* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Emergence and Applications of RNA Interference
Gene desert wikipedia , lookup
Messenger RNA wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Molecular evolution wikipedia , lookup
Polyadenylation wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Genome evolution wikipedia , lookup
X-inactivation wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Gene expression profiling wikipedia , lookup
Transcriptional regulation wikipedia , lookup
List of types of proteins wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Gene regulatory network wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Gene expression wikipedia , lookup
Non-coding RNA wikipedia , lookup
Epitranscriptome wikipedia , lookup
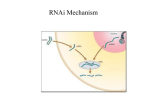
Emergence and Applications of RNA Interference Omar Memon, Vandana Sekhar, Varnika Roy, Yizhou Yin, Alison Heffer University of Maryland, College Park APPLICATIONS HISTORY More than a decade ago, a surprising observation was made in petunias. While trying to deepen the purple color of these flowers, Rich Jorgensen and colleagues introduced a pigment-producing gene under the control of a powerful promoter. Instead of the expected deep purple color, many of the flowers appeared variegated or even white. This phenomenon was considered to be posttranscriptional gene silencing (PTGS), since the expression of both the introduced gene and the homologous endogenous gene was suppressed. Years later, experiments in Caenorhabditis elegans by Andrew Fire and Craig Mello revealed that injection of either “sense” or “anti-sense” mRNA molecules encoding muscle protein, led to no behavioral changes in the worms. But when they injected sense and antisense RNA together, they observed that the worms displayed peculiar, twitching movements. Similar movements were seen in worms that completely lacked a functioning gene for the muscle protein. CRITIQUES Functional Genomics RNAi Advantages Different from classical forward genetics, RNAi is a very powerful technique to investigate gene function in the reverse genetics way. Because of its convenience, high efficiency and economy, it is ideal for analyzing the functions of large numbers of genes and whole genome-wide screens. Based on the completion of sequencing of several organisms and the development of techniques such as cell microarrays, highthroughput RNAi screen is an invaluable tool for functional genomics in a wide range of different species. Use of RNAi in genome-wide screening Step 1 Choose organisms or cell lines Step 2 Choose RNAi reagents: Long dsRNA, synthetic siRNA, plasmid or viruses based shRNA Step 3 Screening with some specific paradigm and format Finding potential therapy targets Easy study of gene function Elucidating cellular mechanisms Analyze many genes at once; RNAi libraries Knock-downs can show a variety of phenotypes More specific than most other therapies Can be used to target many cell and tissue types Efficiently transfers the gene to its target Functional genomics Therapeutics N.benthamiana C.elegans D.melagonaster Arabidopsis Mouse Human Step 4 Read out and analyze results, microarray can be imaged or stained Large-scale RNAi screens have been done: Fire and Mello tested the hypothesis that injection of sense and antisense RNA molecules resulted in the formation of double-stranded RNA (dsRNA). In every experiment, injection of double-stranded RNA carrying a genetic code led to silencing of the gene containing that particular code. From this, they deduced that dsRNA can silence genes and that this RNA interference is specific for the gene whose code matches that of the injected RNA molecule, and that RNA interference can spread between cells and even be inherited. •About 90% genes on C.elegans chromosome III for several basic cellular processes, •Screen on C.elegans chromosome I for embryonic lethal genes, Fire and Mello published their findings in the journal Nature on February 19, 1998. Their discovery clarified many confusing and contradictory experimental observations and revealed a natural mechanism for controlling the flow of genetic information. This research awarded them a Nobel prize and heralded the start of a new research field. •Functional screen for RNAi itself in C.elegans Meanwhile, high-throughput screens and RNAi libraries have proved to be very useful to therapeutic research MECHANISM 1. Introduction of ds RNA in the cell by viral infection or by artificial means using vectors based short hairpin RNA (shRNA) ds RNA virus 1 shRNA 2 3. Duplexes of siRNA of 21-24 nucleotides length formed by Dicer DICER 3 4 miRNA ATP ADP + Pi 5 RISC ds DNA 6 ATP ADP + Pi RISC activation 7 Target mRNA 8 Degraded target mRNA 2. Recognition and processing of long dsRNA by Dicer, an RNase III enzyme 4. miRNA are naturally synthesized long ds RNA in the nucleus, which are processed by Drosha enzyme into small pre-miRNA and exported to cytoplasm. Disease Therapy Wide therapeutic applications of siRNA are the new sensation in the biotechnology drug world. Major traditional drug targets have been proteins (enzymes and receptors), which are targeted at the post translational level. But siRNA drug selectively silences a disease causing gene, at the post transcriptional level itself. Side-effects are decreased by targeting a disease inducing gene in which genetic polymorphisms distinguish it from the RNA of wild type alleles. Unlike the antisense approach, dsRNA employs a normal cellular process thus it is more specific and allows a cell-cell spreading of the gene silencing effect. The knockdown of the target gene by RNAi is heritable and stable. Taking RNAi from Bench to Bedside- First Trial Treating Age Related Macular Degeneration DELIVERY OF DRUG Local intravitreal injection of siRNA (100-800µg) per eye diluted in phosphate buffer saline) 7. Recruitment of RISC along with antisense strand to target mRNA 8. Cleavage of target mRNA by an unidentified RNase (Slicer) within RISC. Degrades mRNA at sites not bound by siRNA siRNA duplex RISC activation VEGF target recognized PHENOTYPIC EFFECT The biggest problem with the use of RNAi is its successful delivery to the target. RNAi must be stable in a cell for prolonged activity without getting degraded. Nonspecific interactions can occur because, though siRNA can be designed to target a specific sequence, a difference in one or two base-pairs is sufficient to cause off-target binding. Table 2. Different delivery methods of RNAi and the advantages and disadvantages of each Knock-downs do not completely inhibit gene expression or activity Number of genes targeted at one time limited; RNAi overload RNAi doesn’t always work Difficult to deliver siRNA to the target Side effect of activating the interferon response (IR) siRNA stability is a concern for effective therapy Concern that the siRNA might interfere with natural RNAi mechanisms in the cell May bind non-specifically to some tissues Delivery system The research conducted over past few years has shown the promising potential of RNAi. This powerful genetic tool has been used to a certain level of success in both proteomics and drug therapy. Though both will continue to be active fields of scientific research, the drawbacks must also be considered (Table 1). Table 1. Advantages and disadvantages of using RNAi in two applications Major characteristics Promising results in cancer therapy through in vitro cell studies Related to retroviruses; eliminate some disadvantages Effective in targeting genes in the brains of Alzheimer’s patients Retrovirus Viral Lentivirus Vector based on adenoassociated viruses Can target tumors For transient expression Balances stability of siRNA without influencing RNAi mechanisms Modifications: locked nucleic acids (LNA), phosphothioates (PS), 2’ modifications to ribose siRNA packaged into an envelope with a signal for target cells Adenovirus Chemically modified siRNA Nonviral Liposomes Naked siRNA Advantages siRNA is injected directly into the organism Very efficient Genes are passed on during mitosis Applies to non-dividing cells Virus can be specific for recognizing one type of tissue Good for carrying larger genes Small risk of host genome integration since replication occurs outside of the nucleus Stability of siRNA is increased More specific targeting Easy to obtain Targets many different types of organs Accomplished with relatively little work Disadvantages Gene is integrated into the genome; risk of mutagenesis Only for dividing cells Can only target specific areas; no systemic applications Risk of mutagenesis Since the DNA is outside of the nucleus, it is less stable and can be lost after many cell divisions No in vivo studies on many modified RNAs 5’ modifications might interfere with silencing Bulky modifications may hinder RNA unwinding Non-specific targeting Liposome electric charge on may interfere with tissue uptake Encounters RNAses in serum Delivery to non-specific sites SUMMARY RNAi is a powerful and attractive genetic approach because of the diversity of its applications. The potential uses currently in progress include the identification of specific gene functions in living systems and creation of genome wide screens. Development of antiviral and anticancer therapies are broadening the horizons of the therapeutic arena. Another value of RNAi screens is in combining it with other functional genomic assays enabling mapping of biochemical pathways. Impact of RNAi is also being extended to the field of agriculture for example by increasing disease resistance in plants. Many potential obstacles in the path of RNAi therapeutics can be overcome, but further insight into the non-coding functions of RNA in vivo will provide better understanding of mechanisms underlying RNAi. Future applications of RNAi technology will revolutionize genetic, genomic and proteomic aspects of biology and will take the field of medicine into new scientific realms. Target cleaved SIRNA-027 Target = VEGFR-1 REFERENCES 5. Incorporation of both synthetic siRNA or endogenously expressed miRNA into RNA-induced silencing complex(RISC) 6. Unwinding of duplex siRNA by a helicase in RISC and removal of passenger strand (RISC activation) IN VIVO MECHANISM RNAi Disadvantages Ocular angiogenesis Reduced Dose dependent Improvement of Vision Common RNAi Targeted Diseases ONCOGENESIS •siRNA drugs directly target cancer promoting genes •Chemotherapeutic avoidance of tumors is decreased by targeting clusterin (antiapoptotic gene). •Ex:-Imatinib drug for Philadelphia chromosome target BCR-ABL fusion protein causes chronic myelogenous leukemia NEURODEGENERATIVE DISEASES •RNAi is an important process in normal neuronal function •Its manipulation is important for treating many untreatable neurological disorders • Ex:-Mouse models for Alzheimer's disease, DYT1 dystonia, and polyglutamine disease in progress VIRAL DISEASES •Targets are viral and host genes that are essential for entry of the virus •Hepatitis B and C, Influenza and HIV are common targets •Ex:- Silencing of the HIV chemokine receptor (CCR5) by RNAi therapy is under trial by Benetic and City of Hope company Bantounas I, Phylactou L, and Uney J. 2004. RNA interference and the use of small interfering RNA to study gene function in mammalian systems. J. Mol. Endo. 33: 545-57. Beal J, 2005, Silence is golden: can RNA interferance therapeutics deliver?, Drug Discovery Today, 10 (3), 169-172 Bargmann C I, 2001. High-throughput reverse genetics: RNAi screens in Caenorhabditis elegans Genome Biology, 2(2): 1005.1-1005.3 Caenorhabditis elegans experimental illustration: Annika Rohl Echeverri C J, Perrimon N, 2006.High-throughput RNAi screening in cultured cells: a user’s guide , Nature Reviews Genetics, (7), 373-384 Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, and Mello CC. (1998). Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391: 806-811. Jorgensen RA, Cluster PD, English J, Que Q, and Napoli CA. (1996) Chalcone synthase cosuppression phenotypes in petunia flowers: comparison of sense vs. antisense constructs and single-copy vs. complex T-DNA sequences. Plant Mol Biol 31: 957-973. Leung R K M, Whittaker P A, 2005. RNA interference: from gene silencing to gene specific therapeutics, Pharmacology and Therapeutics, 107: 222239 Li C, Parker A, Menocal E, Xiang S, Borodyansky L, and Fruehauf J. 2006. Delivery of RNA interference. Cell Cycle. 5(18): 2103-9. Lieberman J, Song E, Lee S, and Shankar P. 2003. Interfering with disease: opportunities and roadblocks to harnessing RNA interference. Trends in Mol. Med. 9(9): 397-403. Miller V, Paulson H, and Gonzalez-Alegre P. 2005. RNA interference in neuroscience: progress and challenges. Cellular and Molecular Neurobiology. 25(8): 1195-1207. Shuey D L, Mc Callus D E and Giordano T, 2002. RNAi: gene-silencing in therapeutic intervention, Drug Discovery Today, 7(20): 1040-1046 Sonnichsen B, et al. Full-genome RNAi profiling of early embryogenesis in Caenorhabditis elegans , (2005), Nature Vol 434(24), 460-469 Stevenson M, 2002. Therapeutic Potential of RNA Interference, The New England Journal of Medicine, 351(17), 1772-1777 Tuschl T, 2003.Functional genomics RNA sets the standard, Nature Vol.421 16 January, 220-221 Wheeler D B, Carpenter A E, Sabatini D M, 2005. Cell microarrays and RNA interference chip away at gene function, Nature Genetics Supplement, (37) 25-30 Whelan Jo, 2005. First Clinical data on RNAi, Drug Discovery Today, 10(15), 1014-1015 Poster: RNA Silencing, (2005) Science 309, 1518