* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download HIF-2α phosphorylation by CK1δ promotes erythropoietin secretion
Survey
Document related concepts
Endomembrane system wikipedia , lookup
Cell nucleus wikipedia , lookup
Extracellular matrix wikipedia , lookup
Signal transduction wikipedia , lookup
Tissue engineering wikipedia , lookup
Cell growth wikipedia , lookup
Cytokinesis wikipedia , lookup
Cell encapsulation wikipedia , lookup
Cell culture wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Cellular differentiation wikipedia , lookup
Protein phosphorylation wikipedia , lookup
Phosphorylation wikipedia , lookup
Transcript
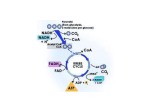
© 2016. Published by The Company of Biologists Ltd. HIF-2α phosphorylation by CK1δ promotes erythropoietin secretion in liver cancer cells under hypoxia Evanthia Pangou1, Christina Befani1, Ilias Mylonis1, Martina Samiotaki2, George Panayotou2, George Simos1and Panagiotis Liakos1* 1 Laboratory of Biochemistry, Faculty of Medicine, University of Thessaly, 41500, Biopolis, Larissa, Greece Protein Chemistry Laboratory, Biomedical Sciences Research Center “Alexander Fleming,” Vari 16672, Greece 2 * Author for correspondence ([email protected],gr) KEY WORDS: HIF-2, CK1, Phosphorylation, EPO, CRM1, Liver cancer cells Summary Statement Journal of Cell Science • Advance article Casein Kinase 1δ-mediated phosphorylation of Hypoxia Inducible Factor-2α increases erythropoietin production by impairing a CRM1-dependent mechanism in liver cancer cells. JCS Advance Online Article. Posted on 29 September 2016 ABSTRACT Hypoxia Inducible Factor 2 (HIF-2) is a transcriptional activator implicated in the cellular response to hypoxia. Regulation of its inducible subunit, HIF-2α, involves post-translational modifications. Here, we demonstrate that casein kinase 1δ (CK1δ) phosphorylates HIF-2α at Ser383 and Thr528 in vitro. Disruption of these phosphorylation sites and silencing or chemical inhibition of CK1δ reduced the expression of HIF-2 target genes and the secretion of erythropoietin (EPO) in two hepatic cancer cell lines, Huh7 and HepG2, without affecting levels of HIF-2α protein expression. Furthermore, when CK1δ-dependent phosphorylation of HIF-2α was inhibited, we observed significant cytoplasmic mislocalization of HIF-2α, which was reversed upon addition of the nuclear protein export inhibitor Leptomycin B. Taken together these data suggest that CK1δ enhances EPO secretion from liver cancer cells under hypoxia by modifying HIF-2α and promoting its nuclear accumulation. This modification represents a novel mechanism of HIF-2 regulation that may allow differentiation of HIF Journal of Cell Science • Advance article isoform function. INTRODUCTION Hypoxia Inducible Factors (HIFs) are transcriptional activators that orchestrate the cellular response to hypoxia. The HIF family contains HIF-1 and HIF-2, both of which function as heterodimers, composed of an oxygen-regulated α subunit and a stably expressed β subunit, also known as ARNT. HIF-1α is ubiquitously expressed and regulates many adaptation responses and metabolic enzymes (Majmundar et al., 2010). The second less studied isoform (HIF-2α) is encoded by the EPAS1 gene, is restricted to specific cell types and is more involved in angiogenesis and metastasis (Keith et al., 2012). HIF-2 is essential in physiological situations such as embryonic development and especially erythropoiesis, a process in which HIF-2 has a predominant role. This has been shown by studies in HIF-2α knock-out mice documenting HIF-2α as the main regulator of hepatic EPO production and essential for the maintenance of systemic EPO and iron homeostasis (Rankin et al., 2007). HIF-2 also plays significant roles in pathological situations, like the progression of solid tumors (Patel and Simon, 2008, Zhao et al., 2015). HIF-2α is regulated by oxygen in a similar way as HIF-1α. Under normoxia, HIF-2α is modified by the Prolyl-4-hydroxylases (PHDs), which then leads to recruitment of the von Hippel-Lindau (VHL) tumor suppressor E3 ligase complex, poly-ubiquitylation and finally proteasomal degradation (Ohh et al., 2000). In addition, under normoxia HIF-2α is also hydroxylated at an asparagine residue by Factor Inhibiting HIF (FIH), which inhibits HIF-2α interaction with the transcriptional co-activator CBP/p300 (Kaelin and Ratcliffe, 2008). During hypoxia, hydroxylation of HIF-2α is inhibited because PHDs and FIH, which use oxygen as substrate, become inactive. This leads to HIF-2α stabilization, transport inside the an active transcriptional complex. Interestingly, HIF isoforms are also subject to oxygen independent and differential regulation, both in terms of expression levels and activity. This regulation includes response to various signaling pathways and functional interplay with other transcription factors which involve a variety of post-translational modifications. While HIF-1α regulation by major signal transduction pathways has been extensively studied (Kietzmann et al., 2016) our knowledge about how HIF-2α is regulated remains quite limited. It has been reported that HIF-2α is acetylated at several lysine residues by CPB (Chen et al., 2012) and de-acetylated by SIRT1 (Dioum et al., 2009, Lim et al., 2010) but there is contradictory evidence concerning the role of these modifications. Moreover, HIF-2α is phosphorylated at Thr324 by Journal of Cell Science • Advance article nucleus, dimerization with ARNT, DNA binding, co-activators recruitment and formation of PKD1 and, as a result, its interaction with the SP1 transcription factor is inhibited, thus promoting NBS1 expression (To et al., 2006). HIF-2α is also phosphorylated by casein kinase 2 (CK2) at Thr844, which results in increased HIF-2 transcriptional activity possibly by lowering the affinity of HIF-2α for FIH (Gradin et al., 2002). The casein kinases (CK) are serine/threonine kinases and comprise the casein kinase 1 (CK1) and casein kinase 2 (CK2) families (Knippschild et al., 2014). In mammals, seven distinct CK1 isoforms have been characterized, all highly conserved and ubiquitously expressed. Up to now, more than 140 in vitro and in vivo substrates of CK1 have been reported, including proteins involved in cell cycle, apoptosis, DNA repair, mitochondrial function and signal transduction. The important role of CK1 family members has been documented by reports linking CK1 isoforms to modulation of key regulatory proteins such as p53, MDM2, and βcatenin, all of which act as signal integration molecules in stress situations and are connected to tumorigenesis (Schittek and Sinnberg, 2014). Previous work from our lab addressed the role of CK1δ in the hypoxic response and has shown that phosphorylation of HIF-1α by CK1δ destabilizes the HIF-1α/ARNT complex and reduces HIF-1 transcriptional activity. Also, by negatively affecting HIF-1 activity, CK1δ decreases lipin-1 expression and lipid droplet formation and, thus, impairs the metabolic adaptation and proliferation of cancer cells under hypoxia (Kalousi et al., 2010, Kourti et al., 2015). In this report, we investigated how casein kinase 1δ (CK1δ), affects HIF-2α expression and transcriptional activity as well as induction of EPO in hepatic cancer cell lines under hypoxia. Our findings suggest a novel regulatory effect of CK1δ on EPO production through direct Journal of Cell Science • Advance article phosphorylation of HIF-2α. RESULTS Casein kinase 1δ supports HIF-2 transcriptional activity and stimulates EPO secretion To investigate the involvement of CK1δ in the regulation of HIF-2α, its expression was suppressed in Huh7 cells by small interfering RNA (siRNA)-mediated silencing. Treatment of cells with CK1δ siRNA under hypoxic conditions substantially reduced the expression of CK1δ, but did not affect the expression levels of endogenous HIF-2α (Fig. 1A). The same treatment drastically reduced the mRNA expression levels of the HIF-2 specific target gene EPO suggesting inhibition of HIF-2 transcriptional activity (Fig. 1B). To explore further the biological function of the CK1δ-mediated regulation of HIF-2α, we measured EPO secretion and we observed that silencing of CK1δ expression substantially reduced EPO secretion compared to cells treated with control siRNA under hypoxia (Fig. 1C). Similar results were obtained in both Huh7 and HepG2 cells using an alternative CK1δ siRNA sequence (Fig. S1A, S1B, S1C, S1D, S1E). To verify these results, Huh7 and HepG2 cells were treated with the specific CK1δ inhibitor D4476 (10 μM) (Rena et al., 2004) under both normoxia and hypoxia. CK1δ protein was similarly expressed under all conditions and inhibition of its activity by D4476 did not significantly alter the protein expression levels of endogenous HIF2α induced by hypoxia (Fig. 1D, S1F). In line with the results obtained by silencing of CK1δ, treatment of both cell lines with D4476, declined significantly the mRNA levels of the HIF-2 specific target genes EPO and PAI-1 under hypoxic conditions (Fig. 1E, S1G) The requirement of CK1δ for hypoxic stimulation of EPO secretion was verified by treating cells with D4476 under hypoxia (Fig. 1F, S1H). Taken together, these data show that inhibition of CK1δ significantly decreases EPO and PAI-1 gene expression suggesting that CK1δ positively affects HIF-2 activity and HIF-2-dependent EPO production under hypoxia CK1δ phosphorylates HIF-2α in vitro To analyze the observed HIF-2 regulation by CK1δ we examined if CK1δ directly modifies HIF-2α in vitro. Full length HIF-2α and its smaller fragments (amino acids 1-820, 1-679, 1542, 1-497, 1-366, 1-280, 366-870, 366-820, 366-679, 366-542, 542-870 and 542-820), were expressed in bacteria as GST fusion proteins (Fig. 2A), purified and used as substrates in radioactive kinase assays using recombinant CK1δ (Fig. 2B). Very little phosphorylation was detected in the amino-terminal (1-280, 1-366) and carboxyl-terminal (542-820, 542-870) fragments of HIF-2α compared to full length HIF-2α. All other fragments could be modified Journal of Cell Science • Advance article without affecting HIF-2α protein levels in liver cancer cells. by CK1δ suggesting that HIF-2α is directly phosphorylated by CK1δ in vitro predominantly between amino acids 366-542, a region that contains the ODD and the N-TAD domains. Identification of candidate CK1δ phosphorylation sites between amino acids 366-542 In order to determine the nature of the modified residues in the 366-542 region of HIF-2α that is targeted by CK1δ, two-dimensional TLC phospho-amino acid analysis was performed on GST-HIF-2α fragment 366-542 that was previously radioactively labeled by CK1δ and mixed with control phosphoamino acids (pSer, pThr, pTyr). The results of the analysis show that CK1δ phosphorylates both serine and threonine residues in the region 366-542 of HIF-2α (Fig. 3A). To map more precisely the CK1δ phosphorylation sites, recombinant full length GST-HIF-2α, as well as the smaller fragments 1-542 and 366-542 were incubated with or without CK1δ and ATP and analyzed by SDS-PAGE electrophoresis. Incubation with CK1δ caused slower migration of the GST-HIF-2α fragments, indicating quantitatively significant phosphorylation (Fig. 3B). The slower migrating bands were excised, digested by trypsin and subjected to analysis with mass spectroscopy. The analysis identified five unique tryptic peptides of HIF-2α (Tables S1, S2) and peptide GAVSEKSNFLFTK was shown to contain two phosphorylated serine residues (Ser383 and Ser386) (Fig. 3C). Sequence comparison revealed that at least one of the two serine residues was conserved in all known vertebrate HIF-2α homologues as well as in the human HIF-1α isoforms (Fig. 3D). Analysis by mass spectroscopy did not produce a candidate threonine residue in HIF-2α 366-542 region that could be modified by CK1δ, as suggested by the phospho-amino acid analysis. However, multiple sequence alignment analysis of this region of HIF-2α showed the presence of a small highly conserved area corresponding to amino acids 518-542 and the presence in this area of sequence motif E/DxxS/T (Fig. 3D). This residue, which is not present at the same position in HIF-1α, could be the target of CK1δ detected by our previous analysis. CK1δ modifies HIF-2α at Ser383 and Thr528 and enhances its transcriptional activity To verify the mass spectroscopy data for Ser383, Ser386 and test the hypothesis for Thr528 as targets of CK1δ, each one of these residues was converted into alanine by site directed mutagenesis. The three mutant proteins, HIF-2αS383A, HIF-2αS386A and HIF-2αT528A or the corresponding mutant 366-542 fragments were expressed as GST fusions in bacteria, purified and used as substrates for in vitro phosphorylation assays by recombinant CK1δ or Huh7 total cell extracts. Phosphorylation of HIF-2αS383A and HIF-2αT528A by CK1δ was Journal of Cell Science • Advance article a conserved threonine residue (Thr528) that was part of the classical CK1 consensus reduced by approximately 50% and 23%, respectively, relative to wild-type HIF-2α (Fig. 4A, upper panel), while phosphorylation of HIF-2α-366-542 S383A and HIF-2α-366-542 T528A was reduced by 49% and 40%, respectively, showing that CK1δ might preferentially target Ser383 (Fig. 4A, lower panel). When recombinant CK1δ was substituted by total soluble protein extracts of Huh7 cells as a kinase source, the impact on residues Ser383 and Thr528 appeared to be quite similar (Fig. 4B). On the other hand, mutation of Ser386 to Ala, did not cause any significant change in the in vitro phosphorylation by CK1δ (Fig. S2A). Taken together, our analysis identified residues Ser383 and Thr528 as the predominant CK1δ in vitro phosphorylation sites of human HIF-2α. The fact that these sites are also subjected to modification by the kinases present in the Huh7 total cell protein extracts indicates that CK1δ represents one of the kinases that interact with HIF-2α in Huh7 cells. To study the effect of Ser383 and Thr528 phosphorylation on HIF-2α function, we also constructed by site-directed mutagenesis the double SA mutant, (HIF-2αSA/TA), as well as the phospho-mimetic mutants HIF-2αS383D, HIF-2αT528E, HIF-2αSD/TE), the negative charges of which could potentially mimic the charges of the phosphorylated residues. Wildtype HIF-2α and all its mutant forms were sub-cloned into the pFLAG-CMV2 mammalian expression vector and introduced as Flag-tagged proteins into Huh7 and HepG2 cells by transient transfection. Western blotting analysis showed that the protein expression levels of all mutant forms were similar to those of the wild-type HIF-2α (Fig. 4C). Using a luciferase reporter gene assay, we observed that all the phosphorylation deficient mutant forms demonstrated an about 50% decrease in their HRE-dependent transcriptional activity compared with that of the wild-type HIF-2α (Fig. 4D). On the contrary, when the HREdependent transcriptional activity of the phospho-mimetic mutants was determined, we HIF-2α is subject to quantitative phosphorylation at the mutated sites. The data described so far show that both inhibition of CK1δ and the HIF-2αS383Α, HIF2αΤ528Α mutations negatively affect HIF-2 activity suggesting that Ser383 and Thr528 are true in vivo as well as in vitro targets of CK1δ. To further confirm that the exerted CK1δ effect is specific and linked to the modification of the Ser383 and Thr528 residues, the wildtype, phospho-deficient SA/TA and phospho-mimetic SD/TE mutant forms of HIF-2α were expressed as Flag-tagged proteins in HepG2 cells, and HIF-2 transcriptional activity was determined in the absence or presence of the CK1δ specific inhibitor D4476 under normoxia (i.e. in the absence of endogenous HIF-2α) (Fig. 4E). As expected, the wild-type form responded to inhibition of CK1δ by exhibiting significantly lower activity, exactly Journal of Cell Science • Advance article observed that it was similar to that of the wild-type HIF-2α form, suggesting that wild-type reproducing the behavior of endogenous hypoxically induced HIF-2α. However, neither the low transcriptional activities of the phospho-deficient HIF2-α mutant forms nor the normal (compared to wild-type) activities of the phospho-mimetic forms were significantly affected by the CK1δ inhibitor. These results strongly suggest that regulation of HIF-2α by CK1δ in hepatoma cell lines predominantly involves Ser383 and/or Thr528. Phosphorylation at Ser383 and/or Thr528 is required for HIF-2-dependent stimulation of EPO secretion from liver cancer cells To explore further the biological effect of Ser383 and Thr528 phosphorylation on HIF-2α function, we tested the mRNA expression of the endogenous HIF-2 specific target genes EPO and PAI-1 in Huh7 and HepG2 cells and observed that their induction by the phosphodeficient HIF-2α mutant forms was significantly lower relative to the wild-type HIF-2α form, suggesting that both Ser383 and Thr528 are required, most likely as CK1δ phosphorylation targets, for full activation of HIF-2 transcriptional function in both cell lines (Fig. 5A, 5C). Moreover, in agreement with the in vitro phosphorylation results, which did not support modification of Ser386, mutation of this site did not alter the expression of EPO and PAI-1 in relation to wild-type HIF-2α (Fig. S2B, S2C). To further confirm that the effect of CK1δ on EPO production is mediated by HIF-2α phosphorylation at Ser383 and/or Thr528, EPO secretion was analyzed in Huh7 and HepG2 cells expressing wild-type Flag-HIF-2α or its phospho-deficient mutant forms under normoxia. As shown in Fig. 5B&5D, mutations at the CK1δ phosphorylation sites Ser383 or Thr528 inhibited HIF-2α-mediated induction of EPO secretion. Overall, these data suggest that direct phosphorylation of HIF-2α at Ser383 or Thr528 by CK1δ increases HIF-2α transcriptional activity and subsequent EPO production in CK1δ does not affect protein stability of HIF-2α or its interaction with transcriptional co-activator USF2 To resolve the mechanism through which lack of phosphorylation by CK1δ, negatively affects HIF-2 transcriptional activity, we investigated first the effect of CK1δ inhibition on the stability of HIF-2α by determining its half-life after blocking translation or after termination of hypoxia. Huh7 cells that were incubated under hypoxia with or without D4476 were treated with the translation inhibitor cycloheximide, or subjected to re-oxygenation and HIF-2α expression was monitored for different time periods. As shown in Fig. 6A&6B, treatment with D4476 did not alter the protein stability of HIF-2α, which was as rapidly Journal of Cell Science • Advance article liver cancer cells. degraded as in the absence of D4476, both after inhibition of protein translation or reoxygenation. In addition, comparison at the same time point of 15 min suggests that, as expected, HIF-2α is more stable at hypoxia (without new protein synthesis) than at normoxia (despite ongoing protein translation). We then tested the effect of D4476 on the interaction of HIF-2 with the Upstream Stimulatory Factor (USF2), as it has been reported that this is necessary for optimal gene expression of EPO and PAI-1 (Pawlus et al., 2012). Huh7 cells overexpressing Flag-tagged USF2 and incubated under hypoxia in the presence or absence of D4476 were subjected to immunoprecipitation with an anti-Flag antibody. As shown in Fig. 6C, HIF-2α coprecipitated with Flag-USF2, as expected by their known association (Befani et al., 2013). However, its interaction with USF2 was not affected by D4476, suggesting that stimulation of their association is not the reason for the observed positive effect of CK1δ on HIF-2dependent activation of EPO and PAI-1 gene expression. Phosphorylation by CK1δ inhibits CRM1-dependent nuclear export of HIF-2α Having eliminated an effect of phosphorylation by CK1δ on HIF-2α stability or its interaction with the co-activator USF2, we next analyzed HIF-2α subcellular distribution. Wild-type Flag-HIF-2α and its mutant forms were expressed in transiently transfected HepG2 or Huh7 cells and their localization was monitored by immunofluorescence microscopy (Fig. 7A, S3 left panels). Wild-type Flag-HIF-2α was detected exclusively inside the nucleus in around 80% of the transfected cells and displayed both nuclear and cytoplasmic localization in only around 20% of the cells (Fig. 7B) When the same cells were when treated with D4476, wildtype Flag-HIF-2α was distributed both in the nucleus and the cytoplasm in about 50% of the Localization of the phospho-deficient mutant HIF-2α forms was both nuclear and cytoplasmic in about 50% of the cell population again suggesting that lack of modification by CK1δ impairs HIF-2α nuclear accumulation, whereas localization of all the phospho-mimetic mutant HIF-2α forms was indeed mimicking the distribution of the wild-type Flag-HIF-2α and was predominantly nuclear (Fig. S4A left panel, S4B). Increased detection of mutant HIF-2αS383A, HIF-2αT528A and HIF-2αSA/TA proteins or wild type HIF-2α protein in the presence of D4476 in the cytoplasm of the tested hepatic cancer cells, could be attributed either to reduced nuclear import or enhanced nuclear export. To distinguish between these two possibilities, cells were treated with Leptomycin B (LMB), a potent inhibitor of CRM1-dependent nuclear export. Exposure to LMB did not influence the Journal of Cell Science • Advance article cell population, suggesting that inhibition of CK1δ impairs its accumulation in the nucleus. localization of the wild type HIF-2α (Fig. 7A right panel, 7C) or the phosphomimetic mutant forms, which remained accumulated inside the nucleus (Fig. S4A right panel, S4C). However, LMB treatment caused exclusive nuclear localization of all the HIF-2α phosphodeficient mutant forms in almost 90% of the cells and abolished the effect of D4476 on the localization the wild type HIF-2α form (Fig. 7A right panel, 7C). These data indicate, first, that HIF-2α shuttles between nucleus and cytoplasm and, second, that lack of phosphorylation by CK1δ enhances HIF-2α nuclear export by CRM1 explaining the observed HIF-2 partial inactivation. Silencing of CK1δ enhances CRM1-dependent nuclear export of HIF-2α under hypoxia To further verify the above results, we tested the impact of CK1δ silencing on the localization of endogenous HIF-2α under hypoxia. CK1δ expression was suppressed by siRNA-mediated silencing in HeLa cells, which were then incubated under hypoxia in the presence or absence of LMB and HIF-2α localization was monitored by subcellular fractionation (Fig. 8A). As expected, HIF-2α was almost exclusively localized in the nuclear fraction under hypoxia independently of its exposure to LMB. However, in accordance to the results obtained for the overexpressed HIF-2α SA/TA mutant proteins above, a substantial fraction of HIF-2α protein migrated from the nucleus to the cytoplasm when CK1δ was silenced, which then returned to the nucleus upon LMB treatment. Taken together, these data show that loss of CK1δ promotes HIF-2α nuclear export, while LMB rescues the accumulation of HIF-2α inside the nucleus, confirming a CRM1-dependent mechanism controlled by phosphorylation. We next tested whether LMB-dependent re-localization of phospho-deficient HIF-2α mutants to the nucleus, could also restore their low transcriptional activity. Indeed, while LMB did the phospho-deficient mutants was increased and reached levels comparable to the wild-type activity (Fig. 8B). These data show that the predominant reason for the inactivation of HIF2α by mutations in the CK1δ sites is the enhanced exclusion of HIF-2α from the nucleus. We finally examined whether HIF-2α lacking modification by CK1δ (e.g. the ST/AA mutant) may change gene target specificity and activate preferentially HIF-1 target genes (such as PGK1) while being less active on HIF-2 target genes (such as EPO). However, neither overexpression of wild-type HIF-2α nor of any of the mutant forms affected the normoxic expression of the HIF-1-specific PGK1 gene, irrespective of the presence or not of LMB (Fig. S4D, S4E), indicating, that HIF-2 cannot replace HIF-1 following inhibition of CK1δmediated phosphorylation of HIF-2α. Journal of Cell Science • Advance article not affect the activity of wild-type HIF-2α or its phospho-mimetic mutants, the activity of all DISCUSSION In this study, we have identified and characterized a novel regulatory mechanism of HIF-2α activity involving CK1δ-mediated phosphorylation of two distinct residues of HIF-2α, serine at position 383 and threonine at position 528. Our conclusion, that phosphorylation of HIF-2α by CK1δ increases EPO production by blocking CRM1-dependent export of HIF-2α to the cytoplasm (Fig. 8C) is based on the following observations: First, CK1δ silencing or inhibition, decreases hypoxic induction of two HIF-2-specific target genes, EPO and PAI-1, without affecting HIF-2α protein expression. Second, CK1δ modifies HIF-2α residues Ser383 and Thr528 in vitro. Third, blocking Ser383 or Thr528 phosphorylation (by converting them into Ala) inhibits HIF-2 transcriptional activity while phospho-mimetic mutations of the same residues preserve HIF-2 activity. Fourth, CK1δ does not affect HIF-2α protein stability or its interaction with the USF2 co-activator but controls CRM1-dependent nuclear export of HIF-2α. Finally, either suppression of CK1δ activity or mutations of the CK1δ phosphorylation sites on HIF-2α, strongly reduce EPO secretion in two HCC cell lines. The first identified site of CK1δ modification, Ser-383, is between the PAS-B and ODD domains of HIF-2α. The crystal structure of human HIF-2α that contains the PAS-B domain has been determined only until the 350th residue (Erbel et al., 2003, Wu et al., 2015) so it is not certain which of the two domains Ser383 belongs to, which makes prediction of its involvement in HIF-2α regulation difficult. However, the second CK1δ phosphorylation site, Thr528, is localized in a region, in which the ODD and N-TAD domains overlap (Maemura et al., 1999, O'Rourke et al., 1999). The ODD is involved in oxygen-dependent regulation, replacement by the analogous region of HIF-1α switches their specificity (Hu et al., 2007). Therefore, Thr528 phosphorylation could be involved in either of the two processes. Oxygendependent regulation of HIF-2α mediated by its ODD domain involves the well characterized PHD-VHL-proteasome-system and is tightly regulated by hydroxylation events on residues Pro405 and Pro531 by the PHDs. Thr528 is found in close proximity to Pro531 and its modification by CK1δ could affect hydroxylation of HIF-2α and therefore its stability. However, the results from our experiments analyzing the effect of CK1δ inhibition on the half-life of HIF-2α after CHX treatment or re-oxygenation do not favor this scenario as they did not show any significant changes in HIF-2α stability. Alternatively, Thr528 phosphorylation may be involved in the function of N-TAD, and more specifically in the Journal of Cell Science • Advance article while the N-TAD plays an important role in the activation of HIF-2α-specific genes, as its interaction of HIF-2α with other transcriptional co-activators. It is well established that multiple transcription factors (TFs) are required to achieve maximal activation of target genes in response to a specific stimulus. This multifactorial transcription complex has been termed the “enhanceosome” (Thanos and Maniatis, 1995). USF2 has been identified as a HIF-2specific co-transcriptional factor that not only is required for HIF-2 target gene activation but also is required for HIF-2-dependent tumorigenicity in vitro in several cell lines including mES and Hep3B cells (Pawlus et al., 2012). It has additionally been shown that USF2 binds to HIF-2 target genes in vivo and contributes significantly to hypoxia-mediated increase in CBP and p300 binding to the HIF-2 target genes PAI-1 and EPO. It could have, therefore, been possible that inhibition of expression of these genes, that we observed when CK1δ was inhibited, was due to diminished interaction between the under-modified Ν-ΤΑD of HIF-2α and USF2. However our immunoprecipitation experiments showed that inhibition of CK1δ did not affect the interaction between HIF-2 and USF2 under hypoxia, making involvement of the Thr528 in this process rather unlikely. In our effort to identify the reason for diminished activity of the mutant forms of HIF-2α that lacked Ser383 or Thr528, we detected their significant cytoplasmic mislocalization, which was also observed for wild-type HIF-2α in the presence of a CK1δ inhibitor. More strikingly, nuclear accumulation was re-established when nuclear protein export was inhibited by Leptomycin B, suggesting that phosphorylation by CK1δ is required to maintain HIF-2α inside the nucleus. Moreover, LMB treatment completely restored the diminished activity of the phospho-deficient mutants, indicating the critical role of the mutated residues. These results were verified under hypoxia and for endogenous HIF-2α, which translocated to the cytoplasm in CK1δ-silenced cells, but re-accumulated in the nucleus upon LMB treatment. accumulation and full activation of shuttling transcription factors as has been shown before in several cases (Ikuta et al., 2004, Lee and Bai, 2002, Zhang and Xiong, 2001). Moreover, CK1 has been reported to regulate the nucleocytoplasmic transport of the eIF6 translation initiation factor by promoting its CRM1-dependent nuclear export through direct phosphorylation (Biswas et al., 2011). Taking all into account, we suggest that phosphorylation of HIF-2α at Ser-383 or Thr-528 by CK1δ prevents its CRM1-mediated nuclear export and is therefore required for its efficient accumulation inside the nucleus and full HIF-2 activity. How exactly these modifications interfere with the nuclear export of HIF-2α is certainly an interesting issue that merits further future investigation. Nuclear export of HIF-1α is also regulated by phosphorylation, but in this case modification is catalyzed by ERK1/2 that target two serine Journal of Cell Science • Advance article Inhibition of nuclear export by phosphorylation is indeed an efficient way to ensure nuclear residues (Ser641 and Ser643) adjacent to a non-conventional CRM1-dependent nuclear export signal (NES; (Mylonis et al., 2008, Mylonis et al., 2006). However, neither these ERK sites nor the NES of HIF-1α are conserved in HIF-2α. Regulation of HIF-2α and HIF-1α by CK1δ represents a new example of distinct-opposite regulation between the two isoforms. Here we have shown that HIF-2α modification by CK1δ at Ser383 or Thr528 enhances its transcriptional activity by promoting its nuclear accumulation. On the other hand, our lab has previously reported that HIF-1α is modified by CK1 at Ser247 and this inhibits HIF-1 activity by blocking HIF-1/ARNT interaction (Kalousi et al., 2010, Kourti et al., 2015). Comparative sequence analysis reveals that the two isoforms display absolute homology in the PAS-B region where the phosphorylation of HIF-1α occurs (Table S3). However, our mapping experiments excluded phosphorylation of the corresponding HIF-2α domain by CK1δ in vitro and instead led to the identification of Ser383 or Thr528, which lie C-terminally to the PAS-B domain. Ser383 is conserved in the HIF-1α isoform but Thr528 is not. So despite the overall sequence similarities between HIF1α and HIF-2α, they are targeted by CK1δ at distinct sites and with opposite functional outcome. It is not without precedent that a single factor mediates distinct regulation between the two HIF-α isoforms. HIF-associated factor (HAF) has been reported to destabilize HIF1α in proteasome-dependent manner by binding to its N-terminal domain (Koh et al., 2008) but, on the contrary, HAF promotes HIF-2 transcriptional activity by binding to its C-terminal domain (Koh et al., 2011). It is probably important for cells that express both HIF-α isoforms to promote the activity of the one isoform when the other one is inactivated under certain conditions and in a tissue-specific manner and CK1δ may be involved in regulating this balance. So high CK1δ expression and/or activity may mediate preferential HIF-2α activation CK1 family members are evolutionarily conserved serine/threonine kinases members involved in many cellular processes including proliferation, cell cycle progression, DNA repair, chromosome segregation and have been reported to target several transcription factors (such as TP53, Foxo1, HIF-1α) (Bischof et al., 2011, Cheong and Virshup, 2011, Greer and Rubin, 2011, Gross and Anderson, 1998, Kalousi et al., 2010, Knippschild et al., 2005, Schittek and Sinnberg, 2014, Knippschild et al., 2014). HIF-2α is added to this list for the first time. HIF-2α is expressed in hepatocytes, is the main regulator of hepatic EPO production (Rankin et al., 2007) and we have now shown that this function can be controlled by CK1δ. During fetal development, EPO is mainly produced in the liver. Its primary site of Journal of Cell Science • Advance article in cells expressing both HIF-α isoforms. production changes to the kidney during late gestation but EPO continues to be produced in the liver to a lesser extent. Therefore, hepatocytes are the primary source of extra-renal EPO in the adult (Koury et al., 1991) and under conditions of severe hypoxia, liver EPO production increases and may account for more than 33% of total EPO production (Eckardt and Kurtz, 2005). Dysregulated EPO expression results in the development of anemia, when serum EPO levels are inadequate, or polycythemia as a result of EPO overproduction (Rankin et al., 2007). Therefore, the identification of CK1δ as a kinase regulating HIF-2-dependent EPO secretion from liver cancer cells has strong physiological significance and may have important therapeutic applications in these conditions. Moreover, Ribatti et al, have demonstrated that EPO and its receptor are involved in highly vascularized human hepatocellular carcinoma (Ribatti et al., 2007). In this case, EPO is secreted by hepatic tumor cells and it acts on vascular endothelial cells via their EPOR receptors, promoting angiogenesis. It has also been observed that some cases of HCC are associated with abnormally high levels of erythrocytes. This HCC-associated erythrocytosis is thought to result via increased EPO production by the HCC cells and many HCC patients have high plasma EPO levels (Sakisaka et al., 1993). Taking into account the important role of HIF-2 in HCC, many compounds that directly interfere with HIF-2 function or target downstream hypoxia-related pathways have been tested in experimental trials as anti-cancer agents but have proven ineffective (Zhao et al., 2015). The novel regulatory communication between CK1δ and HIF-2 described in this work suggests an alternative means to control HIF-2 activity in disease that needs to be further Journal of Cell Science • Advance article investigated. MATERIALS AND METHODS Plasmid Constructions cDNAs encoding the various fragments of HIF-2α were produced by PCR and were cloned as BamHI fragments into pGEX-4T-1 bacterial expression vector (Amersham Biosciences), yielding pGEX-HIF-2α full length (1-870) and pGEX-HIF-2α fragments 1-820, 1-679, 1-542, 1-366, 1-280, 366-870, 366-820, 366-679, 366-542, 542-870, 542-820. Single point HIF-2α mutants S383A, S383D, S386A, T528A and T528E and double point HIF-2α mutants S383A/T528A, S383D/T528E were constructed using as template pBS-SK(+)-HIF-2α containing the full-length cDNA of HIF-2α, with the QuikChange II site-directed mutagenesis kit (Agilent technologies). The primer sequences used are available upon request. The DNA sequence of the point mutants was confirmed by sequencing performed by VbC Biotech. All mutant forms of HIF-2α were then subcloned as BamHI fragments into the mammalian expression vector pFlag-CMV2 and into the pGEX-4T-1 bacterial expression vector. Using as templates the pFlag-CMV2 HIF-2α point mutant forms, we obtained by PCR the cDNA fragments corresponding to 366-542 S383A, S386A and T528A. The derived mutant fragments were subcloned as BamHI fragments into the pGEX-4T-1 bacterial expression vector. The full-length pCDNA-Flag-USF2 plasmid was kindly provided by CJ Hu (Department of Craniofacial Biology, University of Colorado) (Pawlus et al., 2012). Cell culture, Transfection and Luciferase Assays Human Huh7 and HepG2 cells were purchased from the European Collection of Cell Cultures (ECACC). Human HeLa cells were purchased from the American Type Culture Collection (ATCC). Cells were cultured in DMEM (Biosera) containing 10% FBS and 100 and 6-cm dishes or 12-well plates by using Turbofect (Thermo Scientific) Transfection Reagent. When required, 24 h after transfection, cells were treated for 16 h with the CK1δ inhibitor D4476 (10 μM, Cayman) for 4 h with 10 ng/ml Leptomycin B (Sigma-Aldrich) or with DMSO as solvent control. For hypoxic treatment, cells were exposed for 16 h to 1% O2, 94% N2, and 5% CO2 in an IN VIVO2 200 hypoxia workstation (Ruskinn Life Sciences). For half-life studies after 4 h of hypoxia treatment, cells were treated with 10 μg/ml Cycloheximide (Sigma-Aldrich) or were re-oxygenated in a time-depended manner. Luciferase assays were performed in cells transiently co-transfected with equal amounts of plasmids expressing different forms of HIF-2α (or the empty parental vector pFlag-CMV2 as Journal of Cell Science • Advance article U/ml penicillin–streptomycin (Biosera). Transient transfections were carried out in 10-cm control), the firefly luciferase reporter plasmid pGL3–5HRE-VEGF, kindly provided by Dr. A. J. Giacia (Stanford University) and the Renilla luciferase expressing plasmid pCI-Renilla, generously provided by Dr. M. U. Muckenthaler (University of Heidelberg, Germany). 24 h post-transfection, cells were washed with PBS, lysed, and luciferase activity was determined using the luciferase assay kit or Dual Luciferase Reporter Assay System (Promega, Madison, WI). siRNA-mediated silencing of CK1δ mRNA encoding CK1δ was targeted using two distinct sets of validated siRNA (Qiagen): Hs-CSNK1D5 HP (CCGGTCTAGGATCGAAATGTT) and Hs-CSNK1D6 HP (CTCCCTGACGATTCCACTGTA). AllStarssiRNA (Qiagen) was used as negative control. Cells were incubated in serum-free DMEM for 4 h with siRNA (10 nM) in the presence of Lipofectamine™ RNAiMAX (Invitrogen, Life Technologies) as described by the manufacturer. 24 h post-transfection, cells were exposed to hypoxia and 16 h later they were harvested, and cell lysates were prepared for real time PCR or for western blot analysis. RNA extraction, quantitative PCR and quantitation of EPO production Total RNA from Huh7 and HepG2 cells was isolated using the peqGOLDTriFast reagent (Peqlab) and cDNA was synthesized with the High Capacity Reverse Transcription kit (Applied Biosystems). Real-time PCR was performed with SYBRGreen qPCRSuperMix Universal (Invitrogen) in a MiniOpticon instrument (Bio-Rad). The mRNAs encoding EPO, PAI-1 and HPRT1 were amplified using primers available upon request. Each sample was assayed in duplicate for both target and internal control. Relative quantitative gene expression relation to the respective controls. For quantification of EPO secretion, medium was collected from 105 cells in 6-cm culture dishes and EPO in the medium was measured by using the Human Erythropoietin DRG® EPO (Erythropoietin) ELISA (EIA-3646) Kit (DRG International Inc., USA). SDS-PAGE, Western blot and Subcellular Fractionation Proteins were resolved by 10% SDS-PAGE, and analyzed by Coomassie Blue or Western blotting using the antibodies: anti-HIF-2α pAb (Novus Biologicals, NB100-122), anti-GST pAb, anti-Flag mAb (Sigma, F4042), anti-Tubulin mAb (Cell Signaling #3873), anti-CK1δ pAb (Santa Cruz, sc-55553) and anti-USF2 pAb (Santa Cruz, sc-862). Analysis by Journal of Cell Science • Advance article was calculated using the ΔΔCT method and presented as fold increase or percent activity in immunoblotting was carried out as previously described (Lyberopoulou et al., 2007, Mylonis et al., 2008). Subcellular fractionations were performed at HeLa cells (because of technical problems encountered with the fractionation of hepatoma cells) as previously described (Andrews and Faller, 1991). Western blot images were taken using an Uvitec Cambridge Chemiluminescence Imaging System with the help of Alliance Software (ver. 16.06) and quantified by Uviband Software (ver. 15.03) provided with the instrument (Uvitec Cambridge). Immunoprecipitation and Immunofluorescence For immunoprecipitation experiments, cells were transfected with pCDNA-Flag-USF2 and 24 h post-transfection were incubated for 16 h under hypoxia in the presence or absence of D4476. Immunoprecipitation analysis was carried out as previously described (Mylonis et al., 2006). For immunofluorescence experiments, cells were grown on coverslips and transiently transfected with pCMV2-FLAG vector expressing HIF-2α wild-type and its mutant forms. In some cases, 4 h post-transfection cells were treated with D4476 for 16 h, or treated with Leptomycin B 4 h prior to observation. Coverslips were incubated for 1 h at room temperature with anti-Flag mAb (Sigma), washed twice with PBS, and incubated for 1h at room temperature with Cy3-conjugated anti-mouse secondary antibody (Jackson ImmunoResearch). Analysis by immunofluorescence was carried out as previously described (Mylonis et al., 2008). Images were taken on a Zeiss Axioplan fluorescence microscope using an AxioCam MRm CCD sensor and 40x objective with filters for DAPI and Cy3. Protein Purification and Phosphorylation Assays 366, 1-280, 366-870, 366-820, 366-679, 366-542, 542-870, 542-820 were expressed in Escherichia coli and purified as previously reported for GST-HIF-1α (Chachami et al., 2005). Phosphorylation reactions were carried out as previously described (Mylonis et al., 2006). The reaction was stopped by adding SDS sample buffer and heating at 95 °C for 5 min. Samples were analyzed by SDS-PAGE followed by Coomassie Blue staining and autoradiography. Relative phosphorylation levels were determined by cutting out protein bands from gels and measuring their radioactivity by scintillation counting. Journal of Cell Science • Advance article Wild Type and mutant forms of GST-HIF-2α and its GST fragments 1-820, 1-679, 1-542, 1- Phosphoamino acid analysis. P-labelled GST-HIF-2α (366-542) was analyzed by SDS–PAGE, blotted onto immobilon 32 membrane and the relevant radioactive band was excised. The excised strip was treated with 5.7 M HCl at 110 °C for 90 min, and the supernatant was dried in a Speed-Vac concentrator. The lyophilized powder was reconstituted in a solution containing phospho-serine, phosphothreonine and phospho-tyrosine (10 nmole each; Sigma) followed by two-dimensional chromatographic separation (Grangeasse et al., 1999). Briefly, migration in the first dimension was performed by chromatography in a solvent containing propanol/water at a ratio of 70:30 for 90-120 min on a thin layer silica plate (Merk), and in the second dimension in a solvent containing isobutyl alcool/formic acid/ water at a ratio of 80:30:40 for 90-120 min. After separation and drying, phosphoamino acids were visualized by ninhydrin staining (0.5% in acetone) followed by autoradiography. In-gel tryptic digestion The stained gel band corresponding to the HIF-2α construct (aa366-542) was excised and cut into small pieces in order to proceed with in-gel tryptic protein digestion according to standard procedures (Shevchenko et al., 2006). Briefly, the gel pieces were destained with 100 mM ammonium bicarbonate/ acetonitrile (1:1), dehydrated with acetonitrile (ACN) and then rehydrated with 25 mM ammonium bicarbonate (NH4HCO3) buffer. The dried gel pieces were subsequently rehydrated with 12.5 ng/µl trypsin (trypsin gold, Promega) in 25 mM NH4HCO3buffer and incubated on ice for 45 minutes. The samples were finally transferred to a thermomixer (Eppendorf) and incubated overnight at 37 °C with mild mixing (300 rpm). containing 5% formic acid (v/v) with shaking for one hour. The extracted peptide solution was dried down in a vacuum centrifuge (Savant). The samples were reconstituted with 30 μl solution of 2% (v/v) acetonitrile / 0.1% (v/v) formic acid, sonicated in a water bath for 3 minutes and analyzed with LC-MS/MS. LC-MS/MS analysis The purified peptides were analyzed by HPLC MS/MS using coupled to an LTQ Orbitrap XL Mass spectrometer (Thermo Fischer Scientific, Waltham, MA, USA) equipped with a nanospray source. Ten μl of the peptide mixtures were pre-concentrated at a flow-rate of 5μl/min for 10 min using a C18 trap column (Acclaim PepMap) and then loaded onto a 50 Journal of Cell Science • Advance article Next day, peptides were extracted from the gel pieces by addition of 150 μl acetonitrile cm C18 column (75 μm ID, particle size 2 μm, 100Å, Acclaim PepMap RSLC, Thermo Scientific). The binary pumps of the HPLC (RSLCnano, Thermo Scientific) contained solution A (2% (v/v) ACN in 0.1% (v/v) formic acid) and solution B (80% ACN in 0.1% formic acid). The peptides were separated using a linear gradient of 4% - 40% B in 55 min at a flow rate of 300 nl/min. The column was placed in an oven operating at 35 °C. Full scan MS spectra were acquired in the orbitrap (m/z 300–1600) in profile mode and data-dependent acquisition with the resolution set to 60,000 at m/z 400 and automatic gain control target at 106. The six most intense ions were sequentially isolated and multistage activation was used in order to generate a more sequence-informative fragments detected in the linear ion trap. Dynamic exclusion was set to 60 sec. Ions with single charge states were excluded. Lockmass of m/z 445,120025 was used for internal calibration. The software Xcalibur (Thermo Scientific) was used to control the system and acquire the raw files. Data analysis and database search Protein identification was performed using Proteome Discoverer 1.3 software (Thermo Fischer Scientific, Waltham, MA, USA) equipped with SEQUEST HT search engine. Peak lists were searched against the Uniprot database with an initial mass deviation of 10 ppm, fragmentation mass deviation of 0.8 Da and trypsin digestion specificity allowing up to two missed cleavages. Oxidation of methionine, phosphorylation of threonines and serines and acetylation of the protein N-terminus were used as variable modifications. The peptides were filtered according to their XCorr score versus charge state (XCorr:+2 ≥ 1.8 and +3 ≥ 2.5) high confidence peptides were evaluated for their phosphorylation status. The algorithm Statistical analysis The Graph Pad Instat Statistical package for Windows was used. Data are expressed as mean ± standard error of the mean (± s.e.m). Differences were examined by Student's t-test (twotailed) between two groups or by one-way analysis of variance (ANOVA) within multiple groups. P<0.05 was considered statistically significant. Journal of Cell Science • Advance article phosphoRS was used in order to assign and score the phosphosite probabilities. Acknowledgements We thank Dr. C.J. Hu (University of Colorado) for providing USF2 plasmid. Competing interests: The authors declare that there were no conflicts of interest regarding the publication of this paper. Author contributions E.P. performed experiments and wrote the paper, C.B. performed experiments and assisted with data analysis, I.M. assisted with the in vitro phosphorylation and subcellular fractionation assays and data processing, M.S. and G.P. performed experiments with LCMS/MS and data analysis, G.S. supervised experiments and wrote the paper, L.P. designed and supervised experiments and wrote the paper. Funding This work was partially funded by the ARISTEIA II Action of the Operational Program EDUCATION AND LIFELONG LEARNING (grant HYPOXYTARGET, 3129 to G.S.) co- Journal of Cell Science • Advance article funded by the European Social Fund (ESF) and Greek National Resources. REFERENCES Andrews, N. C. & Faller, D. V. (1991). A rapid micropreparation technique for extraction of DNA-binding proteins from limiting numbers of mammalian cells. Nucleic Acids Res, 19, 2499. Befani, C., Mylonis, I., Gkotinakou, I. M., Georgoulias, P., Hu, C. J., Simos, G. & Liakos, P. (2013). Cobalt stimulates HIF-1-dependent but inhibits HIF-2-dependent gene expression in liver cancer cells. Int J Biochem Cell Biol, 45, 2359-68. Bischof, J., Muller, A., Fander, M., Knippschild, U. & Fischer, D. (2011). Neurite outgrowth of mature retinal ganglion cells and PC12 cells requires activity of CK1delta and CK1epsilon. PLoS One, 6, e20857. Biswas, A., Mukherjee, S., Das, S., Shields, D., Chow, C. W. & Maitra, U. (2011). Opposing action of casein kinase 1 and calcineurin in nucleo-cytoplasmic shuttling of mammalian translation initiation factor eIF6. J Biol Chem, 286, 3129-38. Chachami, G., Paraskeva, E., Georgatsou, E., Bonanou, S. & Simos, G. (2005). Bacterially produced human HIF-1alpha is competent for heterodimerization and specific DNAbinding. Biochem Biophys Res Commun, 331, 464-70. Chen, R., Xu, M., Hogg, R. T., Li, J., Little, B., Gerard, R. D. & Garcia, J. A. (2012). The acetylase/deacetylase couple CREB-binding protein/Sirtuin 1 controls hypoxiainducible factor 2 signaling. J Biol Chem, 287, 30800-11. Cheong, J. K. & Virshup, D. M. (2011). Casein kinase 1: Complexity in the family. Int J Biochem Cell Biol, 43, 465-9. Dioum, E. M., Chen, R., Alexander, M. S., Zhang, Q., Hogg, R. T., Gerard, R. D. & Garcia, J. A. (2009). Regulation of hypoxia-inducible factor 2alpha signaling by the stress- Eckardt, K. U. & Kurtz, A. (2005). Regulation of erythropoietin production. Eur J Clin Invest, 35 Suppl 3, 13-9. Erbel, P. J., Card, P. B., Karakuzu, O., Bruick, R. K. & Gardner, K. H. (2003). Structural basis for PAS domain heterodimerization in the basic helix--loop--helix-PAS transcription factor hypoxia-inducible factor. Proc Natl Acad Sci U S A, 100, 15504-9. Gradin, K., Takasaki, C., Fujii-Kuriyama, Y. & Sogawa, K. (2002). The transcriptional activation function of the HIF-like factor requires phosphorylation at a conserved threonine. J Biol Chem, 277, 23508-14. Journal of Cell Science • Advance article responsive deacetylase sirtuin 1. Science, 324, 1289-93. Grangeasse, C., Riberty, M., Vaganay, E. & Duclos, B. (1999). Alternative procedure for two-dimensional separation of phosphoamino acids. Biotechniques, 27, 62-4. Greer, Y. E. & Rubin, J. S. (2011). The role of centrosomal casein kinase 1 delta in neurite outgrowth and beyond. Cell Cycle, 10, 2605-6. Gross, S. D. & Anderson, R. A. (1998). Casein kinase I: spatial organization and positioning of a multifunctional protein kinase family. Cell Signal, 10, 699-711. Hu, C. J., Sataur, A., Wang, L., Chen, H. & Simon, M. C. (2007). The N-terminal transactivation domain confers target gene specificity of hypoxia-inducible factors HIF-1alpha and HIF-2alpha. Mol Biol Cell, 18, 4528-42. Ikuta, T., Kobayashi, Y. & Kawajiri, K. (2004). Cell density regulates intracellular localization of aryl hydrocarbon receptor. J Biol Chem, 279, 19209-16. Kaelin, W. G., Jr. & Ratcliffe, P. J. (2008). Oxygen sensing by metazoans: the central role of the HIF hydroxylase pathway. Mol Cell, 30, 393-402. Kalousi, A., Mylonis, I., Politou, A. S., Chachami, G., Paraskeva, E. & Simos, G. (2010). Casein kinase 1 regulates human hypoxia-inducible factor HIF-1. J Cell Sci, 123, 2976-86. Keith, B., Johnson, R. S. & Simon, M. C. (2012). HIF1alpha and HIF2alpha: sibling rivalry in hypoxic tumour growth and progression. Nat Rev Cancer, 12, 9-22. Kietzmann, T., Mennerich, D. & Dimova, E. Y. (2016). Hypoxia-Inducible Factors (HIFs) and Phosphorylation: Impact on Stability, Localization, and Transactivity. Front Cell Dev Biol, 4, 11. Knippschild, U., Gocht, A., Wolff, S., Huber, N., Lohler, J. & Stoter, M. (2005). The casein kinase 1 family: participation in multiple cellular processes in eukaryotes. Cell Signal, Knippschild, U., Kruger, M., Richter, J., Xu, P., Garcia-Reyes, B., Peifer, C., Halekotte, J., Bakulev, V. & Bischof, J. (2014). The CK1 Family: Contribution to Cellular Stress Response and Its Role in Carcinogenesis. Front Oncol, 4, 96. Koh, M. Y., Darnay, B. G. & Powis, G. (2008). Hypoxia-associated factor, a novel E3ubiquitin ligase, binds and ubiquitinates hypoxia-inducible factor 1alpha, leading to its oxygen-independent degradation. Mol Cell Biol, 28, 7081-95. Koh, M. Y., Lemos, R., Jr., Liu, X. & Powis, G. (2011). The hypoxia-associated factor switches cells from HIF-1alpha- to HIF-2alpha-dependent signaling promoting stem cell characteristics, aggressive tumor growth and invasion. Cancer Res, 71, 4015-27. Journal of Cell Science • Advance article 17, 675-89. Kourti, M., Ikonomou, G., Giakoumakis, N. N., Rapsomaniki, M. A., Landegren, U., Siniossoglou, S., Lygerou, Z., Simos, G. & Mylonis, I. (2015). CK1delta restrains lipin-1 induction, lipid droplet formation and cell proliferation under hypoxia by reducing HIF-1alpha/ARNT complex formation. Cell Signal, 27, 1129-40. Koury, S. T., Bondurant, M. C., Koury, M. J. & Semenza, G. L. (1991). Localization of cells producing erythropoietin in murine liver by in situ hybridization. Blood, 77, 2497503. Lee, H. & Bai, W. (2002). Regulation of estrogen receptor nuclear export by ligand-induced and p38-mediated receptor phosphorylation. Mol Cell Biol, 22, 5835-45. Lim, J. H., Lee, Y. M., Chun, Y. S., Chen, J., Kim, J. E. & Park, J. W. (2010). Sirtuin 1 modulates cellular responses to hypoxia by deacetylating hypoxia-inducible factor 1alpha. Mol Cell, 38, 864-78. Lyberopoulou, A., Venieris, E., Mylonis, I., Chachami, G., Pappas, I., Simos, G., Bonanou, S. & Georgatsou, E. (2007). MgcRacGAP interacts with HIF-1alpha and regulates its transcriptional activity. Cell Physiol Biochem, 20, 995-1006. Maemura, K., Hsieh, C. M., Jain, M. K., Fukumoto, S., Layne, M. D., Liu, Y., Kourembanas, S., Yet, S. F., Perrella, M. A. & Lee, M. E. (1999). Generation of a dominant-negative mutant of endothelial PAS domain protein 1 by deletion of a potent C-terminal transactivation domain. J Biol Chem, 274, 31565-70. Majmundar, A. J., Wong, W. J. & Simon, M. C. (2010). Hypoxia-inducible factors and the response to hypoxic stress. Mol Cell, 40, 294-309. Mylonis, I., Chachami, G., Paraskeva, E. & Simos, G. (2008). Atypical CRM1-dependent nuclear export signal mediates regulation of hypoxia-inducible factor-1alpha by Mylonis, I., Chachami, G., Samiotaki, M., Panayotou, G., Paraskeva, E., Kalousi, A., Georgatsou, E., Bonanou, S. & Simos, G. (2006). Identification of MAPK phosphorylation sites and their role in the localization and activity of hypoxiainducible factor-1alpha. J Biol Chem, 281, 33095-106. O'rourke, J. F., Tian, Y. M., Ratcliffe, P. J. & Pugh, C. W. (1999). Oxygen-regulated and transactivating domains in endothelial PAS protein 1: comparison with hypoxiainducible factor-1alpha. J Biol Chem, 274, 2060-71. Ohh, M., Park, C. W., Ivan, M., Hoffman, M. A., Kim, T. Y., Huang, L. E., Pavletich, N., Chau, V. & Kaelin, W. G. (2000). Ubiquitination of hypoxia-inducible factor requires Journal of Cell Science • Advance article MAPK. J Biol Chem, 283, 27620-7. direct binding to the beta-domain of the von Hippel-Lindau protein. Nat Cell Biol, 2, 423-7. Patel, S. A. & Simon, M. C. (2008). Biology of hypoxia-inducible factor-2alpha in development and disease. Cell Death Differ, 15, 628-34. Pawlus, M. R., Wang, L., Ware, K. & Hu, C. J. (2012). Upstream stimulatory factor 2 and hypoxia-inducible factor 2alpha (HIF2alpha) cooperatively activate HIF2 target genes during hypoxia. Mol Cell Biol, 32, 4595-610. Rankin, E. B., Biju, M. P., Liu, Q., Unger, T. L., Rha, J., Johnson, R. S., Simon, M. C., Keith, B. & Haase, V. H. (2007). Hypoxia-inducible factor-2 (HIF-2) regulates hepatic erythropoietin in vivo. J Clin Invest, 117, 1068-77. Rena, G., Bain, J., Elliott, M. & Cohen, P. (2004). D4476, a cell-permeant inhibitor of CK1, suppresses the site-specific phosphorylation and nuclear exclusion of FOXO1a. EMBO Rep, 5, 60-5. Ribatti, D., Marzullo, A., Gentile, A., Longo, V., Nico, B., Vacca, A. & Dammacco, F. (2007). Erythropoietin/erythropoietin-receptor system is involved in angiogenesis in human hepatocellular carcinoma. Histopathology, 50, 591-6. Sakisaka, S., Watanabe, M., Tateishi, H., Harada, M., Shakado, S., Mimura, Y., Gondo, K., Yoshitake, M., Noguchi, K., Hino, T. & Et Al. (1993). Erythropoietin production in hepatocellular carcinoma cells associated with polycythemia: immunohistochemical evidence. Hepatology, 18, 1357-62. Schittek, B. & Sinnberg, T. (2014). Biological functions of casein kinase 1 isoforms and putative roles in tumorigenesis. Mol Cancer, 13, 231. Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. (2006). In-gel digestion for 60. Thanos, D. & Maniatis, T. (1995). Virus induction of human IFN beta gene expression requires the assembly of an enhanceosome. Cell, 83, 1091-100. To, K. K., Sedelnikova, O. A., Samons, M., Bonner, W. M. & Huang, L. E. (2006). The phosphorylation status of PAS-B distinguishes HIF-1alpha from HIF-2alpha in NBS1 repression. EMBO J, 25, 4784-94. Wu, D., Potluri, N., Lu, J., Kim, Y. & Rastinejad, F. (2015). Structural integration in hypoxia-inducible factors. Nature, 524, 303-8. Zhang, Y. & Xiong, Y. (2001). A p53 amino-terminal nuclear export signal inhibited by DNA damage-induced phosphorylation. Science, 292, 1910-5. Journal of Cell Science • Advance article mass spectrometric characterization of proteins and proteomes. Nat Protoc, 1, 2856- Zhao, J., Du, F., Luo, Y., Shen, G., Zheng, F. & Xu, B. (2015). The emerging role of hypoxia-inducible factor-2 involved in chemo/radioresistance in solid tumors. Cancer Journal of Cell Science • Advance article Treat Rev, 41, 623-33. Fig. 1. Silencing or inhibition of CK1δ impairs HIF-2 transcriptional activity and EPO secretion. (A) Western blot analysis of lysates from Huh7 cells transfected with CK1δ siRNA (10nM) or scrambled siRNA (10nM). Cells were incubated for 16 h under hypoxia before collection and lysis. (B) Determination of EPO mRNA levels by quantitative real-time PCR in Huh7 Journal of Cell Science • Advance article Figures cells incubated and treated as in A. Results are shown as fold increase in relation to the corresponding normoxic conditions and represent the mean of three independent experiments performed in duplicate (± s.e.m). (C) Determination of EPO secretion by ELISA in Huh7 cells incubated and treated as in A. Values are expressed as mlU/ml protein and represent the mean of three independent experiments performed in duplicate (± s.e.m). (D) Western blot analysis of lysates from Huh7 cells incubated for 16 h under normoxia or hypoxia (1% O2), in the presence or absence of the specific CK1δ inhibitor D4476 (10 μΜ). (E) Determination of EPO and PAI-1 mRNA levels by quantitative real-time PCR in Huh7 cells incubated and treated as in D. Results are shown as in B. (F) Determination of EPO secretion by ELISA in Journal of Cell Science • Advance article Huh7 cells incubated and treated as in D. Values are shown as shown as in C. (A) Schematic representation of HIF-2α, its domains and fragments that were cloned, expressed and purified as GST fusion proteins. The parts of HIF-2α that were identified as substrates for recombinant CK1δ based on the results presented in B are marked by +. (B) The indicated GST tagged HIF-2α fragments were subjected to in vitro phosphorylation by 100 units of recombinant CK1δ and analyzed by SDS-PAGE followed by Coomassie Blue staining (upper panels) and autoradiography (lower panels,32P). Dots indicate the positions of Journal of Cell Science • Advance article Fig. 2. HIF-2α is a phosphorylation target of CK1δ in vitro. the recombinant GST tagged HIF-2α proteins. Numbers on the left indicate the corresponding Journal of Cell Science • Advance article molecular weight markers. (Α) Phospho-amino acid analysis following in vitro phosphorylation of GST-HIF-2α (366542) by CK1δ. 32 P-labelled HIF-2α (366-542) was subjected to hydrolysis, two-dimensional TLC and autoradiography (right panel). The phospho-amino acid standards, phosphoserine Journal of Cell Science • Advance article Fig. 3. HIF-2α Ser383 and Thr528 are candidate CK1δ phosphorylation sites (P-S), phosphothreonine (P-T), and phosphotyrosine (P-Y), were marked after exposure of TLC plate to ninhydrin staining solution (left panel). Circles indicate origin of migration; Arrows indicate direction of migration. (B) Western blot analysis of recombinant full length GST-HIF-2α or fragments incubated with or without 100 units of recombinant CK1δ in the presence of cold ATP, using anti-GST antibody. In the presence of CK1δ slower migrating bands are observed, indicating quantitative HIF-2α phosphorylation. (C) Fragmentation spectrum of a doubly-charged peak at m/z 794.34735 produced a near complete b and y ion series (in blue and red) together with prominent neutral loss ions (in green). The corresponding peptide is identified as GAVS(p)EKS(p)NFLFTK confirming that both Serine 4 and Serine 7 are phosphorylated. Tables explaining in detail the different fragments found in the MS2 CID spectrum can be found in the supplementary material. (D) Schematic representation of the peptide sequence identified as phosphopeptide by mass spectrometry in C and the conserved amino acid sequence from 518 to 542 aligned to corresponding amino Journal of Cell Science • Advance article acid sequences from vertebrate HIF-2α homologues and from human HIF1α. inhibits HIF-2 transcriptional activity (A) Full length GST-HIF-2α (upper panel) or GST-HIF-2α 366-542 (lower panel) wild type or its point mutants (1μg) were subjected to in vitro phosphorylation by 100 units of recombinant CK1δ and analyzed by SDS-PAGE followed by Coomassie Blue staining or autoradiography. Only the relevant parts of the gels are shown. The numbers under each lane represent relative phosphorylation levels measured as indicated under “Experimental Procedures” and are means of three independent experiments. (B) Same as in A, but the substrates were subjected to in vitro phosphorylation assays by 1μg of Huh7 total protein extracts. (C) Western blot analysis of lysates from HepG2 cells transfected with flag-CMV2 alone, wild-type flag-HIF-2α or its mutant forms using anti-flag and anti-tubulin antibodies. Journal of Cell Science • Advance article Fig. 4. Mutation of Ser383 or Thr528 reduces HIF-2α phosphorylation by CK1δ and (D) HIF-2 transcriptional activity determined 24 h after transfection of HepG2 cells with the pGL3–5HRE-VEGF reporter plasmid, the plasmid pCI-Renilla and the pFLAG-CMV2 plasmids carrying the indicated inserts. Values determined as a ratio of firefly luciferase activity over renilla activity are expressed in relation to the results obtained from cells expressing pFLAG-CMV2 alone and represent the mean of three different experiments performed in triplicate (± s.e.m). (E) Determination of HIF-2 transcriptional activity in HepG2 cells transfected as in D in the presence or absence of the specific CK1δ inhibitor Journal of Cell Science • Advance article D4476 (10 μΜ). Results are shown as in D. Fig. 5. Mutation of Ser383 or Thr528 inhibits HIF-2-dependent EPO secretion. (A) Determination of EPO and PAI-1 mRNA levels by quantitative real-time PCR in Huh7 cells transfected with flag-CMV2 alone, wild-type flag-HIF-2α or its phosphorylation and represent the means of three independent experiments performed in duplicate (± s.e.m). (B) Huh7 cells were transfected as in A and EPO secretion was measured by ELISA. Results are shown as in A. (C) Determination of EPO and PAI-1 mRNA levels by quantitative realtime PCR in HepG2 cells transfected as in A. Results are shown as in A. (D) HepG2 cells were transfected as in A and EPO secretion was measured by ELISA. Results are shown as in A. Journal of Cell Science • Advance article deficient mutants. Results are shown as percent of activity in relation to wild-type HIF-2α USF2. (Α) Western blot analysis of total extracts from Huh7 cells incubated for 4 h in normoxic or hypoxic (1% O2) conditions, with or without the CK1δ specific inhibitor D4476 (10 μΜ), using anti-HIF-2α and anti-tubulin antibodies as indicated. Cells were lysed after treatment with CHX (10μg/ml) for the indicated time points. (B) Western blot analysis of total extracts from Huh7 cells incubated as in A. Cells were lysed after re-oxygenation for the indicated time points. (C) Huh7 cells were transfected with flag-USF2 and 24 h post transfection were incubated for 16 h in normoxic or hypoxic (1% O2) conditions, with or without the CK1δ specific inhibitor D4476 (10 μΜ). Cells were lysed and subjected to immunoprecipitation Journal of Cell Science • Advance article Fig. 6. Inhibition of CK1δ does not affect HIF-2α protein stability or its interaction with with anti-flag antibody. Total cell extract (input) and precipitated proteins (IP) were analyzed Journal of Cell Science • Advance article by western blot using anti-USF2 and anti-HIF-2α antibodies as indicated. (A) Transfected HepG2 cells expressing flag-CMV2 alone, wild-type flag-HIF-2α, or its phosphorylation deficient mutants were fixed, stained with DAPI and subjected to immunofluorescence with anti-Flag mouse monoclonal antibody. Cells were treated with 10 ng/ml LMB for 4 h before fixation (as indicated). In one case, cells were treated with (10 μΜ) D4476 for 16 h before fixation (as indicated). White scales bars equal to 10 Μm. (B & C) Journal of Cell Science • Advance article Fig. 7. Phosphorylation of HIF-2α by CK1δ blocks its CRM1-dependent nuclear export. Number of cells showing nuclear or cytoplasmic staining under each condition was counted, and the percentage calculated. At least 100-150 transfected cells were examined in each case. Each experiment depicted in this figure was performed three separate times and data plotted Journal of Cell Science • Advance article are the average of the three independent experiments (± s.e.m). (A) Subcellular fractionation of HeLa cells transfected with CK1δ siRNA (10nM) or scrambled siRNA (10nM). Cells were incubated for 16 h under hypoxia and treated with 10 ng/ml LMB for 4 h before collection and lysis. (B) HIF-2 transcriptional activity determined 24 h after transfection of HepG2 cells with the pGL3-5HRE-VEGF reporter plasmid, the plasmid pCI-Renilla and the pFLAG-CMV2 plasmids carrying the indicated inserts. Cells Journal of Cell Science • Advance article Fig. 8. CK1δ impairs CRM1-dependent nuclear export of HIF-2α under hypoxia. were treated with 10 ng/ml LMB for 4 h before collection and lysis. Values determined as a ratio of firefly luciferase activity over renilla activity are expressed in relation to the results obtained from cells expressing pFLAG-CMV2 alone and represent the mean of three different experiments performed in triplicate (± s.e.m). (C) Under hypoxia, HIF-2α stabilizes and enters the nucleus but can be exported back into the cytoplasm by a CRM1-dependent process. CΚ1δ directly phosphorylates HIF-2α on Ser383 and Thr528, blocks its nuclear export and promotes formation of a heterodimer with ARNT, binding to HRE sequences in the enhancer regions of its target gene EPO and subsequent stimulation of erythropoietin Journal of Cell Science • Advance article expression and production. J. Cell Sci. 129: doi:10.1242/jcs.191395: Supplementary information B + + siRNA CK1δ - + C ** mRNA levels % Hypoxia HIF-2α CK1δ Actin Hypoxia D siRNA CK1δ - + CK1δ Actin 1st exp 2nd exp - siCK1δ G - + + D4476 - + - + H * Hypoxia D4476 * * + - + + EPO + - + + PAI-1 Hypoxia + + D4476 - + Fig. S1. Impaired CK1δ activity decreases HIF-2 target genes mRNA and EPO secretion levels. (A) Western blot analysis of lysates from HepG2 cells transfected with CK1δ siRNA (10nM) or scrambled siRNA (10nM). Cells were incubated for 16 h under hypoxia before collection and lysis. (B) Determination of PAI-1 mRNA levels by quantitative real-time PCR in HepG2 cells incubated and treated as in A. Results are shown as percent of activity in relation to hypoxia and represent the means of three independent experiments performed in duplicate (± s.e.m). (C) Determination of EPO secretion by ELISA in HepG2 cells incubated and treated as in A. Results are shown as in B. (D) Western blot analysis of lysates from Huh7 cells transfected with a different set of CK1δ siRNA (10nM) or scrambled siRNA (10nM). Cells were incubated for 16 h under hypoxia before collection and lysis. (E) Determination of EPO mRNA levels by quantitative real-time PCR in Huh7 cells incubated and treated as in (D). Results are shown as percent of activity in relation to hypoxia induced EPO and represent the means of two independent experiments performed in duplicate (± s.e.m). (F) Western blot analysis of lysates from HepG2 cells incubated for 16 h under normoxia or hypoxia (1% O2), in the presence or absence of the specific CK1δ inhibitor D4476 (10 μΜ) hypoxia before collection and lysis. (G) Determination of EPO and PAI-1 mRNA levels by quantitative real-time PCR in HepG2 cells incubated and treated as in A. Results are shown as fold increase in relation to the corresponding normoxic conditions and represent the mean of three independent experiments performed in duplicate (± s.e.m). (H) Determination of EPO secretion by ELISA in HepG2 cells incubated and treated as in A. Values are expressed as mlU/ml protein and represent the mean of three independent experiments performed in duplicate (± s.e.m). Journal of Cell Science • Supplementary information - + EPO mIU/mL Hypoxia mRNA Fold increase F Actin + % EPO mRNA levels + HIF-2α CK1δ + + - E - siCK1δ HIF-2α * Hypoxia + + + - siRNA CK1δ EPO secretion levels % SUPPLEMENTARY MATERIAL Fig. S1 A J. Cell Sci. 129: doi:10.1242/jcs.191395: Supplementary information A HIF-2α 366-542 WT S383A S386A C FLAG HIF-2α Tubulin WT 51 100 S383A S386A n.s WT S386A EPO WT S386A PAI-1 Fig. S2. Mutation of HIF-2α Ser 386 to Ala does not affect HIF-2α phosphorylation and activity. (A) GST-HIF-2α 366-542 wild type or its point mutants (1μg) were subjected to in vitro phosphorylation by 100 units of recombinant CK1δ and analyzed by SDS-PAGE followed by Coomassie Blue staining (upper panel) or autoradiography (lower panel). Only the relevant parts of the gels are shown. The numbers under each lane represent relative phosphorylation levels measured as indicated under “Experimental Procedures” and are means of three independent experiments. (B) Western blot analysis of lysates from Huh7 cells transfected with flag-CMV2 alone, wild-type flag-HIF-2α or its phosphorylation deficient mutants using anti-HIF-2α and antitubulin antibodies. (C) Determination of EPO and PAI-1 mRNA levels by quantitative real-time PCR in Huh7 cells transfected as in B. Results are shown as percent of activity in relation to wild-type HIF-2α and represent the mean of three independent experiments performed in duplicate (± s.e.m). Journal of Cell Science • Supplementary information 100 B mRNA levels % n.s J. Cell Sci. 129: doi:10.1242/jcs.191395: Supplementary information -LMB +LMB HIF-2α WT HIF-2α WT +D4476 HIF-2α S383A HIF-2α T528A DAPI Merge Flag-HIF2α DAPI Merge Fig. S3. HIF-2α phosphorylation deficient mutant localization in Huh7 cells. Transfected Huh7 cells expressing flag-CMV2 alone, wild-type flag-HIF-2α, or its phosphorylation deficient mutants (as indicated) were fixed, stained with DAPI and subjected to immunofluorescence microscopy with anti-Flag mouse monoclonal antibody. Cells were treated with 10 ng/ml LMB for 4 h before fixation (as indicated). In one case, cells were treated with 10 μM D4476 for 16 h before fixation (as indicated). White scale bars equal to 10 μm. Journal of Cell Science • Supplementary information Flag-HIF2α J. Cell Sci. 129: doi:10.1242/jcs.191395: Supplementary information A -LMB +LMB FLAG HIF-2α WT HIF-2α S383D HIF-2α T528E HIF-2α DM SD/TE Flag-HIF2α B DAPI Merge Flag-HIF2α DAPI Merge HIF-2α WT + - - - + - - - HIF-2α WT + - - - + - - - HIF-2α S383D - + - - - + - - HIF-2α S383D - + - - - + - - HIF-2α T528E - - + - - - + - HIF-2α T528E - - + - - - + - HIF-2α DM SD/TE - - - + - - - + HIF-2α DM SD/TE - - - + - - - + LMB Nuclear Cytoplasmic E EPO mRNA levels D -LMB + FLAG + HIF2 WT - + DM SA/TA - + DM SD/TE +LMB Cytoplasmic PGK1 mRNA levels Nuclear LMB + FLAG + HIF2 WT - + DM SA/TA - + DM SD/TE Fig. S4. HIF-2α phosphomimetic mutations does not affect its localization. (A) Transfected HepG2 cells expressing flag-CMV2 alone, wild-type flag-HIF-2α, or its phosphomimetic mutants (as indicated) were fixed, stained with DAPI and subjected to immunofluorescence microscopy with anti-Flag mouse monoclonal antibody. Cells were treated with 10 ng/ml LMB for 4 h before fixation (as indicated). In one case, cells were treated with 10 μM D4476 for 16 h before fixation (as indicated). White scale bars equal to 10 μm. (B & C) Number of cells showing nuclear and cytoplasmic staining under each condition was counted, and the percentage calculated. At least 100–150 transfected cells were examined in each case. Each experiment depicted in this figure was performed three separate times and data plotted are the average of the three independent experiments (± s.e.m). (D & E) Determination of EPO (D) and PGK1 (E) mRNA levels by quantitative real-time PCR in Huh7 cells transfected as indicated. Cells were treated with 10 ng/ml LMB for 16 h before lysis. Results are shown as fold increase in relation to the empty FLAG vector and represent the means of two independent experiments performed in duplicate (± s.e.m). Journal of Cell Science • Supplementary information % transfected cells % transfected cells C J. Cell Sci. 129: doi:10.1242/jcs.191395: Supplementary information Table S1. Annotation of the different fragment ions derived from the CID spectrum of the monoisotopic m/z: 794.34735 Da (identified as GAVSpEKSpNFLFTK). (A) b and y ions (B) Ions with phosphorylation losses and (C) Precursor ions. #1 b⁺ b²⁺ C y⁺ y²⁺ #2 1 58.02875 29.51801 G 2 129.06587 65.03657 A 1530.66416 765.83572 12 3 228.13429 114.57078 V 1459.62704 730.31716 11 4 395.13265 198.06996 1360.55862 680.78295 10 5 524.17525 262.59126 SPhospho E 1193.56026 597.28377 9 6 652.27022 326.63875 K 1064.51766 532.76247 8 7 819.26858 410.13793 936.42269 468.71498 7 8 933.31151 467.15939 SPhospho N 769.42433 385.21580 6 9 1080.37993 540.69360 F 655.38140 328.19434 5 10 1193.46400 597.23564 L 508.31298 254.66013 4 11 1340.53242 670.76985 F 395.22891 198.11809 3 12 1441.58010 721.29369 T 248.16049 124.58388 2 K 147.11281 74.06004 1 13 B Seq. #1 b⁺-P b⁺-2P 13 b²⁺-P b²⁺-2P Seq. y⁺-P y⁺-2P y²⁺-P y²⁺-2P A 1432.68727 1334.71037 716.84727 667.85882 12 V 1361.65015 1263.67325 681.32871 632.34026 11 1262.58173 1164.60483 631.79450 582.80605 10 1 G 2 3 #2 13 4 297.15575 149.08151 5 426.19835 213.60281 SPhospho E 1095.58337 548.29532 9 6 554.29332 277.65030 K 966.54077 483.77402 8 7 721.29168 623.31479 361.14948 312.16103 838.44580 419.72654 7 8 835.33461 737.35772 418.17095 369.18250 SPhospho N 9 982.40303 884.42614 491.70516 442.71671 F 5 11 1242.55552 1144.57863 621.78140 572.79295 F 3 12 1343.60320 1245.62631 672.30524 623.31679 T 2 [M + 2H]²⁺+H 794.85037 [M + 2H]²⁺-H 793.84254 [M + 2H]²⁺-P-H₂O-NH₃ 727.83945 [M + 2H]²⁺-2P-H₂O-NH₃ 678.85100 [M + 2H]²⁺-P-NH₃ 736.84473 [M + 2H]²⁺-2P-NH₃ 687.85629 [M + 2H]²⁺-P-H₂O 736.35273 [M + 2H]²⁺-2P-H₂O 687.36428 [M + 2H]²⁺-P 745.35801 [M + 2H]²⁺-2P 696.36956 [M + 2H]²⁺-H₂O-NH₃ 776.82790 [M + 2H]²⁺-NH₃ 785.83318 [M + 2H]²⁺-H₂O 785.34117 [M + 2H]²⁺ 794.34645 6 Journal of Cell Science • Supplementary information A J. Cell Sci. 129: doi:10.1242/jcs.191395: Supplementary information Table S2. Sequences of HIF-2α specific peptides identified by the LC-MS/MS analysis and their corresponding scorings. phosphoRS Site Probabilities XCorr Charge MH+ [Da] ΔM [ppm] 4.62 3 4007.94504 1.29 4.34 2 1587.68743 1.13 SNFLFTK 2.10 2 856.45616 -0.23 AILPPSQPWATELR 1.92 2 1578.86540 0.93 1.80 2 1041.49175 -0.42 Sequence Modifications LKEEPEELAQLAPTPGDAIIS LDFGNQNFEESSAYGK GAVsEKsNFLFTK S4, S7 S(4): 100.0; S(7): 100.0; T(12): 0.0 LFAmDTEAK M4(Oxidation) Journal of Cell Science • Supplementary information (Phospho) J. Cell Sci. 129: doi:10.1242/jcs.191395: Supplementary information Table S3. Schematic representation of human HIF-2α PAS-B domain. Amino acid sequence of the HIF-2α region 236 to 262 aligned with the corresponding amino acid Journal of Cell Science • Supplementary information sequence from human HIF-1α.