* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Doug Juvinall December 8, 2009 Bradley University Bio 464 Lab
Oncogenomics wikipedia , lookup
RNA silencing wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Genome evolution wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Point mutation wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene expression programming wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Microevolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Designer baby wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Gene expression profiling wikipedia , lookup
Primary transcript wikipedia , lookup
Genome editing wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
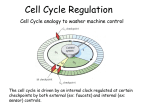
Doug Juvinall December 8, 2009 Bradley University Bio 464 Lab Abstract: Cyclins play an important role in the life of a cell. An RTPCR gel can be used to determine their activity at different time points in the cell cycle. The activity of the cyclin TTHERM 00192000 was measured during conjugation of the ciliate Tetrahymena. TTHERM 00192000 was named CYC5. RNA was collected from Tetrahymena at different time points of conjugation. Primers were made for the TTHERM 00192000 gene which was then used for the RTPCR. The intensities of the gel were determined which corresponds to the amount of gene activation at specific time points. This shows the times during conjugation where the cyclin is supposed to be active. The times of expression from this gel did not correspond to that of the Tetrahymena Gene Expression Database. This suggests there was error in either or both of the data. Introduction: Cyclins are proteins that are expressed at different times of the cell cycle. They allow certain events to occur which permit the cell to continue the process of cell division or conjugation as in the case of Tetrahymena. Cyclins work by binding to a cyclin dependent kinases (CDKs) which activates it. Then the cyclin-CDK complex can phosphorylate target proteins around the cell to allow the cell to continue its cell cycle/conjugation (Lodish et al, 2008). When ciliates are stressed, they will undergo sexual reproduction to try and produce individuals that are able to adapt to the stress. The conjugation process can be over 14 hours long and it has many steps that involve multiple nuclear divisions (Miao , 2009) Figure 1. Events in the Tetrahymena thermophila conjugation cycle. Taken from Miao, et al., PLoS ONE. 2009; 4(2): e4429 Figure 2. Expression data for TTHERM 00192000 in T. thermophila. This is the microarray expression profiles the gene from TGED. L = vegetative log phase growth; S = hours under starved conditions; C = conjugation (hours post-mixing). The TGED data suggests that the cyclin TTHERM 00192000 appears to have peak expression at around 5 to 5.5 hours into conjugation. TTHERM 00192000 was named CYC5. 5 hours into conjugation is the selection of one of the four meiotic products. The 5.5 hour time point is haploid mitosis and the 5.75 time point is the exchange of pronuclei. It is likely that the cyclin sets the cell up for either of these processes. Because haploid mitosis is occurring around the peak activity of this cyclin, it is possible that it is involved in initiating any of the mitosis steps of this haploid mitosis. However, the cyclin activity does not drop off immediately but instead is gradually decreases until around the 12 hour time point which includes exconjugation, loss of Old MAC, and 1 MIC genome rearrangement. The conjugation changes around the 5.75 hour time point which is the exchange of the pronuclei. This means the cells create openings to allow the exchange. The cyclin could be involved in the creation of the openings or the persistence of it. When the cell closes the opening after the pronuclei have been exchanged, it may change the nature of the conjugation. Since the cyclin gradually decreases after the 5.5 hour time point all the way until the 12 hour point, it is not unreasonable to assume that the cyclin could also be involved maintaining the binding of the two Tetrahymena cells after the exchange of pronuclei. Because of the gradual decrease in activity, this seems to be the most likely process the cyclin is involved in. >TTHERM_00192000 (Gene) ATGGCTCAAAAAATTTAAGAGTTAGAAGAATAGTTTTAATAATTTGAATAATCAATTGGAATTAACGATTAAAATTTTGCTAATAAATTCATCTTTT AGTAATTGAGGGAGAATTTGATTAAGGCTTCCGCAGAATCGAATGATAAATTGCAGAAATTTTTAGATTAGGCTCATACAAATTTGTACTTGGAAAT TAGTTCAAAGATTTAAGCCGATTTAGATGGATTCACTAGAGAAGAATAAAACATGCTTCATTTTTATGGCGCATCAATAATTTCTGATGCATGCTAA TATCTTTAGCTTCCAATTACAACTTGTATATCAGCCTAAACTATTTTCCACAGATTTTATACCAAGTAATTTATTTTATTTTGAATGCATTAGAAAT ATTTTTTAATTGATTCAAAACATTAAAAATAATAAATAATTTTTTTTTTTTCTATAAAAAGGTGCTCTTTCTTGAAGCATGATATCAGGGATGTAGC AATGGGATCAGTTTTTATAGCTGGTAAGGCTTAAGAAACAATTAAAAAGCCAAGAGATTTAGCCTATGTCTTTGATCAAATTTTTAAAGTAAATTAT CAGTTTATATTAGATTGCTATCAGCAAAGGAGATTCCTTCAAGAATCAAATCAACCCTCCTAAATCAAACTTTTCAATAATTTTTTTAAAAAAACTT TGAAGGAAAAAAATTAGATCTTAGCTTATTTGCTTTATCTTATGAAAATAATTATTTTAATCATTTGAACTAATTAATAATATAATTATAGGGTAAA TTTATTTTTATGATCTATCATCATCTTCATCATCTTGCTAAATTGAAAGTATTAATATATTAAACACCTTCCCTAAACACTTTCAATGAAAATGTGG CTGGTGTTAGTTAATAAGATTGAATTAAAGATTAATTTGTTCATCAAAATACACAAACACTACCAATTATCATTATCACTATCAACCAACAACATCA ACGAAATAATCTATTTTGTTTGATTCCCTAGTTCCTAAAATTATATGAAAACTTCTTACTGAAGTGATAATAACAATAGATGAATTAAAATCCTTAA TCTTATTTTTAAATACAATTATGTTAGGAATTTTTTAATTTCTCTTATTCAAAGCACAGGCCAATAAATGCTATTTTTAAGCAGAAATAGTAAGTTA ATTCAGTTTTAGAGAAGCAAGAAAAGCTATGAAAATAATTGAATAAAAAAAGTCTTCCAAAAACAAAAATAAAATTATTTATTTCCTTCAAATAAAA GATTTTTAATTAATTTTTAAAAAAAGCGCCCTTTTTAAATTTACTGAAAAACAAACTGTCTCTATTCATGATAGCAAATCGATATAATTTAAAAAGT AAATTTCATGCTTTTTATTTTAAAAAATAGATTGAAGATGGTATACAACGTCCAGTTCCAATTTTAGATGATAAATCATTTAAATTTAATCATTTAA AATAAGTGGTTTAAGACATGGAAAGAGAGATTTTAAAAGAATTAGGCTTTGAGTTATATCAAATTACATGGAATGAACAACCTCATAGATTAATGTA CTTCTATATTAATTTATTTAAACCAAATACCAGTAATTAATCGAGTTCCTTTTAGAATTTGACTAGAACAGCATTCAACTATTTGAATGATAGTTAC AAGACAAACATCTGTATTTTCTTCCCTTTCTAGATGATAGTAGCAAGCTGCATTTACCTAGCCTTTAGAAAGACAGGTACTGAAATGCCAAGAATTG CTTGGTGGACAATTATGGAAACTTCTCTGAACAATCTTAAGCTAGGAGCGGGTAAAATCTAATATATTTATAATCAATTTAAATAACTTCCTAAATA TGAGGACTGCTTCGAAATTCTTAATAAACTTGCAAAACAAAAGGAAATAGAGTTGAAAATCACTCTTAAATATCATGAATTTACAGATAGATTAAAC CAAAGTTCTGCATCTTAGAATCCTATGTTAGCTGTCCAAGAAGCTATCGCCAAATCAAAATTAAGGGGATAAAATGGATTGGCTGCAGGAGCATCAT CATCAGACAAGTTGTTATCATCAGATTAGCTTGAAAAAAAAGAAAATTAAGATGAAACTGCAGGAGTTACTAGTTTAAGAGATGTTACCAGTACTAA AAATGATAAGCATAGCAAATTTTCTACGGCTGGTAGATCAGCATCAAGAGAAAGAGATAGACGTAAAAGGTCCAGAAGCAATTCCCACAAAAAATCC CATAAAAGAAGATCAAGTCGTAGTAGAAGTCGTAATCGTAATAGAGATCGTAAGGATAAAGATAAAAAGCATTCTTCTCGTCACAGTCGCAGCAGAG ATCGTCGTGATAGAAGTAATTCCCATAAAGATAAAAAAAAAAGAAGTAAAAGTCGTGAACGTGAACGCGAACGTAGAAAAGAAAAGTAAGAAGTCCA AAAGCCTACAAGTGCTGAGCGTTAATAAAACGATTAATAAAAGTCTTAAGAGGAATAAAAAACCAACGGTAATGGTGCATCATCAAGTATAAAAACT ACTTCTGAAACACCAGATATTAAGTAAGATAAAACTTCACCTTTAAATTCTAAAAAAGATAAGAATTAAAGAGATCGTGAATCTGCATCAACAGTAA GGGATGTAAGAGAAGGTAGGGAAAAGGAAAAGGACAGAAAAGAAAGAAAGAAGAGTAGAAGCCGTAGTAGAAGTCATCATAAAAAATCTAAAAAGTA TTATAACTCTCCTGATGATAGGGATAGTAAGAGAAAGCGCAGTAAAGATAGAAAAAAAGACTATAAAAAATCAAGGAGCAACAGCAGAGAAAAATAG TCTAAATTTTCAGTTCCTACTTCATCTACTCAATAGATGGAATCAATTGCAAAAATTACACCTTCTCCTCCCTTGAAATAAGATAAATTTACTACAG TTCTAAATGAAAAAAGTAATGATTTTTCATCTGTATAAAATAAAATACTTCCTTCTGATAATAAAAATATTCATTAAGAAAATCAAATAAATGGCAA TCCAAGTATTATTGAAACAGCTGCTCCTGCTCCCAACTAAAAATTTATTTAAAAGTAAGATTTGTTTCATAATTTATAAAATAATAGCAATTTAACT CTAATTAATAACATCAAATTACTAATAATAAAAAGAACTGATTAAATTAAGCCTCAAGAATAAAATTCTTTATTAGATGGAAAGAAAACACTTGACA TTATCTAATAAATAAAATCTAAAATTTAATCAAAGGCTACTAATGAGGAAAGTTCGAATTAAAAAATATCAAGTTCTAATGTCACTTAAGAAACTAA CCTAACATAGAAGGCTCAGACAAACCCAACCAAACCAACTTTAGATTTAAGCGCTAAATTGAGTGAAAAGTTTAAATAGTACGATAACAAACTGCCC TCTTCATCATCATCAACAACAACATCAACTACAACATCAACTACAACAGCAACAAAATTAGTAGCAGGAGCCAGTTAGCCATAGTTGTCTAAGTAAT TGTCAAAAGAAAGTATGACTTCAAACGATGATGAAAATAATGACAACGATGACTATCTGATGAGATTGAAAATGCTTCGTAAATAATTTTTACATAT TGTTTAGATAAAATAGTGATAAAATTTAACTGTCTAAAAATTAGGTGAACAAAATAAATAAAAGGCTCAACAAGGAAACTAATGA Figure 3. Genomic sequence of TTHERM_00192000. Red letters correspond to introns. Yellow letters correspond to the primer sequences. Sequence from Tetrahymena Genome Database (www.ciliate.org) Materials and Methods: Cyclin genes were identified at the Tetrahymena Genome Database (www.ciliate.org) by searching for proteins with the keyword “cyclin”. A BLAST search with a cyclin protein sequence ensured that all cyclin genes were identified using this method. Microarray data during conjugation (Miao, et al., PLoS ONE. 2009; 4(2): e4429) were collected for each gene from the Tetrahymena Gene Expression Database (TGED; http://tged.ihb.ac.cn). PCR primers flanking an intron were generated for each gene using Primer3 (Steve Rozen and Helen J. Skaletsky 2000) and ordered from Integrated DNA Technology (Coralville, IA). Oligo-dTprimed M-MLV reverse transcription (RT; Ambion) was performed on RNA collected from control cells and from cells at various stages of conjugation using the Trizol reagent (Invitrogen) according to the manufacturer’s protocol. 1 mL of cells (2.1 x 103 cells/mL) was collected at each time point, pelleted at 6k rpm, supernatant discarded, and cells resuspended in 1 mL of Trizol. 180 ng of each template RNA was used per reverse transcription reaction. cDNA was diluted 1:5 and used as a template for PCR (How much RNA for RT? ) PCR was performed in 25 uL reactions using GOTaq (Fisher, Hampton, NH) with 1 uL of each primer (10 uM). 15 uL of completed PCR reaction products were separated on a 2% agarose gel. DNA bands were visualized using ethidium bromide and photographed with a Kodak EDAS290 imaging system. Band intensities were determined using ImageJ (Abramoff, M.D., Magelhaes, P.J., Ram, S.J. "Image Processing with ImageJ". Biophotonics International, volume 11, issue 7, pp. 36-42, 2004). Results: The RTPCR data for TTHERM 00192000 is vastly different from the TGED. This data appears to have more activity at hours 1, 7, 9, 11, 14, and 16. There does not seem to be anything at these time points that are similar. This means the experiment failed and should be repeated. However, the gene appears to be expressed in vegetative, starved, and conjugating cells based on the appearance of bands in the RTPCR. The primers do appear to be working correctly because bands do show up on the gel. This means that future work could use the same primers. There was a more pronounced experimental error on time points for hours 4, 17, and 18 of conjugation. M G V V S S 0 1 2 3 4 5 6 7 M 8 9 10 11 12 13 14 15 16 17 18 Figure 4. RT-PCR analyses. Lane 1 = DNA MW Marker. Lane 2 = CU428 genomic DNA template (contains intron). Lanes 3,4 = CU427 and CU428 vegetative. Lanes 5,6 = CU427 and CU428 starved 24 hours. Lanes 7-25 = conjugating (0-18 hours post-mixing). RNA concentrations were standardized using Nanodrop prior to RT-PCR. Figure 5. Expression data for TTHERM 00192000 in T. thermophila. This is the from the RTPCR gel. Band Intensities were determined by ImageJ software. Discussion: The results from the gel and the TGED data do not compare well. It seems more likely that the TGED data for TTHERM 00192000 is closer to being true because it shows an obvious increase in expression at a certain time in conjugation while the RTPCR data appears to have expression at many time points of conjugation that have little or no relation in the cellular activities. Similar experiments using RTPCR of Tetrahymena genes predicted the expression charts from TGED. Because those experiments have reinforced TGED results, then it is likely that this RTPCR experiment has failed and needs to be repeated. References: Abramoff, M.D., Magelhaes, P.J., Ram, S.J. "Image Processing with ImageJ". Biophotonics International, volume 11, issue 7, pp. 36-42, 2004 Lodish et al. Molecular Cell Biology. 6th Edition. W.H. Freeman Publishing. 2008 Miao, et al., PLoS ONE. 2009; 4(2): e4429