* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download HMW glutenin subunits in multiploid Aegilops species: composition
Vectors in gene therapy wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Genomic imprinting wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Multilocus sequence typing wikipedia , lookup
Molecular cloning wikipedia , lookup
Genetic code wikipedia , lookup
Molecular Inversion Probe wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Biosynthesis wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
SNP genotyping wikipedia , lookup
Non-coding DNA wikipedia , lookup
Genomic library wikipedia , lookup
Point mutation wikipedia , lookup
Molecular ecology wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Molecular evolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
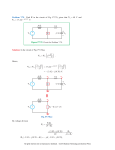
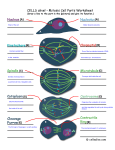
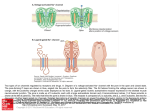
NOTES HMW glutenin subunits in multiploid Aegilops species: composition analysis and molecular cloning of coding sequences XIE Ruili, WAN Yongfang, ZHANG Yan & WANG Daowen Institute of Genetics, Chinese Academy of Sciences, Beijing 100101, China Correspondence should be addressed to Wang Daowen (e-mail: [email protected]) Abstract The Aegilops genus contains species closely related to wheat. In common with wheat, Aegilops species accumulate high molecular weight (HMW) glutenin subunits in their endospermic tissue. In this study, we investigated the composition of HMW glutenin subunits in four multiploid Aegilops species using SDS-PAGE analysis. Furthermore, by working with Ae. ventricosa, we established an efficient genomic PCR condition for simultaneous amplification of DNA sequences coding for either x- or y-type HMW glutenin subunits from polyploid Aegilops species. Using the genomic PCR condition, we amplified and subsequently cloned two DNA fragments that may code for HMW glutenin subunits in Ae. ventricosa. Based on an analysis of the deduced amino acid sequences, we concluded that the two cloned sequences encode one xand one y-type of HMW glutenin subunit, respectively. Keywords: Aegilops, Ae. ventricosa, HMW glutenin subunit, molecular cloning. The seed storage proteins of common wheat are composed of three categories of polypeptides: high molecular weight (HMW) glutenin subunits, low molecular weight (LMW) glutenin subunits and gliadins. HMW glutenin subunits make up 10% of the total protein content in the mature wheat grain, its composition and function can have a profound influence on the baking quality of wheat flour[1 3]. In hexaploid wheat (genome AABBDD), three loci (Glu-1A, Glu-1B, Glu-1D), located on the long arm of the homeologous group one chromosomes 1A, 1B and 1D, respectively, contain genes encoding HMW glutenin subunits. Within each locus, there are two closely linked genes, one coding for a higher molecular weight xtype subunit and the other for a lower molecular weight ytype subunit. Owing to gene silencing, in the Glu-1A locus, the y-type subunit gene is not expressed in hexaploid wheat. The remaining five genes are all active and each of them possesses multiple alleles (including null). As a consequence, hexaploid wheat varieties usually express four to five HMW glutenin subunits, and often differ in their composition of HMW glutenin subunits. Chinese Science Bulletin Vol. 46 No. 4 February 2001 Through genetic analysis, it was demonstrated in 1981 that the composition of HMW glutenin subunits in a given common wheat variety could affect the baking quality of its flour[4]. Recent biotechnological investigations on transgenic expression of the genes encoding the functionally superior HMW subunit (such as the 1Dx5 subunit specified by the Glu-1D locus of the D genome) have not only supported the conclusion drawn through genetic analysis, but also provided direct examples for improving wheat flour baking quality by changing HMW glutenin subunit compositions[5 7]. At present, the number of functionally superior HMW subunits identified in common wheat is very limited. This necessitates the search for more and better subunits from alternative genetic resources. The Aegilops genus is closely related to the Triticum genus, to which common wheat belongs. Many Aegilops species possess either a D genome (Ae. tauschii, DD) or a D genome component (multiploid Aegilops species)[8]. Cytogenetic analysis has shown that the D genome component present in common wheat and in those multiploid Aegilops species is interrelated and may be originally derived from a common ancestor (Ae. tauschii). This evolutionary relatedness points to the possibility that Aegilops species may contain genes coding for polypeptides structurally similar among the HMW glutenin subunits of common wheat. This supposition has been confirmed by SDS-PAGE analysis of Aegilops seed storage proteins and by the cloning of a gene from Ae. tauschii encoding a polypeptide (1Dty subunit), which shows a high level of amino acid sequence similarity to the 1Dy subunits specified by the Glu-1D locus[9 11]. In contrast to the studies carried for Ae. tauschii, little is known about the composition of HMW glutenin subunits and the genes encoding these polypeptides in polyploid Aegilops species (such as Ae. crassa, Ae. juvenalis, Ae. vavilovii and Ae. ventricosa). The primary structure (i.e. amino acid sequence) of thus far published HMW glutenin subunits is very conserved, the amino acid sequence in both the N and C terminus is highly similar among the different subunits[12]. The middle region of the primary sequence is named the repetitive domain because this region consists of highly repeated short amino acid sequence motifs. Different HMW glutenin subunits usually differ in the length of the repetitive domain, the type of the short amino acid sequence motifs contained, and the number of repeats for individual motifs. In the x-type subunit, the repetitive motifs are mainly of three types: GQQ, PGQGQQ and GYYPTSP/SQQ. The presence of the GQQ motif is unique to the x-type subunits. In the y-type subunit, the repetitive motifs are also of three types: PGQGQQ, GYYPTSLQQ and GHYPASLQQ. As a result of the conservation in terminal amino acid sequences, the nucleotide sequences in the 5 and 3 ends of the DNA region coding for HMW glutenin subunit are also highly homologous. This high level of nucleotide sequence homology 309 NOTES facilitated the design of PCR primers for the amplification of HMW glutenin subunit coding sequences[13 16]. However, this type of PCR analysis has so far limited to only a few species of wheat and Ae. tauschii. Furthermore, there is currently no report on simultaneous amplification of the coding sequences for both x- and y-type subunits in single PCR reactions. We investigated the composition of HMW glutenin subunits in four multiploid Aegilops species, and established an optimized PCR condition for simultaneous amplification of the coding sequences for both x- and y-type subunits of Ae. ventricosa in single PCR reactions. Our results contributed to the understanding of HMW glutenin subunits and their genes in polyploid Aegilops species. This knowledge may be an aid to the exploitation of Aegilops HMW glutenin subunits in wheat quality improvement in the future. 1 Materials and methods ( ) Plant materials. The seeds of the four polyploid species, Ae. crassa (DDD2D2McrMcr), Ae. juvenalis (DDMjMjUU), Ae. vavilovii (DDMcrMcrSpSp) and Ae. ventricosa (DDMvMv), were provided by the Institute of Crop Germplasm Resources of the Chinese Academy of Agricultural Sciences. Common wheat materials possessing known HMW glutenin subunits were obtained from the Long Ashton Research Station in UK. ( ) SDS-PAGE analysis. The endosperm proteins were extracted from single seeds for SDS-PAGE analysis according to the published method[17] . For each Aegilops species, two accessions were analyzed. For each accession, MW glutenin subunit composition was determined for three seeds in independent experiments. Gel electrophoresis results were recorded using a digital camera. ( ) Genomic DNA extraction and genomic PCR reactions. DNA extraction from the seedlings of Ae. ventricosa was performed as previously described[18]. Two pairs of PCR primers were designed according to the published nucleotide sequences of HMW glutenin genes (fig. 1). P1 (5 -ATGGCTAAGCGGC/TTA/GGTCCTCTTTG-3 ) and P2 (5 -CTATCACTGGCTG/AGCCGACAATGCG-3 ) were based on the conservation of terminal nucleotide sequences in the coding region of HMW glutenin genes and for amplifying the complete coding region of the gene encoding either x- or y-type subunits. P3 (5 ATCACCCACAACACCGAGCA-3 ) and P4 (5 -AGCTGCAGAGAGTTCTATCA-3 ) were based on the conservation of two stretches of nucleotide sequences located, respectively, in the 5 and 3 untranslated region of HMW glutenin genes and for amplifying the DNA fragment containing both the coding region and the untranslated regions. For both primer pairs (P1/P2, P3/P4), two genomic PCR reactions were carried out. Except for the DNA template and the nucleotide primers, the other com- 310 ponents (polymerase, nucleotides and buffer) required by the PCR reaction were provided by either the kit (Taq DNA polymerase PCR kit) from the Gibco BRL Company or the kit (Advantage-GC Genomic PCR kit) from the Clontech Company. PCR reactions using the Taq DNA polymerase PCR kit in conjunction with P1/P2, P3/P4, respectively, were conducted in 50 µL volume with 300 ng genomic DNA, 0.2 mmol/L of each nucleotide and 20 µmol/L of each primer and 5 units of Taq polymerase. The cycling parameters were: 95 for 5 min, followed by 34 cycles of 95 for 1 min, 55 for 1 min and 72 for 3 min, and a final extension at 72 for 10 min. PCR reactions using the Advantage-GC Genomic PCR kit in conjunction with P1/P2, P3/P4, respectively, were also conducted in 50 µL volume with 300 ng genomic DNA, 0.2 mmol/L of each nucleotide and 10 µmol/L of each primer and 2.5 units of Taq polymerase. The cycling parameters were: 94 for 1 min, followed by 34 cycles of 94 for 0.5 min, 68 for 3 min, and a final extension at 68 for 3 min. The PCR products were analyzed in 1% ethidium bromide (EB) containing agarose gels. The results were recorded using the Bio-Rad Gel Doc-1000 scanner. Fig. 1. A schematic representation of the positions of the four nucleotide primers (P1, P2, P3 and P4) in relation to the coding region of an HMW glutenin gene. ATG and TAG denote the start and stop codon of the coding region, respectively. The TATA box of the promoter region is also shown. The diagram is not drawn to scale. ( ) Southern hybridization. To examine the presence of DNA sequences bearing homology to HMW glutenin genes in the PCR reactions, PCR products, after agarose gel electrophoresis, were vacuum-blotted onto Hybond N+ membrane (Amersham Pharmacia Biotech). The resultant membrane was hybridized with a radioactive probe, prepared using the Prime-A-gene ® kit (Promega) and the plasmid DNA containing the coding sequence of an x-type HMW glutenin subunit from Ae. tauschii (AtDx2.5, Wan Yongfang and Wang Daowen, unpublished result). The protocol of Southern hybridization and the optimization of post hybridization washing conditions were based on the published procedures[19]. The hybridization results were recorded using a phosphor-imager (Molecular Dynamics). ( ) Cloning of PCR products and nucleotide sequence analysis. The desired PCR products were recovered from agarose gels, then cloned into the pGEM-T vector (Promega). Nucleotide sequence of positive clones was determined by a commercial company (TaKaRa). For nucleotide sequence comparison, the programs (such as Chinese Science Bulletin Vol. 46 No. 4 February 2001 NOTES Fig. 2. SDS-PAGE analysis of the composition of HMW glutenin subunits in the four polyploid Aegilops species. Lanes 1 and 2, Ae. crassa (three subunits detected); lanes 3 and 4, Ae. juvenalis (four subunits detected); lane 5, common wheat (cv. Chinese Spring, four subunits detected); lane 6, common wheat (MG7249, five subunits detected,); lanes 7 and 8, Ae. ventricosa (four subunits detected, the arrow indicated subunits possessing an electrophoretic mobility slower than that of the 1Dx2.2 subunit); lanes 9 and 10, Ae. vavilovii (five subunits detected). Fig. 3. Analysis of PCR products by agarose gel electrophoresis (a) and Southern hybridization (b). Lane 1, DNA markers (kb); lane 2, empty track; lane 3, PCR products obtained with the primer pair P1/P2 and the Taq DNA polymerase PCR kit; lane 4, PCR products obtained with the primer pair P3/P4 and the Taq DNA polymerase PCR kit (* denotes non-specific amplification products); lane 5, PCR products obtained with the primer pair P1/P2 and the Advantage-GC genomic PCR kit; lane 6, PCR products obtained with the primer pair P3/P4 and the Advantage-GC genomic PCR kit. Long arrows indicate the fragments with a size more than 1.8 kb, the short arrow indicates the 1.0 kb product. ORF Finder, Blast, etc.) in the NCBI network service were used. 2 Results ( ) SDS-PAGE analysis. Fig. 2 shows the result of SDS-PAGE analysis of HMW glutenin subunits in four polyploid Aegilops species. Several preliminary conclu) Three HMW glutesions could be drawn from fig. 2: nin subunits were expressed in the hexaploid Ae. crassa species; ) in Ae. juvenalis, four subunits were detected for each of the two accessions. However, the two accessions differed in the composition of the subunits they expressed. The electrophoretic mobility of the Jx1 subunit was faster than that of 1Dx2.2, whereas that of the Jx2 subunit was similar to what was displayed by the 1Dx2.2 subunit. Considerable difference in electrophoretic mobil) In ity also existed between the Jy1 and Jy2 subunits. Ae. ventricosa, four subunits were detected with largest one possessing an electrophoretic mobility similar to that of the 1Dx2.2 subunit. ) In Ae. vavilovii, five subunits were detected. The findings described above demonstrated the following points: ) Except for Ae. ventricosa, there was no correlation between the number of subunits expressed and the ploidy level of the genome. For example, in theory, the hexaploid species Ae. juvenalis should express six subunits. However, only four subunits were detected in the SDS-PAGE analysis. ) Different accessions of the same Chinese Science Bulletin Vol. 46 No. 4 February 2001 species may differ in their composition of HMW glutenin subunits. This was clearly shown by the two accessions of Ae. juvenalis. ) In two of the four species analyzed, the presence of the subunits possessing an electrophoretic mobility similar to, or larger than, that of the 1Dx2.2 subunit was observed. However, it was not known if these subunits were encoded by the D genome component in the relevant Aegilops species. ( ) Genomic PCR reactions. Based on the above SDS-PAGE analysis, we chose Ae. ventricosa as a model species to establish an optimized PCR condition that may permit simultaneous amplification of the coding sequences for both x- and y-type subunits. Among the four different PCR conditions tested, the reactions performed with the Advantage-GC genomic PCR kit were generally better than those with the Taq DNA polymerase PCR kit. In agarose gel analysis of the PCR products derived from reactions with the Advantage-GC genomic PCR kit, there were more DNA fragments with a size in the range of 1.8 3.0 kb, and the yield of individual fragments was also higher (fig. 3(a), lanes 5 and 6, arrows). In contrast, in agarose gel analysis of PCR products obtained from reactions with the Taq DNA polymerase PCR kit, there were no DNA fragments with a size in the range of 1.8 3.0 kb (fig. 3(a), lanes 3 and 4). The two primer pairs affected the amplification results obtained with the Advantage-GC genomic PCR kit. In the reaction performed with P1/P2, a higher yield was obtained for the DNA fragment with a 311 NOTES Table 1 accession numbera) X12928 X61009 M22208 X13927 g543541 AF216868 X03041 X12929 U39229 AF216869 X61026 Comparison of the deduced, N-terminal amino acid sequences of Aevenx2.5 and Aeveny1.9 subunits to those of published x- and y-type subunits Aevenx2.5 Aeveny1.9 plant, subunit, type wheat, Dx5, x-type wheat, Ax1, x-type wheat, Ax2*, x-type wheat, Bx7, x-type wheat, Bx17, x-type rye, x-type wheat, Dy12, y-type wheat, Dy10, y-type Ae. tauschii, y-type rye, y-type wheat, By9, y-type identity (%) 92 76 72 71 70 67 64 64 63 63 59 accession numbera) X03041 X12929 U39229 AF216869 X61026 M22208 X13927 X61009 g543541 X12928 AF216868 plant, subunit, wheat, Dy12, wheat, Dy10, Ae.tauschii, rye, wheat, By9, wheat, Ax2*, wheat, Bx7, wheat, Bx17, wheat, Ax1, wheat, Dx5, rye, type y-type y-type y-type y-type y-type x-type x-type x-type x-type x-type x-type identity (%) 99 98 98 93 92 64 63 62 62 61 60 a) Except for g543541,which represents EMBO accession number for a protein sequence, the remaining accession numbers all represent nucleotide sequences. higher molecular weight (approximately 2.5 kb) (fig. 3(a), lane 5). In the case of P3/P4, the yield of the lower molecular weight DNA fragment (approximately 1.9 kb) was higher (fig. 3(a), lane 6). Southern hybridization, with the probe prepared the coding sequence for the AtDx2.5 subunit, revealed three cross-hybridizing DNA fragments in the PCR products with a size in the range of 1.0 2.5 kb (fig. 3(b)), confirming that he amplified DNA fragments were indeed related to HMW glutenin genes. ( ) Cloning of PCR products and amino acid sequence analysis. The 2.5 and 1.9 kb PCR fragments were cloned into the pGEM-T vector, yielding plasmids pAeven2.5 and pAeven1.9, respectively. The 5 end sequence of the 2.5 and 1.9 kb inserts was determined. The resultant nucleotide sequences (615 bps for the 2.5 kb fragment, 537 bps for the 1.9 kb fragment) were translated into amino acid sequences using the ORF Finder program. The deduced amino acid sequences were compared with those of published HMW glutenin genes using the Blast program. The deduced N-terminal amino acid sequence (composed of 204 amino acids) of the 2.5 kb fragment showed higher than 90% identity with those of x-type subunit genes, whereas its identity with those of the y-type subunit was comparatively lower (around 70%). This suggests that the subunit specified by the 2.5 kb fragment was an x-type subunit, for which a tentative name Aevenx2.5 was therefore given. A limited analysis of the amino acid sequence in the repetitive domain of the Aevenx2.5 subunit revealed the presence of three types of repeated, short amino acid sequence motifs, namely, GQQ, PGQGQQ and GYYPTS/PQQ. The existence of the repeated tripeptide GQQ in the Aevenx2.5 subunit further confirmed that this subunit was an x-type subunit. The deduced N terminal amino acid sequence (composed of 179 amino acids) of the 1.9 kb fragment exhibited more than 90% identity with those of y-type subunit genes. In contrast, its identity with those of the x-type subunit was less than 70%. This sug- 312 gests that the subunit specified by the 1.9 kb fragment was a y-type subunit, for which a tentative name Aeveny1.9 was subsequently given. A preliminary analysis of the amino acid sequence in the repetitive domain of the Aeveny1.9 subunit showed the presence of two types of repeated, short amino acid sequence motifs, namely, PGQGQQ and GYYPTS/PQQ. The absence of the repeated tripeptide GQQ in the Aeveny1.9 subunit provided additional evidence, supporting that this subunit was a ytype subunit (see table 1). 3 Discussion Many agronomically important genes, such as the mildew resistance genes Pm2, Pm12, Pm13, Pm19 and the fertility restoration gene Rf6, are originally derived from the genome of Aegilops species[20 23]. These Aegilops genes have contributed to the genetic improvement of common wheat varieties. Based on these examples, the potential value of the Aegilops HMW glutenin subunits in wheat quality breeding merits a thorough investigation. Some investigations on the HMW glutenin subunits and their coding genes in Ae. tauschii have been conducted[9, 10]. A preliminary evaluation of the genetic materials derived from a common wheat × Ae. tauschii cross has already shown that some progeny lines possess quality properties superior to those of the common wheat parent[24]. In this study, we analyzed HMW glutenin subunit composition of four multiploid Aegilops species. In common with bread wheat varieties, multiple subunits (at least three) were expressed in the seeds of different Aegilops species. Because it was observed that there was generally no correlation between the number of subunits expressed and the ploidy level of the species investigated, we deduced that null alleles and/or gene silencing might also affect the expression of HMW glutenin subunit genes in Aegilops species. Owing to a limited number of accessions analyzed, our finding on the difference of HMW glutenin Chinese Science Bulletin Vol. 46 No. 4 February 2001 NOTES subunit composition between different accessions of a single species requires further investigation. Although PCR reactions have been employed to study HMW glutenin genes in common wheat and Ae. tauschii materials, successful attempts in simultaneous amplification of the coding sequences for both x- and ytype subunits in single PCR reactions have not been reported by previous investigators. By designing two pairs of PCR primers and testing four different PCR conditions, we found a PCR condition that gave rise to the amplification of both x- and y-type subunit genes in single PCR reactions. There were three different DNA fragments (2.5, 1.9 and 1.0 kb, respectively) in the PCR reactions conducted with the Advantage-GC genomic PCR kit. Southern hybridization indicated that they all possessed nucleotide homology to HMW glutenin genes. DNA cloning and amino acid sequence analysis confirmed that the 2.5 kb fragment represented the coding sequence for an x-type subunit (Aevenx2.5) and the 1.9 kb fragment for a y-type subunit (Aeveny1.9). However, the identity of the 1.0 kb fragment remains to be determined. The PCR reactions performed with the Advantage-GC genomic PCR kit were generally more efficient in terms of product yield. Considering that the Advantage-GC genomic PCR kit was developed especially for genomic PCR reactions using GC-rich templates and that the coding sequence of HMW glutenin genes was generally rich in GC content, it was natural to find that this kit performed better than the Taq DNA polymerase PCR kit in our experiments. Because of the limited sequence information generated in this study, it is not known at this stage how different the two Ae. ventricosa subunits are from the subunits published previously in overall amino acid sequence. Considering that Ae. ventricosa is a tetraploid species with two component genomes (D and MV), further studies are needed to find out to which genome(s) the 2.5 and 1.9 kb fragments belong. The simultaneous amplification of the coding sequences for both Aevenx2.5 and Aeveny1.9 subunits in single PCR reactions demonstrates that the PCR condition established in this study is highly efficient, and will be useful in further molecular genetic studies on HMW glutenin subunit in the multiploid Aegilops species. Acknowledgements This work was supported by a special fund for biotechnology research from the Chinese Academy of Sciences (Grant No. STZ-3-11). The nucleotide sequences for the Aeveny1.9 and Aevenx2.5 subunit have been deposited in the EMBO database with the accession numbers AF226698 and AF226699, respectively. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. 17. 18. 19. 20. 21. 22. References 1. 2. 3. 4. Lawrence, G. J., Shepherd, K. W., Chromosomal location of genes controlling seed proteins in species related to wheat, Theor. Appl. Genet., 1981, 59: 25. Shewry, P. R., Miflin, B. J., Seed storage proteins of economically important cereals, Adv. Cereal. Sci. Technol., 1985, 7: 1. Weegels, P. L., Hamer, R. J., Schofield, J. D., Functional properties of wheat glutenin, J. Cereal. Sci., 1996, 23: 1. Payne, P. I., Corfield, K. G., Holt, L. M., et al., Correlation between the inheritance of certain high-molecular-weight-subunits of glutenin and bread-making quality in progenies of six crosses of breat-wheat, J. Sci. Food Agric., 1981, 32: 51. Chinese Science Bulletin Vol. 46 No. 4 February 2001 23. 24. Blechl, A. E., Anderson, O. D., Expression of a novel highmolecular-weight glutenin subunit gene in transgenic wheat, Nature Biotechnology, 1996, 14: 875. Altpeter, E., Vasil, V., Srivastava, V. et al., Integration and expression of the high-molecular weight-glutenin subunit 1Ax1 gene into wheat, Nature Biotechnology, 1996: 1155. Barro, F., Rooke, L., Békés, F. et al., Transformation of wheat with high molecular weight subunit genes results in improved functional properties, Nature Biotechnology, 1997, 15: 1295. Miller, T. E., Systematics and evolution, in Wheat Breeding: Its Scientific Basis (ed. Lupton, F. G. H.), New York: Chapman and Hall, 1987, 1 49. William, M. D. H. M., Pena, R. J., Mujeeb-Kazi, A., Seed protein and isozyme variations in Triticum tauschii (Aegilops tauschii), Theor. Appl. Genet., 1993, 87: 257. Mackie, A. M., Lagudah, E. S., Sharp, P. J. et al., Molecular and biochemical characterization of HMW glutenin subunits from T. Tauschii and the D genome of hexaploid wheat, J. Cereal. Sci., 1996, 23: 213. Mackie, A. M., Sharp, P. J., Lagudah, E. S., The nucleotide and derived amino acid sequence of an HMW glutenin gene from Triticum tauschii and comparison with those from the D genome of bread wheat, J. Cereal. Sci., 1996, 24: 73. Shewry, P. R., Tatham, A. S., Barro, F. et al., Biotechnology of breadmaking: unraveling and manipulating the multi-protein gluten complex, Bio/Technology, 1995, 13: 1185. D’Ovidio, R., Anderson, O. D., PCR analysis to distinguish between alleles of a member of a multigene family correlated with wheat bread-making quality, Theor. Appl. Genet., 1994, 88: 759. Smith, R. L., Schweder, M. E., Barnett, R. D., Identification of glutenin alleles in wheat and triticale using PCR-generated DNA markers, Crop Sci., 1994, 34: 1373. D’Ovidio, R., Masci, S., Porceddu, E., Development of a set of oligonucleotide primers specific for genes at the Glu-1 complex loci of wheat, Theor. Appl. Genet., 1995, 91: 189. Lafiandra, D., Tucci, G. F., Pavoni, A. et al., PCR analysis of xand y- type genes present at the complex Glu-A1 locus in durum and bread wheat, Theor. Appl. Genet., 1997, 94: 235. Payne, P. I., Holt, I. M., Law, C. N., Structural and genetic analysis on the high-molecular-weight subunits of wheat glutenin, Part 1: Allelic variation in subunits amongst varieties of wheat (Triticum aestivum), Theor. Appl. Genet., 1981, 60: 229. McCouch, S. R., Kochert, G., Yu, Z. H. et al., Molecular mapping of rice chromosomes, Theor. Appl. Genet., 1988, 76: 815. Sambrook, J., Fritsch, E. F., Maniatis, T., Molecular Cloning: A Laboratory Manual, New York: Cold Spring Harbor Laboratory Press, 1989. Lutz, J., Hsam, S. L. K., Limpert, E. et al., Chromosomal location of powdery mildew resistance genes in Triticum aestivum L. (common wheat). 2. Genes Pm2 and Pm19 from Aegilops squarrosa L., Heredity, 1995, 74: 152. Donini, P., Koebner, R. M. D., Ceoloni, C., Cytogenetic and molecular mapping of the weight-Aegilops longissima chromatin breaking points in powdery mildew resistant introgression lines, Theor. Appl. Genet., 1995, 91: 738. Jia, J., Devos, K. M., Chao, S. et al., RFLP-based maps of the homoeologous group-6 chromosomes of wheat and their application in the tagging of Pm12, a powdery mildew resistance gene transferred from Aegilops speltoides to wheat, Theor. Appl. Genet., 1996, 92: 559. Ma, Z. Q., Zhao, Y. H., Sorrells, M. E., Inheritance and chromosomal location of male-fertility restoring gene transferred from Aegilops umbellulata Zhuk to Triticum aestivum L., Mol. Gen. Genet., 1995, 247: 351. Murphy, J. P., Griffey, C. A., Finney, P. L. et al., Agronomic and grain quality evaluations of Triticum aestivum x Aegilops tauschii backcross populations, Crop Sci., 1997, 37: 1960. (Received May 12, 2000) 313