. Denis Fourches Young Innovators 2009 INTRODUCTION: QSAR MODELING CHEMICAL STRUCTURES CHEMICAL DESCRIPTORS CHEMICAL DATABASE PREDICTIVE QSAR MODELS PRORETY/ ACTIVITY QSAR MAGIC VIRTUAL SCREENING ~106 HITS 109 – molecules Young Innovators 2009 NON HITS INTRODUCTION: CONNECTIVITY MAP Genome-wide mRNA Expression data for 1,309 bioactive small molecules in different cell lines (MCF7, PC3, HL60, SKMEL5) 7,056 genome-wide expression profiles representing 6,100 individual treatment instances were produced The Connectivity Map is the most comprehensive effort yet for using genomics in a drug-discovery framework. Young Innovators 2009 INTRODUCTION: QUERYING THE CONNECTIVITY MAP Input Database Output High positive score Biological state 1 Null Signature Control Step1: upload signature High negative score Step2: query the cmap Step3 : list of correlated compounds Lamb, J. et al. Science, 313, 1929-1935 (2006) Young Innovators 2009 INTRODUCTION: 5-HT6 RECEPTOR AS A NOVEL TARGET FOR ALZHEIMER’S Young Innovators 2009 METHODS: 5-HT6 DATASET Modeling set: 79 B (Ki < 10,000.00 nM), 99 NB (Ki ≥ 10000.00 nM) Source: PDSP Ki Database B: Binder NB: Non-binder 5-HT6 Dataset Structure Activity Data 99 NB 79 B External validation sets: 24 compounds, all are binders to 5-HT6 (Ki < 10,000.00 nM) Source: Roth Lab, Department of Pharmacology, UNC-Chapel Hill 15 Young Innovators 2009 METHODS: COMBI QSAR STUDY DESIGN Dragon 5-HT6 Dataset: 176 cps 79B & 97 NB B: Binder NB: Non-binder SG Random splitting of the dataset using 5-fold external validation Compound representation kNN Combi QSAR CBA Accept models with CCRtr & CCRts ≥ 0.70 Split into multiple tr & ts sets Modeling set External set (24 B) Validation set Validate by Predicting Validate by Young Innovators 2009 Y-Randomization 16 METHODS: QUERYING THE CMAP WITH ALZHEIMER’S DISEASE GENE SIGNATURES Step1: upload signature Step2: query the cmap Step3 : list of correlated compounds cmap SCORE Alzheimer’s disease gene signatures “Two different signatures” from hippocampus (S1) and cerebral cortex (S2) from two independent reports Positive Connectivity “possible causes for disease state” 1.00 F 0.00 E Negative Connectivity “possible treatments for disease state” (S1) (S2) cmap Signature database “Pattern Matching” Young Innovators 2009 0.00 C B A -1.00 Identification of possible treatments (A,B,C) and causes (F) RESULTS: QSAR MODELING The performance of different binary QSAR approaches employed as part of the combinatorial QSAR strategy for 5-HT6 receptor ligands and based on external set statistics, is summarized in the figure below. Both QSAR methods: kNNDragon and CBA-SG performed very well with the external prediction accuracy characterized by CCRex of 92% and 78%, respectively. Models built with randomized activities for the training set had average CCRex close to 0.50, indicating that the models built with actual activities were robust . Fig. Comparison of the QSAR approaches to classify 5-HT6 receptor binders vs. non-binders based on CCRex. Young Innovators 2009 RESULTS: TOP CMAP NEGATIVE CONNECTIONS WITH S1 FOR AD Web browser screen shot showing top negative connections from the cmap with S1 Young Innovators 2009 RESULTS: TOP CMAP NEGATIVE CONNECTIONS WITH S2 FOR AD Web browser screen shot from showing top negative connections from the cmap with S2 Young Innovators 2009 RESULTS: VIRTUAL SCREENING Number of compounds Removing duplicates & modeling set Generate Dragon descr. 59000 WDI Database 46754 37127 Similarity filter Use acceptable kNN-Dragon models to classify binders vs. non-binders Use CBA-SG classifier Select compounds that have -ve connectivity with AD in cmap 1500 600 Binders 300 Binders 34 Potential anti-AD agents Young Innovators 2009 RESULTS: SELECTED COMMON VS HITS FROM QSAR AND CMAP WDI Name CLOZAPINE TAMOXIFEN FLUSPIRILENE ZUCLOPENTHIXOL BI-2 CIDOXEPIN NORTRIPTYLINE BI-3 ENCLOMIFENE DO-897 LY-294002 ACEFYLLINE-PRENYLAMINE NISOXETINE IFENPRODIL FENDILINE NAFTIFINE RALOXIFENE MEBEVERINE LOBELANIDINE LOBELINE AZACYCLONOL cmap Name clozapine tamoxifen fluspirilene zuclopenthixol imipramine doxepin nortriptyline clomipramine clomifene Prestwick-559 LY-294002 prenylamine nisoxetine ifenprodil fendiline naftifine raloxifene mebeverine lobelanidine lobeline azacyclonol No. of Models Av. Pred. Value cmap Score S1 900 1.00 -0.398 910 0.99 0.358 854 0.99 -0.493 883 0.98 -0.609 898 0.98 -0.503 908 0.97 -0.463 883 0.96 -0.555 893 Selective0.95 Estrogen-0.768 899 0.91 -0.414 858 0.79 -0.741 Receptor 866 0.70 -0.351 Modulators 679 0.69 -0.589 899 0.68 -0.491 900 0.66 -0.541 765 0.66 -0.388 724 0.66 -0.790 809 0.64 -0.378 820 0.57 -0.543 826 0.56 -0.508 882 0.55 -0.514 840 0.54 -0.448 Young Innovators 2009 cmap Score S2 -0.366 -0.507 -0.551 -0.746 -0.415 -0.777 -0.410 -0.425 -0.611 -0.619 -0.303 -0.457 -0.408 -0.489 -0.683 -0.591 -0.482 -0.798 -0.488 -0.750 -0.556 RESULTS: RADIOLIGAND BINDING ASSAYS PDSP ID cmap score S1 cmap score S2 % Inhibition Ki (nM) Clomipramie 13494 -0.768 -0.425 98.6 112.00 Doxepin 13495 -0.463 -0.777 98.1 105.00 Fluspirilene 13512 -0.493 -0.551 97.7 2,230.00 Lobeline 13501 -0.514 -0.750 16.6 NB LY-294002 13502 -0.351 -0.303 -2.3 NB Nortriptyline 13503 -0.555 -0.410 99.1 214.00 Prestwick-559 13498 -0.741 -0.619 27.9 NB Raloxifene 13505 -0.378 -0.482 88.2 750.00 Tamoxifen 13506 0.358 -0.507 91.1 1,041.00 Zuclopenthixol 13510 -0.609 -0.746 98.4 169 Compound Young Innovators 2009 DISCUSSION: COMMON HITS FROM QSAR AND THE CMAP We have queried the cmap to identify drugs that might be connected to AD. Hits with statistically significant negative connectivity scores can be considered as potential treatments. Although the two gene signatures used to query the cmap shared no common genes, both queries resulted in common list of negative connections. Hits from the cmap are considered potential therapeutics for treating the AD; they could have different mechanisms of action including binding to 5-HT6 receptors. Taking common hits from the QSAR study and the cmap increases the chances of identifying novel 5-HT6 ligands as effective anti-AD treatments. This approach also affords the increase in the true hit rates in the QSAR study. Young Innovators 2009 DISCUSSION: SERMS IDENTIFIED TO BE 5-HT6 RECEPTOR BINDERS Binding assays confirmed that 7 out of a total of 10 predicted binders were true binders for the 5-HT6 receptors. Achieving a success rate of 70% , indicates that our QSAR/VS approach was successful in prioritizing computational hits for biological screening. Surprisingly, four of the computational hits predicted to bind to the 5-HT6 receptor were found to be known selective estrogen receptor modulators (SERMs). These hits included tamoxifen, toremifene, clomiphene and raloxifene. However, further analysis of chemical-protein interaction networks indicated that raloxifene has distinctive protein partners than the rest of the SERMs. Both tamoxifen and raloxifene were proved to be true binders in respective binding assays suggesting that these SERMs may be potentially re-profiled as potential anti-AD drugs. Tests will be done soon for toremifene and clomiphene. Young Innovators 2009 DISCUSSION: RALOXIFENE IS A 5-HT6 BINDER AND POTENTIAL ANTI-ALZHEIMER’S A power of the integrated informatics approach to drug discovery Raloxifene was one of the common hits of QSAR and the cmap. Experimental testing confirmed that raloxifene binds to 5-HT6 receptor with a Ki= 750 nM. A recent study indicated that raloxifene given at a dose of 120 mg/day led to reduced risk of cognitive impairment in postmenopausal women. A newlyfunded clinical trial is ongoing to test whether raloxifene can slow the rate of AD progression. This example can be considered as a proof of concept for our novel chemogenomic approach. Therefore, the rest of the 34 hits should be studied comprehensively in relation to AD. Fig. Raloxifene (blue triangle) and Chlorpromazine (square) versus [3H] LSD competition binding at 5-HT6 receptors. Young Innovators 2009 DISCUSSION: ADDITIONAL STUDIES FOR DEVELOPING FAMILY-BASED RECEPTOR MODELS In our effort to understand the mechanisms of actions of drugs predicted from the cmap as treatments for AD, a new study was conducted where gene signatures for the Alzheimer’s disease (AD) were used to query the cmap to identify potential anti-AD agents. Concurrently, QSAR models were developed for the serotonin, dopamine, muscarinic and sigma receptor families implicated in the AD. The models were used for VS of the World Drug Index database to identify putative ligands. 12 common hits from QSAR/VS and cmap studies were subjected to parallel binding assays against a panel of GPCRs. All compounds were found to bind to at least one receptor with binding affinities between 1.7 9000 nM. Young Innovators 2009 CONCLUSIONS We have developed a novel integrative chemogenomic workflow focused on the discovery of new drug candidates by merging predictions from independent lines of evidence. The current workflow combines QSAR modeling and modelbased virtual screening (VS), gene expression profiling, and text mining of biomedical literature for drug discovery. This approach was shown to be useful in predicting new experimentally confirmed ligands for 5-HT6 receptors. Young Innovators 2009 CONCLUSIONS Our approach led to the identification of 34 computational hits as potential treatments for AD. 10 hits with higher confidence level were tested in binding assays and 7 compounds were confirmed as 5-HT6 receptor ligands achieving a success rate of 70%. Mining of the biological literature indicated possible anti-AD effects (for raloxifene, fluspirilene, nortriptyline and doxepin) or neuroprotective effects (for clomipramine) for most of the predicted hits. This approach could be extended to many similar receptor systems serving as a cost-effective in silico tool for the discovery of novel biologically active compounds acting via clinically relevant targets. Young Innovators 2009 ACKNOWLEDGMENTS ADVISOR MML MEMBERS Prof. Alexander TROPSHA Research Professors Graduate Students Alexander GOLBRAIKH Clark JEFFRIES M. KARTHIKEYAN Simon WANG Hao ZHU System Administrator Chris GRULKE Nancy BAKER Kun WANG Tanarat KIETSAKORN Tong-Ying WU Jui-Hua HSIEH Hao TANG Liying ZHANG Stephen BUSH Man LUO Guiyu Zhao Andrew FANT Dongqiuye PU Tiago MODA Joyce CHANDARAJOT Mihir SHAH Administrative Assistant Adjunct Members: Paula PRESS Postdoctoral Fellows Prof. Bryan ROTH Denis FOURCHES Denise TEOTIKO Eugene MURATOV Georgiy Abramochkin CONSULTATION Visiting Research Scientist Dr. Justin LAMB BROAD INSTITUTE Alex SEDYKH COLLABORATORS FUNDING NIH EPA University of Jordan Research Programmer Theo WALKER Weifan ZHENG, Shubin LIU Young Innovators 2009 REFERENCES Kubinyi, H. Chemogenomics in drug discovery. Ernst. Schering. Res. Found. Workshop 2006, 1-19. Roth, B. L.; Sheffler, D. J.; Kroeze, W. K. Magic shotguns versus magic bullets: selectively non-selective drugs for mood disorders and schizophrenia. Nat. Rev. Drug Discov. 2004, 3, 353-359. Keiser, M. J.; Roth, B. L.; Armbruster, B. N.; Ernsberger, P.; Irwin, J. J.; Shoichet, B. K. Relating protein pharmacology by ligand chemistry. Nature Biotechnology 2007, 25, 197-206. Hood, L.; Perlmutter, R. M. The impact of systems approaches on biological problems in drug discovery. Nature Biotechnology 2004, 22, 1215-1217. Hopkins, A. L. Network pharmacology. Nature Biotechnology 2007, 25, 1110-1111. Cavalli, A.; Bolognesi, M. L.; Minarini, A.; Rosini, M.; Tumiatti, V.; Recanatini, M.; Melchiorre, C. Multi-target-directed ligands to combat neurodegenerative diseases. J. Med. Chem. 2008, 51, 347-372. Rosenbaum, D. M.; Rasmussen, S. G. F.; Kobilka, B. K. The structure and function of G-protein-coupled receptors. Nature 2009, 459, 356-363. Young Innovators 2009 REFERENCES Tropsha, A. Predictive QSAR (Quantitative Structure Activity Relationships) Modeling. In Comprehensive Medicinal Chemistry II; Martin, Y. C. Ed.; Elsevier: 2006; pp 113-126. Tropsha, A.; Golbraikh, A. Predictive QSAR Modeling workflow, model applicability domains, and virtual screening. Current Pharmaceutical Design 2007, 13, 3494-3504. Olsson, T.; Oprea, T. I. Cheminformatics: a tool for decision-makers in drug discovery. Curr. Opin. Drug Discov. Devel. 2001, 4, 308-313. Baker, N. Extracting Drug Activity Terms from Medline Annotations. Proceedings: Summit on Translational Bioinformatics; March, 2008; : American Medical Informatics Association; 2008. Summit on Translational Bioinformatics. 2008. American Medical Informatics Association. Ref Type: Conference Proceeding. Jacoby, E. A novel chemogenomics knowledge-based ligand design strategy Application to G protein-coupled receptors. Quantitative Structure-Activity Relationships 2001, 20, 115-123. Young Innovators 2009 RIMA HAJJO: BIOS/CONTACT INFO CONTACT E-mail: [email protected] http://www.unc.edu/~hajjo/ http://www.hajjo.info/ SCHOOL ADDRESS Laboratory for Molecular Modeling, 2069 Genetic Medicine, UNC Eshelman School of Pharmacy ,University of North Carolina at Chapel Hill ,Chapel Hill, NC 27599, USA RESEARCH INTERESTS Rima is interested in cheminformatics, chemogenomics, datamining, polypharmacology, GPCRs and CNS-Diseases. She is working on developing and exploiting, by the means of computational data analysis, chemogenomics approaches to drug discovery. Her PhD thesis project is focusing on polypharmacology, network pharmacology and computational screening of the GPCR receptorome. Young Innovators 2009 Rima Hajjo RIMA HAJJO/BIOS EDUCATION 2005-present, Graduate Program in Pharmaceutical Sciences, University of North Carolina at Chapel Hill, Chapel Hill, NC 1999-2003 M.Sc., Pharmaceutical Sciences, University of Jordan, Amman, Jordan 1994-1999 B.Sc., PharmD, University of Jordan, Amman, Jordan HONORS AND AWARDS 2009 Awarded AAPS Young Innovator Award. 2008 Awarded ADDF Young Investigator Scholarship. 2005 Nominated for king Abdullah’s award for best thesis writing. 2004 Awarded a scholarship from the University of Jordan to proceed with PhD studies. 1994 Awarded a scholarship from the Jordanian Ministry of Higher Education to get a PharmD. 1993 Awarded an academic prize for the scientific creativity of school students, from the Ministry of Education in Jordan. Young Innovators 2009 RIMA HAJJO/PUBLICATIONS RECENT PUBLICATIONS Hajjo, Rima; Grulke, Christopher; Golbraikh, Alexander; Huang, Xi-Ping; Roth, Bryan L.; and Alexander Tropsha. Predictive QSAR models of 5-HT2B receptor ligands identify potentially valvulopathic compounds. (Submitted to J. Med. Chem.) Hajjo, Rima; Roth, Bryan L.; and Tropsha Alexander. In silico strategies to identify 5-HT6 receptor ligands as potential anti-Alzheimer’s treatments. (In preparation) Hajjo, Rima; Setola, Vincent; Roth, Bryan L.; and Tropsha Alexander. A novel chemogenomics approach to identify receptor mediated clinical effects of chemicals. (In preparation) Fourches, Denis; Hajjo, Rima; Wang, Xiang; and Tropsha, Alexander. Multi-task versus single task modeling studies on dopamine receptor ligands. (In preparation) Hajjo, Rima; Roth, Bryan L.; and Tropsha, Alexander. Modeling promiscuity of GPCR ligands. (In preparation) ABSTRACTS/POSTERS Hajjo, Rima; Roth, Bryan L.; and Tropsha Alexander. A novel chemogenomics approach to identify receptor mediated clinical effects of chemicals. Accepted for the 2009 AAPS Annual Meeting and Exposition, November, 2009. Hajjo, Rima; Fourches, Denis; Roth, Bryan L.; Tropsha, Alexander. In silico strategies to identify novel 5-HT6 receptor ligands as potential anti-Alzheimer's and anti-obesity treatments. Abstracts of Papers, 238th ACS National Meeting, Washington, DC, United States, August 16-20, 2009. Wang, Xiang S.; Hajjo, Rima; Tropsha, Alexander. A computational workflow to identify and validate the druggable allosteric binding sites. Abstracts of Papers, 238th ACS National Meeting, Washington, DC, United States, August 16-20, 2009. Hajjo, Rima; Grulke, Christopher; Golbraikh, Alexander; Roth, Bryan L.; Tropsha, Alexander. QSAR models of 5HT2B receptor ligands and their application to predicting compounds that could cause valvulopathy. Abstracts of Papers, 237th ACS National Meeting, Salt Lake City, UT, United States, March 22-26, 2009 Young Innovators 2009