* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Accelerated Evolution of Sex Chromosomes in
Group selection wikipedia , lookup
Genetic drift wikipedia , lookup
Non-coding DNA wikipedia , lookup
Point mutation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Ridge (biology) wikipedia , lookup
Public health genomics wikipedia , lookup
Human genetic variation wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Adaptive evolution in the human genome wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
History of genetic engineering wikipedia , lookup
Minimal genome wikipedia , lookup
Genomic imprinting wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
Microsatellite wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Gene expression programming wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Neocentromere wikipedia , lookup
Genome evolution wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Y chromosome wikipedia , lookup
Koinophilia wikipedia , lookup
Genome (book) wikipedia , lookup
X-inactivation wikipedia , lookup
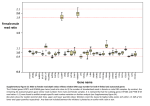
Accelerated Evolution of Sex Chromosomes in Aphids, an X0 System Julie Jaquiéry,1,* Solenn Stoeckel,1 Claude Rispe,1 Lucie Mieuzet,1 Fabrice Legeai,1,2 and Jean-Christophe Simon1 1 Institut National de Recherche Agronomique (INRA), Unité Mixte de Recherche 1099, Biology of Organisms and Populations Applied to Plant Protection, Le Rheu, France 2 INRIA Centre Rennes—Bretagne Atlantique, GenOuest, Rennes, France *Corresponding author: E-mail: [email protected]. Associate editor: John Parsch Abstract Key words: effective population size, faster-X effect, genetic diversity, inheritance, sex chromosome, X chromosome. Introduction Sex chromosomes exhibit several unusual properties such as sex-specific inheritance patterns, hemizygosity, and reduced recombination (Vicoso and Charlesworth 2006). Hence, these chromosomes are expected to differ from autosomes in their response to evolutionary factors (e.g., drift, demography, and selection) and can help elucidate the relative importance of these forces to genome evolution. Several empirical studies outlined lower levels of genetic diversity on the X relative to the autosomes in a wide range of species and tried to disentangle the several possible mechanisms that could account for this difference (e.g., Moriyama and Powell 1996; Begun and Whitley 2000; Sundström et al. 2004; Geraldes et al. 2008; Ellegren 2009; see also table 1). In XY systems, the effective population size of the X is three-fourths that of the autosomes because under 1:1 sex ratio and equal variance in reproductive success between sexes, there are three copies of the X per mating pair but four copies of the autosomes (Wright 1931; Caballero 1994, 1995). Thus, the X is more exposed to genetic drift, leading to an expected X:A diversity ratio of three-fourths. However, this ratio might be further affected by several demo- graphic factors. For example, a higher variance in male reproductive success or female-biased dispersal will increase this ratio above three-fourths (the reverse would decrease it below three-fourths) since the Xs are more often transmitted by females than males compared with the autosomes (Caballero 1995; Charlesworth 2001; Laporte and Charlesworth 2002). Bottlenecks or population expansions alter the X:A diversity ratio, too, because the effective population size determines the rate of convergence to the new diversity equilibrium (Wall et al. 2002; Pool and Nielsen 2007). Additionally, both mutation and recombination rates differ between the X and the autosomes. Given that the X is transmitted more often through females than the autosomes, male-biased mutation (i.e., a higher mutation rate in males because of the large number of germ cells divisions required for spermatogenesis, Miyata et al. 1987; Li et al. 2002) will lower mutation rate on the X and hence X-linked diversity relative to autosomes (Ellegren 2007). All else being equal, recombination is also predicted to be lower on the X because autosomes recombine in both males and females and the X only in females (Hedrick 2007) (but note that recombination is predicted to be higher on the X chromosome in Drosophila, since males do not recombine, © The Author 2011. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution. All rights reserved. For permissions, please e-mail: [email protected] Mol. Biol. Evol. 29(2):837–847. 2012 doi:10.1093/molbev/msr252 Advance Access publication October 13, 2011 837 Research article Sex chromosomes play a role in many important biological processes, including sex determination, genomic conflicts, imprinting, and speciation. In particular, they exhibit several unusual properties such as inheritance pattern, hemizygosity, and reduced recombination, which influence their response to evolutionary factors (e.g., drift, selection, and demography). Here, we examine the evolutionary forces driving X chromosome evolution in aphids, an XO system where females are homozygous (XX) and males are hemizygous (X0) at sex chromosomes. We show by simulations that the unusual mode of transmission of the X chromosome in aphids, coupled with cyclical parthenogenesis, results in similar effective population sizes and predicted levels of genetic diversity for X chromosomes and autosomes under neutral evolution. These results contrast with expectations from standard XX/XY or XX/X0 systems (where the effective population size of the X is threefourths that of autosomes) and have deep consequences for aphid X chromosome evolution. We then localized 52 microsatellite markers on the X and 351 on autosomes. We genotyped 167 individuals with 356 of these loci and found similar levels of allelic richness on the X and on the autosomes, as predicted by our simulations. In contrast, we detected higher dN and dN/dS ratio for X-linked genes compared with autosomal genes, a pattern compatible with either positive or relaxed selection. Given that both types of chromosomes have similar effective population sizes and that the single copy of the X chromosome of male aphids exposes its recessive genes to selection, some degree of positive selection seems to best explain the higher rates of evolution of X-linked genes. Overall, this study highlights the particular relevance of aphids to study the evolutionary factors driving sex chromosomes and genome evolution. MBE Jaquiéry et al. · doi:10.1093/molbev/msr252 Table 1. Factors Influencing X:A Diversity Ratio in Standard XX/XY or XX/X0 Systems and in Aphid-Like Systems. Effect on X:A Diversity Ratio in Standard XX/XY or XX/XO Systems Effect on X:A Diversity Ratio in Aphid-Like Systems (XX/X0) Mechanisms Mechanisms Number of chromosome copies (X vs. A) Yes (2) NeX 5 3/4 NeA No NeX 5 NeA Variance in reproductive success between the sexes Yes (1/2) X is transmitted two-thirds of the time by females; A half of the time No Both X and A are transmitted half of the time by females Sex-biased dispersal Yes (1/2) X is transmitted two-thirds of the time by females; A half of the time Noa Both X and A are transmitted half of the time by females Bottleneck or population expansion Yes (1/2) Ne determines the rate to reach 6 NeA equilibrium 1 NeX ¼ No Ne determines the rate to reach equilibrium 1 NeX 5 NeA Male-biased mutation (i.e., higher mutation rate m in males) Yes (2) X is transmitted two-thirds of the time by females; A half of the time (so mX < mA) No Both X and A are transmitted half of the time by females (so mX 5 mA) Recombination rate r Yes (1/2) If only females recombine (e.g., Drosophila): d Both X and A recombine only in females No In aphids, only females recombine: d Both X and A recombine only in females d Both X and A are transmitted half of the time by females 0rX 5 rA d X is transmitted two-thirds of the time by females; A half of the time 0rX > rA Then selection similarly affects diversity on the X and autosomes through hitchhiking effects because rX 5 rA If both males and females recombine (e.g., mammals): d X recombines in females; autosomes in both sexes d X is transmitted two-thirds of the time by females; A half of the time 0rX < rA Then for both cases mentioned, selection differentially affects linked diversity on the X and autosomes through hitchhiking effects because rX 6¼ rA Positive or background selection on new or standing variation Yes (1/2) The hemizygosity of the X in males exposes X-linked genes to selection more than autosomal genes Yes (1/2) The hemizygosity of the X in males exposes X-linked genes to selection more than autosomal genes NOTE.—Signs in brackets show the direction of the effect. a A marginal effect (þ or ) can be expected under sex-biased dispersal combined to an extreme success of reproduction of migrant individuals. Connallon 2007). Recombination in turn determines how positive (Maynard-Smith and Haigh 1974) and background selection (Charlesworth et al. 1993) impact on linked genes. Furthermore, X-linked recessive alleles are exposed to selection in the heterogametic sex, whereas autosomal recessive alleles are not exposed until they reach frequencies large enough to allow them to appear as homozygote. Therefore, selection is predicted to be more efficient on genes on the X (Charlesworth et al. 1987; Vicoso and Charlesworth 2006), and this will in turn increase or decrease the X:A diversity ratio, depending on whether selection acts on standing variation or new mutations (Charlesworth et al. 1987; Orr and 838 Betancourt 2001; Betancourt et al. 2004). However, it is important to stress that the various evolutionary factors discussed here do interact (Ellegren 2009). Hence, disentangling the evolutionary causes (e.g., drift, demography, and selection) of unequal diversity levels between the X and the autosomes in XX/XY systems remains challenging. In this context, organisms with alternative modes of inheritance of sex chromosomes can be valuable models to better understand forces driving sex chromosome evolution. For instance, the X chromosome of aphids shares some characteristics with the X of species with XX/XY sex-determining Faster-X Evolution in Aphids · doi:10.1093/molbev/msr252 MBE FIG. 1. Annual life cycle of the pea aphid and ploidy levels for autosomes (A) and sexual chromosome (X). Overwintering eggs, diploid for both types of chromosomes (AA and XX), give birth to an asexual female. After 10–20 generations of apomictic parthenogenesis, asexual females produce (by parthenogenesis) sexual females and males. Males inherit the same autosomal genome as asexual females but receive only one of the female Xs: hence, they are diploid for the autosomes and haploid for the X (represented as AAX0). Ovules (haploid for both the autosomes and the X) are generated by a normal meiosis, but males produce only X-bearing sperm (AX). The fusion of male and female gametes restores the diploid level at both the X and the autosomes and generates asexual females. systems (e.g., the X is present in single copy in male aphids), but its inheritance pattern is different; hence, the forces driving aphids’ X chromosome evolution must differ. Aphids have an XX/X0 sex determination system whereby females carry two X chromosomes and males only one (whereas both sexes have three diploid autosomes) (fig. 1). A major difference between aphids and other XX/XO organisms such as some nematodes (e.g., Caenorhabditis elegans, Pires-DaSilva 2007), insects (e.g., grasshopper, crickets, and cockroaches, Tatsuta et al. 2006; Kaiser and Bachtrog 2010), or molluscs (Thiriot-Quievreux 2003) comes from the peculiar pattern of inheritance of the aphid X chromosome in a life cycle combining sexual and asexual reproduction (see fig. 1). Indeed, the aphid life cycle involves several rounds of apomictic parthenogenesis, followed by a single event of sexual reproduction in autumn (fig. 1). Sexual individuals are produced by apomictic parthenogenesis, the males inheriting randomly only one of the asexual female Xs (Wilson et al. 1997; Caillaud et al. 2002). Since males produce exclusively X-bearing gametes (i.e., haploid for the X and for the autosomes), the fusion of male and female gametes restores the diploidy of the X and autosomes to generate an asexual female (figs. 1 and 2). Therefore, in the asexual progeny produced from sexual reproduction, half of the autosomes and X chromosomes originate from the mother and the other half from the father (fig. 2C). This differs from the pattern of X chromosome inheritance in standard XX/X0 or XX/XY systems whereby female offspring receive one maternal and one paternal X chromosome, but males only inherit the maternal X copy (fig. 2A and B). Though a priori minor, this difference between aphids and other standard systems has deep consequences for the evolutionary trajectory of the aphid X chromosome (table 1). Since both male and sexual female aphids transmit one set of autosomes and one of their X chromosomes to the progeny during sexual reproduction, we predict similar effective population sizes for X chromosome and autosomes (i.e., NeA 5 NeX), and so an X:A diversity ratio equal to one (this hypothesis is validated by simulations in the present study). Hence, bottlenecks or population expansions do not affect differentially the genetic diversities on the X and on the autosomes, since these diversities converge to the new equilibrium at the same rate. In addition, any variance in sex-specific reproductive success will have NeX the same effect on the X as on the autosomes, leaving Ne A unchanged. This result arises because the same proportion (50%) of the X and autosomes in the progeny is paternally inherited, so that both the X and the autosomes are similarly impacted by any sex-specific variance in reproductive success. This contrasts to standard XX/XY or XX/XO systems where autosomes and X are differentially affected because half of the autosomes are paternally inherited in the progeny but only 1/3 of the Xs. A similar argument can be raised to explain why sex-biased dispersal similarly affects X NeX and the autosomes in aphids, also leaving Ne unchanged. A Interestingly, we also expect male-biased mutation to similarly affect X chromosome and autosomes in aphids, since the same proportion (50%) of autosomes and X chromosomes are paternally inherited and thus have experienced the higher mutation rate occurring in males. Additionally, we predict no a priori difference in recombination rate between X and autosomes in aphids. This prediction also differs from that for standard XX/XY systems, where the X is predicted to recombine less than autosomes when males recombine (Ellegren 2009) or more than autosomes in species with male achiasmy (e.g., Drosophila, Connallon 2007). The aphid system resembles that of Drosophila because 839 MBE Jaquiéry et al. · doi:10.1093/molbev/msr252 FIG. 2. Inheritance of the X chromosome in XX/XY, standard XX/X0, and aphid-like XX/X0 sex-determining systems. Whereas males transmit their X chromosome to only half of their progeny (i.e., only to daughters) in XX/XY (A) and standard XX/X0 sex-determining systems (B), male aphids transmit their X to 100% of their progeny (C). there is also no recombination during spermatogenesis (Sloane et al. 2001; Hales et al. 2002). Nevertheless, we predict different X:A recombination rates between aphids and Drosophila. The key point is not to consider the proportion of Xs and autosomes that recombine in the parental generation (two-thirds of the Xs and half of the autosomes recombine in the parents in both aphids and Drosophila) but to consider the proportion of chromosomes actually transmitted to the progeny that have recombined in the parents. Indeed, in Drosophila, two-thirds of the Xs found in the progeny have recombined (against half of the autosomes), resulting in a larger rate of recombination for the X. By contrast, in aphids, the progeny from sexual crosses always inherits one X from the mother and the second from the father since male aphids transmit their X to all their progeny (fig. 2C). Autosomes are also evenly transmitted by mothers and fathers. As a result, 50% of the Xs and 50% of the autosomes found in the progeny have recombined in the parents, leading to a 1:1 X:A expected recombination rate in aphids. Therefore, we expect X-linked and autosomal diversities to be similarly eroded by positive or background selection acting on linked genes. By contrast, a shared feature between aphid X chromosome and the X from standard XX/X0 and XX/XY systems is the hemizygosity of the X in males, which exposes X-linked recessive genes to selection. This in turn affects X:A diversity ratio through the influence of selective sweeps or background selection on linked loci. Note also that the faster-X effect (i.e., an accelerated rate of evolution of Xlinked genes, Charlesworth et al. 1987) due to the fixation of beneficial mutations is predicted to be particularly strong in aphids that combine large effective population NeX sizes and a large Ne ratio compared with other species A (Vicoso and Charlesworth 2009; Mank, Vicoso, et al. 840 2010). Overall, a difference in X:A diversity ratio would be more straightforward to interpret in aphids than in conventional XX/XY systems, since fewer factors affect this ratio (table 1). This study aims to disentangle the different evolutionary factors driving X chromosome evolution in the pea aphid (Acyrthosiphon pisum), whose genome has recently been sequenced (International Aphid Genomics Consortium 2010). To this end, we first verified through simulations that the effective population size of the X chromosome equals that of autosomes in an aphid-like sex determination system. In a second step, we assessed the genetic diversity in these two types of chromosomes using 45 X-linked and 311 autosomal microsatellite loci typed on 167 individuals. Finally, we compared the rate of evolution of genes on the X and on the autosomes by identifying orthologs between A. pisum and three other aphid species. Materials and Methods Assessing X and Autosome Effective Population Sizes in Aphids NeX To compare the Ne : NeX ratio in organisms with an aphidA like XX/X0 system (e.g., aphids and the nematode Strongyloides ratti, Harvey and Viney 2001) and in those with an XX/XY system, we simulated the action of genetic drift on X-linked and autosomal genes in R (R Development Core Team 2010). At cycle 0, two alleles at one X-linked locus and one autosomal locus were randomly assigned to 100 asexual females (AA XX) from a uniform distribution considering 30 possible allelic states. Then, the 10–20 generations of asexual reproduction occurring in spring and summer (fig. 1) were modeled by drawing the number of clones generated by each of the 100 initial female lineage Faster-X Evolution in Aphids · doi:10.1093/molbev/msr252 from a geometric distribution with mean 5 0.02 (with this value, each initial lineage produced 48.7 ± 49.1 [mean ± standard deviation] asexual females that reached the last generation before the production of sexual females and males). Each of these newly generated asexual females then produced on average (Poisson distribution) five sexual females and five males parthenogenetically (males randomly lost one of their X chromosome). Finally, offspring (asexual females) were generated by randomly drawing with replacement one female gamete and one male gamete produced by sexual individuals (see figs. 1 and 2). We repeated this step until 100 offspring were generated, to start a new cycle. Simulations were run for ten cycles (a cycle corresponds to several rounds of asexual reproduction followed by one event of sexual reproduction, e.g., fig. 1), and 50,000 independent replicates were performed. Expected heterozygosities (Nei 1987) were computed for the autosomal and the X-linked loci at every cycle t after sexual reproduction (HA,t and HX,t) and were then averaged over the 50,000 replicates. Effective population sizes (Nei) for both the autosomal (NeA) and the X-linked (NeX) loci were estimated from the rate of loss of genetic diversity during the last cyHi;t cle as Nei 5 2ðHi;t H . In our simulations, fecundities were i;tþ1 Þ chosen within the lower tail of values observed in aphids to ensure conservative results. By contrast, the number of allelic states and the size of the population do not influence NeA:NeX ratio and were thus chosen arbitrarily. Since we estimated NeA and NeX from the rate of loss of diversity at one locus within one cycle in a small population (i.e., 100 individuals), we expect a large variance for this estimate because of the stochastic process of inheritance of alleles. We therefore performed an important number of replicates (i.e., 50,000) to obtain an accurate estimation of mean NeA and NeX (alternatively, we could have estimated Ne over a larger number of cycles, but this would have led to similar results). We ran analogous simulations to compute effective population sizes for X-linked and autosomal loci in a standard XX/XO or XX/XY system (i.e., where males transmit their X chromosome only to their daughters). In that case, the adult population consisted of 50 males and 50 females, and 100 offspring (50 males and 50 females) were randomly generated by drawing with replacement one female gamete (AX) and one spermatozoid (either AX to generate females or AY to generate males) from the pool of gametes generated by the adults. Comparison of Genetic Diversities on the X versus Autosomes in Pea Aphid Populations DNA extraction was carried out as in Peccoud et al. (2008) on 167 individuals from nine European pea aphid populations collected on three different host plants (supplementary table S1, Supplementary Material online). Several independent extractions of genetically identical individuals (clones) from each of the 167 lineages were required to obtain enough DNA to genotype these 167 individuals at 403 loci (for primer sequences, see supplementary table MBE S2, Supplementary Material online). Three hundred and seventeen of the microsatellite loci were located within EST contigs or at an average distance of 1.4 kb from the closest predicted gene (International Aphid Genomics Consortium 2010) (hereafter referred to as loci in ‘‘coding region’’). The 86 remaining loci were located at more than 30 kb from any predicted genes (average distance to the closest gene is 47 kb) (hereafter referred to as loci in ‘‘noncoding region’’). Polymerase chain reaction (PCR) amplification and genotyping for loci AlA09M, AlB07M, AlB12M, ApF08M, ApH10M, and AlB08M were performed as in Peccoud et al. (2008). The 397 remaining loci were amplified individually in 10 ll PCR. The mix included approximately 20 ng of DNA, 0.4 U of Taq, PCR buffer at 1 final concentration, 2 mM of MgCl2 (all products from Promega), 0.2 mM of each dNTP, 0.4 lM of forward and reverse primers, and 0.36 lM of a universal primer labeled with either 6-FAM, VIC, NED, or PET fluorescent dyes (DyeGTGCTGCAACATTTTGCTG) (Schuelke 2000). Amplifications were performed on Tetrad PTC-225 (MJ Research) or C1000 (Biorad) Thermo Cyclers. Conditions of the PCR amplification are as follows: 95 °C (5 min), then 20 cycles at 95 °C (1 min)/58 °C (1 min)/72 °C (1 min 30 s), followed by 20 cycles 95 °C (1 min)/53 °C (1 min)/72 °C (1 min 30 s), and a final extension at 72 °C for 10 min. Two microliters of PCR product from four loci with nonoverlapping amplicon sizes and each labeled with a different dye were added to 150 ll H2O. Two microliters of this solution were then added to 5 ll of formamide and 0.05 ll of 500 Liz DNA Ladder (Applied Biosystems). Samples were run on an ABI 3730 or ABI 3130 and we determined allelic scores with Genemapper 3.7 (Applied Biosystems). Allelic richness was estimated for each of the nine populations assuming a minimal sample size of six individuals with FSTAT 2.9.4 (Goudet 2005) on the 356 loci (of 403) that successfully amplified in most of the 167 individuals (see supplementary table S2, Supplementary Material online). The chromosomal localization of each microsatellite marker was performed as in Wilson et al. (1997) by looking at the pattern of inheritance of alleles in 6–28 families made up of an asexual mother and 2 of her sons (we used the families generated by Frantz et al. 2006). Briefly, a heterozygote female will produce heterozygote sons at autosomal loci but her sons will appear homozygote for X-linked loci since males inherit only one of their mother Xs. The effects of the chromosome (X chromosome vs. autosomes) and type of loci (within coding regions or not) on allelic richness were assessed in R (R Development Core Team 2010) using the following linear mixed-effect model (R package Lme4, Bates and Sarkar 2007) fitted with REML (REstricted Maximum Likelihood): Allelic richness ; Number of Repeats þ Motif Size þ Chromosome þ Type of Loci þ Population þ Host Plant þ Loci. Population, Host Plant, and Loci were set as random factors, accounting for the structure of the data. The other effects, namely the Number of Repeats (measured on the genome sequence of the pea aphid, International Aphid Genomics Consortium 2010), Motif Size (di-, tri-, or tetranucleotide), Chromosome, 841 MBE 1 0 −1 −2 Residuals for allelic richness 2 3 Jaquiéry et al. · doi:10.1093/molbev/msr252 Autosomes X Within non−coding area Autosomes X Within coding area FIG. 3. Effects of chromosome type (X or autosomes) and locus position (within coding areas or not) on allelic richness estimated as residuals from the partial mixed-effects model. and Type of Loci were set as fixed effects. The effect of each variable was tested by likelihood ratio tests based on the drop in Akaike’s information criterion following the inclusion of the focal variable into the partial model comprising all other variables. Since Motif Size had no effect of allelic richness (P 5 1), it was removed from the models. To graphically look at the effects of Chromosome (X vs. autosomes) and Type of Loci (within coding regions or not) on allelic richness (fig. 3) once other effects were accounted for, residuals for allelic richness for each locus were extracted from the partial model: Allelic richness ; Number of Repeats þ Population þ Host Plant þ Loci. Linkage disequilibrium between pairs of loci was calculated with FSTAT 2.9.4 (Goudet 2005). We then measured the proportion of pairs showing significant linkage disequilibrium (at P , 0.05) for loci that were both located on 1) the X or 2) autosomes (in that case, we are not able to distinguish whether both loci are located on the same autosome). As a result, even under similar X and autosomes recombination rates, we expect a slightly larger linkage disequilibrium for loci on the X than for those on autosomes. Evolutionary Rates of Genes Located on the X versus Autosomes The 403 microsatellite loci that had been assigned to the X (52 loci) or to the autosomes (351 loci) were then located on the genome sequence of the pea aphid (Acyr 1.0 assembly, International Aphid Genomics Consortium 2010), allowing us to assign 47 scaffolds (genomic DNA fragments) to the X chromosome and 323 to the autosomes. Orthologs between A. pisum genes located on these scaffolds (International Aphid Genomics Consortium 2010) and coding sequences (CDSs) reconstituted from ESTs databases from three aphid species (Acyrthosiphon kondoi: 842 23,046 ESTs, Myzus persicae: 27,728 ESTs, and Aphis gossypii: 88,851 ESTs) were then identified following reciprocal best blast hit criterion (RBH). Translated sequences were aligned using T-coffee (Notredame et al. 2000), nucleic sequences being then aligned using the proteic alignment as a guide. Alignments were trimmed using Gblocks (Castresana 2000; Talavera and Castresana 2007) with default parameter values. We then estimated maximum likelihood pairwise synonymous (dS) and nonsynonymous (dN) evolutionary rates for each gene, using a codon-based model (Codeml from PAML, Yang 1997). For each species comparison, genes located in the upper 5% tail for dS values (when considering X-linked and autosomal genes together) were eliminated as well as those with dN . 0.2. We combined these two thresholds because it allowed eliminating paralogous genes and those containing errors in their EST-based sequence. Overall, 26 genes (of 512) were eliminated for the comparison between A. pisum and A. kondoi (50/ 842 for A. pisum vs. M. persicae and 67/864 for A. pisum vs. A. gossypii, respectively). The eliminated genes represented 5%, 5%, and 7% of autosomal loci and 4%, 15%, and 12% of X-linked loci for comparisons between A. pisum–A. kondoi, A. pisum–M. persicae, and A. pisum–A. gossypii, respectively. Differences between dN (dS and dN/dS, respectively) for X-linked and autosomal genes were tested using two-sided Mann–Whitney tests, independently for each pair of species. A global analysis (overall species) was also performed using the following linear mixed-effects model (fitted with REML): y ; Chromosome þ Gene Expression Level þ Gene þ Species Comparison, where y is dN, dS, or dN/dS. Gene and Species Comparison were set as random factors and Chromosome (X vs. autosomes) as fixed. Gene Expression Level (log[x þ 1] transformed) was included as a fixed-effect covariable and was roughly estimated by counting the ESTs from A. pisum supporting each gene used for dN/dS estimation by using Blastn between the published approximately 204,800 A. pisum ESTs and the 34,603 predicted genes (International Aphid Genomics Consortium 2010). The significance of Chromosome and Gene Expression Level were tested by likelihood ratio tests. Finally, we also tested for a difference in expression levels (based on ESTs counts) between X-linked and autosomal genes using a two-sided Mann–Whitney test. Results Similar X and Autosome Effective Population Sizes in Aphid-Like Systems From our simulations on an aphid-like sex determination system, genetic diversities did not differ between X-linked and autosomal loci after ten cycles (HX,t510 5 0.8566744, HA,t510 5 0.8566305, two-sample t-test: P 5 0.85, n 5 50,000) (see also supplementary fig. S1, Supplementary Material online). Effective population sizes were estimated as NeX NeA 5 50.25 and NeX 5 49.72, providing a ratio of Ne close A to 1 (0.99). By contrast, our simulations of genetic drift in an XX/XY or standard XX/X0 system led to a smaller MBE Faster-X Evolution in Aphids · doi:10.1093/molbev/msr252 Table 2. Allelic Richness Averaged over Nine Populations for X-Linked and Autosomal Microsatellite Loci Located within Coding and Noncoding Genomic Areas. Type of Chromosome Number of microsatellite loci Mean number of repeats (median 6 SD) Mean allelic richness (median 6 SD) Mean allelic richness (corrected for number of microsatellite repeats) (median 6 SD) Within Noncoding Areas Within Coding Areas Autosomes X Chromosome Autosomes X Chromosome 64 14 247 31 11.9 (10 6 4.8) 16.5 (14 6 8.3) 9.3 (9 6 3.1) 9.8 (9 6 3.4) 4.54 (4.59 6 1.43) 5.15 (5.21 6 1.35) 3.30 (3.15 6 1.18) 3.53 (3.52 6 1.06) 0.50 (0.43 6 1.04) 0.15 (0.38 6 1.49) 20.13 (20.25 6 0.86) 20.03 (20.08 6 0.91) NOTE.—SD, standard deviation. genetic diversity on the X than on autosomal loci (twosample t-test: n 5 50,000, P , 1015) and to an X:A ratio of effective population sizes of 0.74, close to the threefourths expectation. Similar Genetic Diversity on the X and Autosomes Mean allelic richness (averaged over populations) reached 3.53 for X-linked and 3.30 for autosomal loci located within coding regions, but allelic richness was slightly higher on the X (5.15) than on the autosomes (4.54) for loci located within noncoding areas. X-linked loci also had a higher number of microsatellite repeats than loci on the autosomes (table 2) and more especially so for loci located within noncoding regions. In contrast, when allelic richness was corrected for the number of microsatellite repeats (since both parameters covary: R2 5 0.43, P , 1015), we observed no difference (P 5 1) of allelic richness between X-linked and autosomal loci (fig. 3). Loci located within coding regions were less diverse (P , 106) than those far from genes (fig. 3). We also showed that 17% of the pairs of loci located on the X chromosome were significantly linked against 12% for pairs on autosomes. These values suggest similar recombination rates on the X and autosomes (for details, see Materials and Methods). Faster Evolutionary Rates for X-Linked Genes In the three pairs of aphid species, we found higher rates of nonsynonymous mutations (dN) for X-linked genes than for autosomal genes (two-sided Mann–Whitney test, P , 0.04 for all comparisons, table 3). Synonymous rate of substitution (dS) did not differ between X-linked and autosomal genes (P . 0.38 in all comparisons). Also, we found higher dN/dS ratio for X-linked genes in all pairs of species, the effect being significant in two pairs of species (P , 0.04) but not in the third one (P 5 0.08) (table 3 and supplementary fig. S2, Supplementary Material online). The global analysis (that accounted for Gene Expression Level, Gene, and Species Comparison) also showed higher dN and dN/ dS ratio for X-linked genes (P 5 0.029 and P 5 0.034) but no difference for dS values (P 5 0.42). We observed particularly high dN/dS values for X-linked genes compared dNX =dSX with autosomes, with dN reaching 1.76 for A. pisum–A. A =dSA kondoi, 1.45 for A. pisum–M. persicae, and 1.34 for A. pisum–A. gossypii. These values decreased with phylogenetic distances between the pair of species compared, A. kondoi being the closest from A. pisum and A. gossypii the more distantly related (Ortiz-Rivas and Martinez-Torres 2010). We also found a significant effect of gene expression levels on dN/dS (P 5 103), dN (P 5 106), and dS (P 5 106), with a negative relation between gene expression and evolutionary rates in all cases. The amount of explained variance was nevertheless small (R2 , 0.005 for dN/dS, and ;0.035 for dN and dS). Finally, we observed slightly lower (but not significantly) EST support for X-linked genes (mean: 19.8) compared with those on the autosomes (mean: 25.6; Mann–Whitney test, n 5 1,188, P 5 0.07). Discussion Effective Population Size Does Not Differ between Aphid X Chromosome and Autosomes Here, we showed by simulations that the aphid X chromosome and autosomes have similar effective population sizes because of the peculiar X chromosome inheritance. This result contrasts with expectations from standard XX/XY or XX/X0 systems, where NeX is only three-fourths of NeA (assuming 1:1 sex ratio and equal variance in reproductive success between sexes) because of the sex-specific inheritance pattern of the X (Wright 1931; Caballero 1994, 1995). Consequently, in aphids, the rate of genetic drift should not differ among the two categories of chromosomes, so that the ratio of X:A diversity should be 1. Interestingly, this result, coupled to the arguments presented before, implies that the ratio of X:A diversity in aphids is not affected by the various demographic (i.e., expansion, bottleneck, sex-biased dispersal, or mating systems) and genetic (recombination and mutation) factors influencing the X:A diversity ratio in XX/XY and standard XX/X0 (fig. 1). Indeed, in such standard systems, several neutral factors such as the number of chromosome copies (e.g., Begun and Whitley 2000; Schofl and Schlotterer 2004; Berset-Brändli et al. 2007), sex-biased dispersal (Laporte and Charlesworth 2002; Ellegren 2009), population demography (e.g., Andolfatto 2001; Schofl and Schlotterer 2004; Handley, Berset-Brandli and Perrin 2006; Baines and Harr 2007), mating system (e.g., 843 MBE Jaquiéry et al. · doi:10.1093/molbev/msr252 Table 3. Number of Genes, Median Values for Rates of Nonsynonymous (dN) and Synonymous (dS) Substitution, and Ratio of dN/dS for the Three Pairs of Species Compared for X-Linked and Autosomal Genes (standard deviation in brackets). Species Pair Comparison Acyrthosiphon pisum– Acyrthosiphon kondoi Acyrthosiphon pisum– Myzus persicae Acyrthosiphon pisum– Aphis gossypii A X P A X P A X P n 463 23 753 39 755 46 dN 0.008 (0.026) 0.014 (0.018) 0.038* 0.020 (0.031) 0.032 (0.035) 0.039* 0.031 (0.034) 0.043 (0.049) 0.032* 0.100 0.091 0.58 0.253 0.271 0.38 0.414 0.396 0.60 dS (0.052) (0.053) (0.087) (0.068) (0.137) (0.131) dN/dS 0.088 (0.104) 0.155 (0.354) 0.040* 0.079 (0.121) 0.115 (0.181) 0.085 0.077 (0.095) 0.102 (0.105) 0.038* NOTE.—P values from two-sided Mann–Whitney tests. *P, 0.050 Baines and Harr 2007; Broquet et al. 2009), and mutation rate (e.g., Handley et al. 2006; Qiu et al. 2010) must be accounted for.SincetheaphidX:Adiversityratioisnotaffectedbyallthese neutral evolutionary factors, aphids are ideal models to investigate the effects of selection on sex chromosome evolution. Similar Levels of Diversity on Aphid X Chromosome and Autosomes We found similar genetic diversity on the X compared with autosomes once the number of microsatellite repeats (that was higher for X-linked loci, and particularly, so for those located within noncoding regions) was accounted for. Such a correction was required since mutation rate, hence genetic diversity, increases with the number of repeats (Ellegren 2000; Xu et al. 2000; Seyfert et al. 2008), an effect also observed in our data where repeat number accounted for 43% of the variance in allelic richness. The larger number of microsatellite repeats observed on X-linked loci corroborates previous evidence that aphid X chromosomes are enriched in heterochromatin and repetitive DNA sequences compared with autosomes (Mandrioli and Borsatti 2007; Monti et al. 2010). The similar diversity on the X and autosomes corresponds to the 1:1 X:A diversity ratio predicted under neutral evolution. However, these results contrast with those of Brisson et al. (2009) who used a smaller data set of noncoding DNA sequences and found that X-linked loci were less diverse than autosomal ones in the pea aphid. Microsatellite markers have been widely used to compare X and autosomes diversities in other taxa (e.g., Payseur et al. 2002; Schofl and Schlotterer 2004), but in aphids, these markers may not be the most appropriate to investigate such patterns because as mentioned above, aphid X chromosomes appear to be enriched in repetitive DNA sequences compared with the rest of the genome. Faster Evolution of X-Located Genes: Relaxed or Positive Selection? We observed higher dN values and dN/dS ratio for X-linked genes, whereas dS, that can be equated to the rate of neutral substitution, did not differ between the X and the autosomes. This suggests similar mutation rates for genes located on these two types of chromosomes. A fast844 er-X effect (i.e., increased dN and dN/dS for X-linked genes) can result from relaxed selection: this is indeed the case when the X has a smaller effective population size than autosomes (e.g., Mank, Nam, et al. 2010), resulting in a reduced effectiveness of selection against slightly deleterious mutations. Alternatively, faster evolutionary rates on the X could result from a higher efficiency of selection (because of the haploid state of the X in males, Vicoso and Charlesworth 2006). As we found here, the similar effective population size between the X and the autosomes in aphids argues against a role of drift and reduced efficiency of selection on the X (Vicoso and Charlesworth 2009). Second, we observed no significant difference in EST supports of A. pisum genes on the X and on the autosomes, suggesting that X-linked genes do not evolve faster because they are less expressed. Gene expression levels also explained less than 0.5% of the variance in dN/dS. Therefore, positive selection seems to best explain the higher evolutionary rates of X-linked genes. However, our measurements of evolutionary rates are only based on pairwise estimates of synonymous and nonsynonymous distances, and few pairwise estimates of the dN/dS ratio were above 1, the usual criteria to conclude that there has been positive selection at the scale of the entire sequence. But such approach (based on pairwise estimates) lacks power to actually detect positive selection (Yang et al. 2005), and a more complete set of taxa and genes would help to better evaluate the potential for positive selection. Ongoing genome sequencing projects in other aphid species will help filling this gap in the near future. Also, positive selection often concerns only specific regions or domains of a gene. With more sequences and taxa available, we will be able to test models of site variation of the rates and to determine if X-linked genes more often contain domains or regions under the influence of positive selection. Interestingly, Vicoso and Charlesworth (2009) showed that the scope for faster-X effect due to the fixation of benNeX eficial mutations increases when Ne exceeds three-fourths, A with faster-X effect occurring for a wider range of dominance values (h) and the effects being stronger (fig. 2 in Vicoso and Charlesworth 2009). Mank, Vicoso, et al. (2010) and Vicoso and Charlesworth (2009) also showed with a theoretical model that the fixation of X-linked MBE Faster-X Evolution in Aphids · doi:10.1093/molbev/msr252 beneficial mutation is predicted to increase with effective population size. From these results, we expect aphid Xlinked genes to show a higher rate of evolution compared NeX with autosomal genes because 1) Ne 51 in aphids and 2) A aphid effective population size is large compared with the effective population sizes of vertebrates. We indeed observed particularly high dN/dS values for X-linked genes compared with autosomes, with X:A dN/dS ratio ranging from 1.34 to 1.76 in the three pairs of aphid species. These values are located within the upper tail of values generally observed (ranging from 0.995 in some Drosophila species to 1.359 in rodents, Begun et al. 2007; Mank, Vicoso, et al. 2010). We located genes on the X and on the autosomes in A. pisum, but we ignore whether their orthologs in A. kondoi, M. persicae, and A. gossypii are also X-linked. This is not necessarily always the case, but the X is thought to be the structurally most conserved chromosome among aphid species (Wilson et al. 2004), possibly because it should be clearly identified for its elimination in males. The hypothesis of high homology of the X between species is further supported because most X-linked microsatellite loci amplify and remain X-linked across distantly aphid related species (Wilson et al. 2004). Yet, any gene shuffling between X and autosomes should reduce the X:A dN/dS ratio and thus be conservative regarding our conclusions. Along this line, we observed that the X:A dN/dS ratio decreases with increasing phylogenetic distances between the pair of species compared, presumably because of increased probability of gene shuffling between the X and the autosomes. Our data on effective population size and expression rate suggest that positive selection (rather than relaxed selection) is responsible for increased rate of evolution of Xlinked genes. Yet, our diversity data from X-linked and autosomal microsatellite loci neither support nor refute this hypothesis. Indeed, theoretical models have shown that both increased and reduced levels of diversity can be expected on linked genes on the X relative to autosomes, depending on whether positive selection acts on newly arisen or standing variation (Orr and Betancourt 2001; Betancourt et al. 2004). Furthermore, even under the hypothesis of positive selection on new mutations, we still could expect similar levels of diversity between the X and the autosomes if recombination rate is high and thus prevents hitchhiking effect to act on linked loci. Interestingly and in contrast to our results, Brisson et al. (2009) found lower rate of diversity on the X than on autosomes on a data set based on noncoding DNA sequences obtained on the pea aphid, giving some support to the hypothesis of positive selection on newly arisen mutations. Resolving the issue of the nature of selection driving X chromosome evolution may be achieved in the future when more extensive genomic data sets from multiple aphid individuals and species will be available. Then comparing within species patterns of polymorphism with patterns of divergence among species (fixed substitutions)—using for example McDonald and Kreitman tests (McDonald and Kreitman 1991)—could provide key insights into the dominant selective forces that influence X chromosome and autosomes. Conclusion This study highlights the relevance of organisms with peculiar modes of inheritance of sex chromosomes, such as aphids, as complementary models to disentangle the various evolutionary forces that drive sex chromosome evolution. Indeed, we have shown here that aphids are particularly well-suited for covering such an issue because 1) clear predictions for X:A diversity ratio can be made under neutral evolution because no factor (apart from selection) should influence this ratio and 2) the similar effective population sizes of the X and autosomes allows to sort out causes for faster-X evolution. Our study therefore highlights the particular relevance of the aphid system to compare the different evolutionary factors that drive sex chromosome and genome evolution in general. Supplementary Material Supplementary tables S1 and S2 and figures S1 and S2 are available at Molecular Biology and Evolution online (http:// www.mbe.oxfordjournals.org/). Acknowledgments We thank L. Buechi, A. Juilland, G. Evanno, Y. Outreman, the Jaquiéry, and Evanno families for help with sampling, J. Bonhomme for rearing aphids, N. Bernard, A. Bonvoisin, F. Mahéo, and N. Prunier-Leterme for help with lab work, and M. Ollivier and J.-P. Gauthier for help with statistical analyses. L. Handley, N. Perrin, D. Tagu, Associate Editor J. Parsch, and two anonymous referees contributed constructive comments to a previous draft of this manuscript. This work was supported by INRA-AIP BioRessources, the Département INRA Santé des Plantes et Environnement, the Genoscope, the Fondation pour la Recherche sur la Biodiversité, the ANR GW-Aphid, and the Swiss National Science Foundation (grant number PBLAA-122658 to J.J.). References Andolfatto P. 2001. Contrasting patterns of X-linked and autosomal nucleotide variation in Drosophila melanogaster and Drosophila simulans. Mol Biol Evol. 18:279–290. Baines JF, Harr B. 2007. Reduced X-linked diversity in derived populations of house mice. Genetics 175:1911–1921. Bates D, Sarkar D. 2007. lme4: linear mixed-effects models using S4 classes. R package version 0.999375-28. Available from: http:// lme4.r-forge.r-project.org/ Begun DJ, Holloway AK, Stevens K, et al. (13 co-authors). 2007. Population genomics: whole-genome analysis of polymorphism and divergence in Drosophila simulans. PLoS Biol. 5:2534–2559. Begun DJ, Whitley P. 2000. Reduced X-linked nucleotide polymorphism in Drosophila simulans. Proc Natl Acad Sci U S A. 97:5960–5965. Berset-Brändli L, Jaquiéry J, Perrin N. 2007. Recombination is suppressed and variability reduced in a nascent Y chromosome. J Evol Biol. 20:1182–1188. Betancourt AJ, Kim Y, Orr HA. 2004. A pseudohitchhiking model of X vs. autosomal diversity. Genetics 168:2261–2269. Brisson JA, Nuzhdin SV, Stern DL. 2009. Similar patterns of linkage disequilibrium and nucleotide diversity in native and introduced 845 Jaquiéry et al. · doi:10.1093/molbev/msr252 populations of the pea aphid, Acyrthosiphon pisum. BMC Genet. 10:22. Broquet T, Jaquiéry J, Perrin N. 2009. Opportunity for sexual selection and effective population size in the lek-breeding European treefrog (Hyla arborea). Evolution 63:674–683. Caballero A. 1994. Developments in the prediction of effective population-size. Heredity 73:657–679. Caballero A. 1995. On the effective size of populations with separate sexes, with particular reference to sex-linked genes. Genetics 139:1007–1011. Caillaud MC, Boutin M, Braendle C, Simon JC. 2002. A sex-linked locus controls wing polymorphism in males of the pea aphid, Acyrthosiphon pisum (Harris). Heredity 89:346–352. Castresana J. 2000. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 17:540–552. Charlesworth B. 2001. The effect of life-history and mode of inheritance on neutral genetic variability. Genet Res. 77:153–166. Charlesworth B, Coyne JA, Barton NH. 1987. The relative rates of evolution of sex-chromosomes and autosomes. Am Nat. 130: 113–146. Charlesworth B, Morgan MT, Charlesworth D. 1993. The effect of deleterious mutations on neutral molecular variation. Genetics 134:1289–1303. Connallon T. 2007. Adaptive protein evolution of X-linked and autosomal genes in Drosophila: implications for faster-X hypotheses. Mol Biol Evol. 24:2566–2572. Ellegren H. 2000. Microsatellite mutations in the germline: implications for evolutionary inference. Trends Genet. 16: 551–558. Ellegren H. 2007. Characteristics, causes and evolutionary consequences of male-biased mutation. Proc R Soc B Biol Sci. 274: 1–10. Ellegren H. 2009. The different levels of genetic diversity in sex chromosomes and autosomes. Trends Genet. 25:278–284. Frantz A, Plantegenest M, Simon JC. 2006. Temporal habitat variability and the maintenance of sex in host populations of the pea aphid. Proc R Soc B Biol Sci. 273:2887–2891. Geraldes A, Basset P, Gibson B, Smith KL, Harr B, Yu HT, Bulatova N, Ziv Y, Nachman MW. 2008. Inferring the history of speciation in house mice from autosomal, X-linked, Y-linked and mitochondrial genes. Mol Ecol. 17:5349–5363. Goudet J. 2005. FSTAT, a program to estimate and test gene diversities and fixation indices. Version 294. Lausanne (Switzerland): University of Lausanne, Department of Ecology and Evolution. Hales DF, Wilson ACC, Sloane MA, Simon JC, Le Gallic JF, Sunnucks P. 2002. Lack of detectable genetic recombination on the X chromosome during the parthenogenetic production of female and male aphids. Genet Res. 79:203–209. Handley LJL, Berset-Brandli L, Perrin N. 2006. Disentangling reasons for low Y chromosome variation in the greater white-toothed shrew (Crocidura russula). Genetics 173:935–942. Harvey SC, Viney ME. 2001. Sex determination in the parasitic nematode Strongyloides ratti. Genetics 158:1527–1533. Hedrick PW. 2007. Sex: differences in mutation, recombination, selection, gene flow, and genetic drift. Evolution 61:2750–2771. International Aphid Genomics Consortium. 2010. Genome sequence of the pea aphid Acyrthosiphon pisum. PLoS Biol. 8:e1000313. Kaiser VB, Bachtrog D. 2010. Evolution of sex chromosomes in insects. Annu Rev Genet. 44:91–112. Laporte V, Charlesworth B. 2002. Effective population size and population subdivision in demographically structured populations. Genetics 162:501–519. Li WH, Yi SJ, Makova K. 2002. Male-driven evolution. Curr Opin Genet Dev. 12:650–656. 846 MBE Mandrioli M, Borsatti F. 2007. Analysis of heterochromatic epigenetic markers in the holocentric chromosomes of the aphid Acyrthosiphon pisum. Chromosome Res. 15:1015–1022. Mank JE, Nam K, Ellegren H. 2010. Faster-Z evolution is predominantly due to genetic drift. Mol Biol Evol. 27:661–670. Mank JE, Vicoso B, Berlin S, Charlesworth B. 2010. Effective population size and the faster-X effect: empirical results and their interpretation. Evolution 64:663–674. Maynard-Smith J, Haigh J. 1974. Hitch-hiking effect of a favorable gene. Genet Res. 23:23–35. McDonald JH, Kreitman M. 1991. Adaptive protein evolution at the ADH locus in Drosophila. Nature 351:652–654. Miyata T, Hayashida H, Kuma K, Mitsuyasu K, Yasunaga T. 1987. Male-driven molecular evolution—a model and nucleotidesequence analysis. Cold Spring Harb Symp Quant Biol. 52:863–867. Monti V, Manicardi GC, Mandrioli M. 2010. Distribution and molecular composition of heterochromatin in the holocentric chromosomes of the aphid Rhopalosiphum padi (Hemiptera: Aphididae). Genetica 138:1077–1084. Moriyama EN, Powell JR. 1996. Intraspecific nuclear DNA variation in Drosophila. Mol Biol Evol. 13:261–277. Nei M. 1987. Molecular evolutionary genetics. New York: Columbia University Press. Notredame C, Higgins DG, Heringa J. 2000. T-Coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol. 302:205–217. Orr HA, Betancourt AJ. 2001. Haldane’s sieve and adaptation from the standing genetic variation. Genetics 157:875–884. Ortiz-Rivas B, Martinez-Torres D. 2010. Combination of molecular data support the existence of three main lineages in the phylogeny of aphids (Hemiptera: Aphididae) and the basal position of the subfamily Lachninae. Mol Phylogenet Evol. 55: 305–317. Payseur BA, Cutter AD, Nachman MW. 2002. Searching for evidence of positive selection in the human genome using patterns of microsatellite variability. Mol Biol Evol. 19:1143–1153. Peccoud J, Figueroa CC, Silva AX, Ramirez CC, Mieuzet L, Bonhomme J, Stoeckel S, Plantegenest M, Simon JC. 2008. Host range expansion of an introduced insect pest through multiple colonizations of specialized clones. Mol Ecol. 17:4608–4618. Pires-DaSilva A. 2007. Evolution of the control of sexual identity in nematodes. Semin Cell Dev Biol. 18:362–370. Pool JE, Nielsen R. 2007. Population size changes reshape genomic patterns of diversity. Evolution 61:3001–3006. Qiu S, Bergero R, Forrest A, Kaiser VB, Charlesworth D. 2010. Nucleotide diversity in Silene latifolia autosomal and sex-linked genes. Proc R Soc B Biol Sci. 277:3283–3290. R Development Core Team. 2010. R: a language and environment for statistical computing. Vienna (Austria): R Foundation for Statistical Computing. Schofl G, Schlotterer C. 2004. Patterns of microsatellite variability among X chromosomes and autosomes indicate a high frequency of beneficial mutations in non-African D. simulans. Mol Biol Evol. 21:1384–1390. Schuelke M. 2000. An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol. 18:233–234. Seyfert AL, Cristescu MEA, Frisse L, Schaack S, Thomas WK, Lynch M. 2008. The rate and spectrum of microsatellite mutation in Caenorhabditis elegans and Daphnia pulex. Genetics 178:2113–2121. Sloane MA, Sunnucks P, Wilson ACC, Hales DF. 2001. Microsatellite isolation, linkage group identification and determination of recombination frequency in the peach-potato aphid, Myzus persicae (Sulzer) (Hemiptera: Aphididae). Genet Res. 77:251–260. Sundström H, Webster MT, Ellegren H. 2004. Reduced variation on the chicken Z chromosome. Genetics 167:377–385. Faster-X Evolution in Aphids · doi:10.1093/molbev/msr252 Talavera G, Castresana J. 2007. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol. 56:564–577. Tatsuta H, Hoshizaki S, Bugrov AG, Warchalowska-Sliwa E, Tatsuki S, Akimoto SI. 2006. Origin of chromosomal rearrangement: phylogenetic relationship between X0/XX and XY/XX chromosomal races in the brachypterous grasshopper Podisma sapporensis (Orthoptera: Acrididae). Ann Entomol Soc Am. 99:457–462. Thiriot-Quievreux C. 2003. Advances in chromosomal studies of gastropod molluscs. J Molluscan Stud. 69:187–202. Vicoso B, Charlesworth B. 2006. Evolution on the X chromosome: unusual patterns and processes. Nat Rev Genet. 7:645–653. Vicoso B, Charlesworth B. 2009. Effective population size and the faster-X effect: an extended model. Evolution 63:2413–2426. Wall JD, Andolfatto P, Przeworski M. 2002. Testing models of selection and demography in Drosophila simulans. Genetics 162:203–216. MBE Wilson ACC, Massonnet B, Simon JC, et al. (11 co-authors). 2004. Cross-species amplification of microsatellite loci in aphids: assessment and application. Mol Ecol Notes. 4:104–109. Wilson ACC, Sunnucks P, Hales DF. 1997. Random loss of X chromosome at male determination in an aphid, Sitobion near fragariae, detected using an X-linked polymorphic microsatellite marker. Genet Res. 69:233–236. Wright S. 1931. Evolution in Mendelian populations. Genetics 16:97–159. Xu X, Peng M, Fang Z, Xu XP. 2000. The direction of microsatellite mutations is dependent upon allele length. Nat Genet. 24:396–399. Yang ZH. 1997. PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci. 13:555–556. Yang ZH, Wong WSW, Nielsen R. 2005. Bayes empirical Bayes inference of amino acid sites under positive selection. Mol Biol Evol. 22:1107–1118. 847