* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Evolutionary origin and consequences of uniparental mitochondrial
Deoxyribozyme wikipedia , lookup
DNA barcoding wikipedia , lookup
Transgenerational epigenetic inheritance wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Genome (book) wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Genomic library wikipedia , lookup
Human genome wikipedia , lookup
Minimal genome wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Non-coding DNA wikipedia , lookup
History of genetic engineering wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome editing wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Frameshift mutation wikipedia , lookup
Oncogenomics wikipedia , lookup
Genome evolution wikipedia , lookup
Population genetics wikipedia , lookup
Koinophilia wikipedia , lookup
Microevolution wikipedia , lookup
Point mutation wikipedia , lookup
Mitochondrial Eve wikipedia , lookup
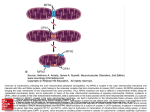
Human Reproduction, Vol. 15, (Suppl. 2), pp. 102-111, 2000 Evolutionary origin and consequences of uniparental mitochondrial inheritance Rolf F.Hoekstra1 Laboratory, of Genetics, Department of Plant Sciences, Wageningen University, Wageningen, the Netherlands ! To whom correspondence should be addressed at: Laboratory of Genetics, Department of Biomolecular Sciences, Wageningen University, Dreijenlaan 2, NL-6703 HA Wageningen, the Netherlands. E-mail: [email protected] In the great majority of sexual organisms, cytoplasmic genomes such as the mitochondrial genome are inherited (almost) exclusively through only one, usually the maternal, parent. This rule probably evolved to minimize the potential spread of selfish cytoplasmic genomic mutations through a species. Maternal inheritance creates an asymmetry between the sexes from which several evolutionary consequences follow. Because natural selection on mitochondria operates only in females, mitochondrial mutations may have more deleterious effects in males than in females. Strictly uniparental inheritance creates asexual mitochondrial lineages that are vulnerable to mutation accumulation (Muller's ratchet). There is evidence that over evolutionary time mitochondrial genomes have indeed accumulated slightly deleterious mutations. Mutation accumulation in animal mitochondrial genomes is probably slowed down mainly by two processes: a severe reduction in germline mitochondrial genome copy number at some point in the life cycle, enabling more effective elimination of mutations by natural selection, and occasional recombination between maternal and paternal mitochondrial genomes following paternal leakage. 102 Key words: evolution/maternal inheritance/ mitochondria/mitochondrial DNA/mtDNA mutations Introduction Because mitochondria cannot be synthesized de novo, these organelles must originate from pre-existing ones and they must be passed on from one generation to the next. A striking and almost universal feature of mitochondrial inheritance is uniparental transmission: in many eukaryotic organisms the offspring from a sexual cross receives mitochondria from only one of the two parents (for a recent review, see Birky, 1995). In animals and flowering plants, it is the maternal parent that transmits the mitochondria. The vast difference in the amount of cytoplasm contributed by the male and female gamete to the zygote almost guarantees effective uniparental inheritance, but active elimination of male-derived mitochondrial DNA from zygotes has been demonstrated (e.g. Kaneda et al., 1995). Uniparental mitochondrial inheritance has also been shown in some species lacking sexual gamete dimorphism, such as the unicellular green alga Chlamydomonas reinhardtii, in which the zygotes are formed by fusions © European Society of Human Reproduction & Embryology Uniparental mitochondria! inheritance between two equally sized gametes that differ in so-called + and - mating type. The mitochondrial DNA derived from the + mating type gamete is degraded in the early zygote (Gillham, 1994). Uniparental inheritance of mitochondrial DNA is in sharp contrast to the familiar biparental inheritance of nuclear DNA. From an evolutionary perspective these two contrasting inheritance patterns have far-reaching consequences. Uniparental inheritance results in asexual mitochondrial DNA lineages, while biparental inheritance creates sexual lineages due to meiotic recombination of paternally and maternally derived DNA. This review will first discuss ideas about the functional importance and evolutionary origin of uniparental mitochondrial inheritance, and then it will concentrate on some of the consequences of the male-female asymmetry in inheritance and of the general lack of recombination between animal mitochondrial genomes. Functional significance of uniparental mitochondrial inheritance Several authors have noted that the mixing of cytoplasm when gametes fuse ought to produce the potential for a rapid spread of deleterious cytoplasmic elements through a sexual population (Grun, 1976; Eberhard, 1980; Cosmides and Tooby, 1981). The argument assumes a lack of precise control of intracellular replication and subsequent segregation of the cytoplasmic DNA at gametogenesis. Such strict control is not known to exist in presentday organisms, and indeed would presumably be difficult to establish for multi-copy genomes whose replication is not synchronized with that of the cell. Without control but with biparental transmission, a mutant germline mitochondrial genome with a replication advantage over the wild-type genome would out-compete the latter and gain transmission to a majority of the gametes. This segregation advantage, if great enough, could outweigh negative effects on the host's fitness and spread through the population. This is a scenario in evolution that would lead to maladaptation instead of adaptation. Natural selection is thus expected to favour genetic variants that minimise the possibilities for spread of such deleterious mitochondrial mutations. One possible mechanism is to restrict the role of transmitting mitochondria to one of the parents. Evolution of uniparental mitochondrial inheritance The above-mentioned argument has formed the basis for mathematical population genetic models showing that uniparental mitochondrial inheritance could have evolved to minimize cytoplasmic mixing and hence to prevent the spread of deleterious mitochondrial mutations (Hoekstra, 1990; Hastings, 1992; Hurst and Hamilton, 1992; Law and Hutson, 1992). The models take as a starting point a population in which mitochondria are inherited from both parents, and then analyse under which conditions a mutation enforcing uniparental mitochondrial inheritance will increase in frequency and eventually become established. The basic argument is as follows. With biparental mitochondrial inheritance, a 'selfish' mitochondrial mutation (i.e. a mutation that is competitively superior in mitochondrial replication but deleterious to the host organism) can be further transmitted by all descendants who have inherited this mutation. In this way the mutation can come to be established in the whole population. In contrast, under uniparental (maternal) mitochondrial inheritance this mutation can only be transmitted further by the daughters who received this mutation from their mother, and again the next generation only by the daughters of these daughters. Thus, the mutation will remain 103 R.F.Hoekstra restricted to a segment of the population formed by individuals descended along the particular female line that originates from the female in which the mutation originally arose. It can never reach individuals descended from other females. Moreover, because the mutation lowers fitness of individual carriers, or hosts, natural selection will act to remove this segment of the population, leading to extinction of the mutant, selfish genome. Uniparental cytoplasmic transmission and male-female dimorphism Mechanisms of uniparental mitochondrial transmission require a fundamental sexual asymmetry in fusing gametes, because it is compulsory that one of them (and not the other) transmits mitochondria to the zygote. This asymmetry is revealed by male-female gamete dimorphism among so-called anisogamous species (characterized by very small male gametes and large female gametes) and by mating-type dimorphism in isogamous species (where all gametes are the same size). Isogamy is seen mainly in some groups of protists and fungi, and is discussed further below. It is possible that sexual dimorphism evolved for reasons other than the regulation of uniparental transmission of cytoplasmic DNA and that mechanisms regulating uniparental transmission were based on pre-existing sexual asymmetry. However, it is also possible that sexual dimorphism evolved primarily to control uniparental genomic transmission. This possibility has been suggested (Hoekstra, 1987) and further analysed (Hurst and Hamilton, 1992). The latter authors make the interesting empirical observation that the taxa that exhibit a fertilization mechanism in which the cytoplasm of the mating partners is not mixed (a group of ciliates among the protists, and the mushrooms among the fungi) also exhibit a lack of sexual dimorphic (male104 female) differentiation. (These species are nevertheless sexual, and there is the advantage that mating can occur with any other individual, rather than with just the 50% of the species with the opposite sex.) Hurst and Hamilton considered this phenomenon to support their hypothesis that the integrity of organellar inheritance is primary in compelling the evolution of sexual dimorphism. However, irrespective of the precise causal connection between the evolution of sexual dimorphism and the evolution of uniparental cytoplasmic inheritance, the asymmetries involved have important evolutionary consequences. Uniparental (maternal) mitochondrial inheritance can be harmful for male fertility A genetic element such as the mitochondrial genome that is transmitted by only one sex is subjected to natural selection forces different to those that affect a genetic element, such as the nuclear genome, which is transmitted by both sexes. Whether a mitochondrial mutation is favoured or not by natural selection will depend only on its effects in females, not on its effects in males. This is caused by the fact that the mutation is only transmitted to a future generation by females, not by males. Because males are a 'dead end' for mitochondria, mitochondrial effects in males have no influence on the fate of particular mitochondrial genotypes. Therefore a mitochondrial mutation with beneficial effects in females but deleterious effects in males will be favoured by selection and is expected to spread through a population. An instructive example of such a situation is provided by the phenomenon of male sterility in plants. Of angiosperm species (the flowering plants), - 5 % exhibit the condition of gynodioecy, or male sterility. At the phenotypic level, male sterility manifests as the coexistence Uniparental mitochondrial inheritance within the population of (normal) hermaphroditic plants and plants that produce only seeds but no viable pollen. Much genetic and population genetic research on this trait has been carried out in natural populations of Plantago and Thymus, as well as in several agricultural crops. At first sight, the presence of appreciable frequencies of plants deficient in their male function is puzzling. These plants can only reproduce as females, and are expected to have a lower fitness than their hermaphroditic conspecifics, which can function both as male and as female. Selection ought to eliminate the genotypes that cause male sterility. The explanation of the conundrum is that, in almost all cases where the genetics of male sterility has been investigated, the male sterile phenotype is caused by a mitochondrial mutation; elimination of the male function does not affect the transmission prospects of mitochondrial genes (as long as there is no overall shortage of pollen in the population). Indeed, a mitochondrial mutation that eliminates the male function and at the same time somewhat enhances fitness of the female hosts will be selectively favoured, because the fate of mitochondrial genes is affected only by their female carriers' fitness. On the other hand, selection acting on nuclear genes that affect the sex ratio will tend to establish an equal investment in both sexes. If the frequency of male steriles in a population rises because selection favours a mitochondrial gene causing male sterility, there will be increasing selection pressure favouring any nuclear mutation that suppresses the cytoplasmic male sterility and restores a more equal sex ratio. Such nuclear restorer genes are indeed found in gynodioecious species. This is a clear example of genomic conflict: selection at the level of mitochondria favouring mitochondrial male sterility in conflict with selection at the individual level favouring nuclear restorers of male fertility. The long-term outcome of such genetic conflict between mitochondrial male sterility genes and nuclear restorer genes is not easy to predict: it depends on the precise relations between the fitness of the different genotypes. There is indirect evidence that at least in some cases the nuclear genome ultimately 'wins' the conflict. It has been observed that malesterile individuals can result from crosses between related species, while within these species no male sterility occurs. This can be interpreted as evidence for complete withinspecies restoration of male fertility, but involving different mitochondrial mutations and nuclear restorers in the two species. In the hybrids, a nuclear restorer from one species can combine with a mitochondrial malesterility gene from the other species against which it is ineffective. The complexity of the evolutionary dynamics of male sterility is illustrated by the finding that natural populations often can simultaneously harbour several different mitochondrial male sterility mutations together with the corresponding sets of nuclear restorer genes. There are some suggestive findings in humans that allow an interpretation along these lines. Leber's hereditary optic neuropathy (LHON) is associated with mitochondrial mutations and appears to affect males more severely than females. Of individuals affected by LHON, -85% are male (Wallace, 1992). A few other examples of mitochondrial mutations that seem to cause more severe disease effects in males than in females have been mentioned (Frank and Hurst, 1996). These authors also point out that reduced sperm motility and poor male fertility have been associated with mitochondrial mutations. Sperm dysfunction could perhaps be caused by mitochondrial mutations that affect female fitness only weakly or not at all, so that such mutations might reach relatively high frequencies at mutation-selection equilibrium. 105 R.F.Hoekstra Uniparental transmission produces asexual mitochondrial lineages Under exclusive uniparental inheritance, mitochondria remain captured in separate female lines of descent and are excluded from recombination with mitochondria derived from a different line of descent. Thus mitochondria form extremely ancient asexual lineages (the animal mitochondrial lineage must be > 1 billion years old). An intriguing question is how non-recombining mitochondria have survived for so long and apparently managed to escape too severe an accumulation of mutations. Asexual reproduction is basically a copying process in which newly copied genomes cannot contain fewer mutations than were present in the parental genome, except in the rare case of back-mutations. When in an asexual population the best class of individuals (containing the lowest number of deleterious mutations) either produces no surviving offspring or only produces offspring that acquired a new deleterious mutation, the population suffers an irreversible increase in the mean number of deleterious mutations per genome. The previously second-best class of individuals will eventually undergo the same fate, and so on. Muller was the first to consider this mutation accumulation consequence of asexual reproduction formally (Muller, 1964), and the process is now known as Muller's ratchet (Felsenstein, 1974). Quantitative description of Muller's ratchet The process of mutation accumulation in asexual lineages (Muller's ratchet) has been analysed mathematically (Haigh, 1978). If, in a population, the number of individuals in the optimal class (containing genomes with the smallest number of deleterious mutations) equals n0, then the following relationship will hold: n0 = N e~u/s, where N is the total 106 population size, U is the expected number of novel deleterious mutations per genome per generation, and s is the average selective disadvantage per mutation. The model (Figure 1) assumes that different mutations in the same genome interact multiplicatively on organismal fitness. Clearly, there is only a real danger that the optimal class will get lost (and the ratchet will click) if « 0 is small. The formula suggests that mutations with very weakly deleterious effects will have the greatest chance of accumulating. When s is small, g-u/s c a n b e v e r y s m a n anc j UQ W[\I stiii be small even in large populations. Strongly deleterious mutations will not accumulate because they are likely to be eliminated quickly by natural selection. The slighter the deleterious nature of asexual lineage mutations, the more likely it is that such mutations become fixed and accumulate. Evidence of mutation accumulation in mitochondrial genomes Several studies, on both mammalian and Drosophila mitochondrial genomes, have reported an excess of non-synonymous nucleotide polymorphisms (i.e. base changes resulting in amino acid changes) between related species compared with the number expected under the assumption of strictly neutral mutations (Ballard and Kreitman, 1994; Nachman et al, 1996; Rand and Kann, 1996). In other words, mitochondrial genomes seem to contain more mutations with (presumably negative) fitness consequences than expected, probably because, with genetic drift, weakly deleterious mutations can reach appreciable frequencies. Lynch has found molecular evidence for the action of Muller's ratchet in animal mitochondrial genomes (Lynch, 1996). He compared transfer RNA genes in mitochondrial and nuclear genomes, and observed that the mitochondrial genome accumulates nucleotide 10 10 8 8 0 0 Figure 1. Muller's ratchet. The population histograms show the relative number of subpopulations with 0, 1,2, 3... etc. deleterious mutations. In the upper histogram, there is a small proportion of the population still with no mutations. If the population with no mutations (n0) is small in absolute numbers, then there is in each generation a chance that, despite high fitness, all n0 individuals die without leaving offspring. If so, the optimal class is lost, the ratchet will have clicked (middle diagram), and the class can be reconstituted only by back mutation (whereas in a sexual population it can be reconstituted by recombination). In the lower diagram the ratchet has clicked a second time (after Maynard Smith, 1989). 1 10 8 0 G i 65 "3" R.F.Hoekstra substitutions -25 times faster than the nuclear genome does. The average binding energy between complementary strands in the stems of mitochondrial tRNA is less than half that of nuclear tRNA, and tRNA loop sizes are 50 times more variable in the mitochondrial than in the nuclear genome (Lynch, 1996): an observation interpreted as meaning that there has been a loss of fitness of mitochondrial tRNAs compared with nuclear tRNAs. He further argues (Lynch, 1996, 1997) that these findings support the idea that animal mitochondrial lineages are subject to slow and very long-term fitness decline due to the accumulation of slightly deleterious mutations not seen in the nuclear genome. The nuclear genome is less likely to accumulate deleterious mutations because of the purging effect of meiotic recombination. This raises the question for how long on an evolutionary time scale animal mitochondrial lineages can survive the inevitable decline in fitness. It has been estimated that the process is not likely to imperil many species on time scales of less than a million years (Lynch and Blanchard, 1998), but nonetheless it is conceivable that mitochondrial mutations accumulation will have contributed to extinction of some animal species. Features of mitochondrial genetics related to the risk of mutation accumulation Deficient DNA protection and repair Unlike the nuclear chromosomes, animal mitochondrial genomes are not protected by histones, spend significant time as single DNA chains during replication (Clayton, 1991; 2000), and are deficient in several DNA repair enzymes, such as those involved in excision repair, photoreactivation and recombination repair (Lansman and Clayton, 1975). Both features will contribute to an elevated mitochondrial mutation rate. 108 Reduction of genome size Some features that reduce mitochondrial mutation accumulation are related to the reduction of the mitochondrial genome size. The ancestors of mitochondria are thought to derive from the alpha-division of the purple photosynthetic bacteria (Yang et al., 1985). Present day animal mitochondria have much smaller genomes than their bacterial relatives. This is mostly the result over evolutionary time of extensive gene transfer (in most taxa some 90% of the genes) from the mitochondrial genome to the nuclear genome. Additionally, the mitochondrial genome is streamlined in several respects (Kurland, 1992). Mammalian mitochondria do not contain introns and have only a single site for the initiation of transcription. Small genomes will carry fewer mutations than larger genomes and the advantage of a single initiation site (apart from saving the space required for separate regulation of each gene) is that a mutation at this site renders the complete mitochondrial genome (instead of just one gene) defective. In this way such a mutation has a more severe deleterious effect and is more easily eliminated by selection. Multiplicity of mitochondrial genomes The multiplicity of mitochondrial genomes in a cell also contributes to a slowing down of mutation accumulation. Because of the multiplicity of mitochondrial genomes per cell and the 'relaxed control' (Birky, 1983; see also Clayton, 2000) of the partitioning of mitochondria among daughter cells, a heteroplasmic cell with a given number of mitochondrial mutations can give rise to progeny cells that carry fewer (or more) mutations. In this way, variation in mutational load among cells would be produced, enabling natural selection to favour cells with a relatively small mutation load. When an individual starts its develop- Uniparental mitochondrial inheritance ment from a homoplasmic condition, this argument only applies to novel mitochondrial mutations arising during development of an individual organism. The effect of intracellular random genetic drift on reducing the rate at which Muller's ratchet clicks has been analysed (Takahata and Slatkin, 1983). They suggest that this effect of random genetic drift could even halt the ratchet, but their simulations are based on very strong selective effects (10% reduction in fitness of individuals fixed for a deleterious mitochondrial mutation). The effect of random drift is expected to be much smaller when mutational effects on fitness are small, which seems more realistic (Lynch, 1997); it is precisely for very weakly deleterious mutations that the danger of mutation accumulation is highest (as discussed above). Bottleneck in mitochondrial genome copy number Genetic drift in heteroplasmic cell lineages is very much enhanced by the occurrence of bottlenecks in the number of mitochondrial genomes in the cell. If in every generation germline mitochondrial genomes are at one point severely reduced to only a few per cell, drift would be maximal and natural selection could potentially be very effective in eliminating inferior mitochondrial genotypes. This requires that at this stage, the mitochondrial genome can be put to the test, i.e. that all mitochondrial functions are activated and that cells containing suboptimal mitochondria are in some way competitively inferior to cells with optimal mitochondria. Bottlenecking could also make natural selection for advantageous mitochondrial mutations more effective. Kearsey et al. have studied the fate of mutations for mitochondrial chloramphenicol resistance in a mouse cell line. The mutations are selectively advantageous in the presence of this antibiotic but disadvantageous in its absence. Despite the continuous presence of chloramphenicol, mutants did not increase in frequency sufficiently to reach homoplasmy (Kearsey et al, 1985). They suggested that a bottleneck in the oocyte could favour evolution towards fixation of advantageous mitochondrial mutations. Paternal leakage A small amount of biparental mitochondrial transmission (with recombination between the maternal and the paternal mitochondrial genomes) could be sufficient to stop Muller's ratchet. Pamilo et al. concluded from a theoretical analysis that the difficulties created by Muller's ratchet can be alleviated with a very low rate of sex-induced recombination (Pamilo et al, 1987). Recent analyses of human mitochondrial sequences suggest that transmission of paternal mitochondrial genomes and subsequent recombination may occasionally (but probably very rarely) occur (Eyre-Walker et al, 1999; Hagelberg et al, 1999). Here we encounter an interesting evolutionary dilemma: selection for a small percentage of biparental transmission would prevent mitochondrial mutation accumulation, but would on the other hand allow an increase in population frequency of selfish mitochondrial mutations, as discussed above. Using mathematical analysis and computer simulation, Hastings has analysed the effects of a low level of paternal leakage on the spread of selfish cytoplasmic mutations (Hastings, 1999). He concluded that even a small amount of paternal leakage can have devastating effects on host population fitness unless it is either exceedingly rare or tight bottlenecking through the female germline reduces the effective population size to nearly unity (a combination of the two mechanisms is allowable). This result emphasizes the essential role of a mitochondrial bottleneck in rapidly exposing a mutation deleterious to 109 R.F.Hoekstra individual host fitness and allowing natural selection to remove affected genomes. Hastings also investigated the effect of paternal leakage in life cycles with a number of asexual divisions between sexual reproduction, as is common among many protists (protozoa, including algae) and fungi (Hastings, 1999). The threat posed by selfish cytoplasmic elements seems to be greatly diminished if sexual reproduction is a rare event in an otherwise asexual life cycle, because during asexual reproduction lineages containing selfish mutations can be eliminated by selection. This could explain the fact that some degree of biparental cytoplasmic transmission is not uncommon among these groups of organisms (yeast being a well-known example). In conclusion, it seems that the effect of natural selection on mammalian mitochondrial transmission operates in a balance, between the Scylla of mutation accumulation and the Charybdis of invasion by selfish mutations, by allowing just a little bit of paternal genomic leakage from a successful fertilizing spermatozoon (with competitive mitochondrially driven motility and hence, by implication, mitochondrial genomic integrity): too little to allow selfish elements to spread, but enough to contribute significantly to slowing down Muller's ratchet. References Ballard, J.W.O. and Kreitman, M. (1994) Unraveling selection in the mitochondrial genome of Drosophila. Genetics, 138, 757-772. Birky, C.W., Jr. (1983) Relaxed cellular controls and organelle heredity. Science, 222, 468^75. Birky, C.W., Jr. (1995) Uniparental inheritance of mitochondrial and chloroplast genes: Mechanisms and evolution. Proc. Natl Acad. Sci. USA, 92, 11331— 11338. Clayton, D.A. (1991) Replication and transcription of vertebrate mitochondrial DNA. Annu. Rev. Cell Biol, 7, 453-478. Clayton, D.A. (2000) Transcription and regulation of 110 the mitochondrial genome. Hum Reprod., 15 (Suppl. 2), 11-17. Cosmides, L.M. and Tooby, J. (1981) Cytoplasmic inheritance and intragenomic conflict. /. Theor. Biol., 89, 83-129. Eberhard, W.G. (1980) Evolutionary consequences of intracellular organelle competition. Q. Rev. Biol., 55, 231-249. Eyre-Walker, A., Smith, N.H. and Maynard Smith, J. (1999) How clonal are human mitochondria? Proc. R. Soc. Lond. B, 266, 477-483. Felsenstein, J. (1974) The evolutionary advantage of recombination. Genetics, 78, 737-756. Frank, S.A. and Hurst, L.D. (1996) Mitochondria and male disease. Nature, 383, 224. Gillham, N.W. (1994) Organelle Genes and Genomes. Oxford University Press, New York, USA. Grun, P. (1976) Cytoplasmic Genetics and Evolution. Columbia University Press, New York, USA. Hagelberg, E., Goldman, N., Lio, P. et al. (1999) Evidence for mitochondrial DNA recombination in a human population of island Melanesia. Proc. R. Soc. Lond. B, 266, 485^92. Haigh, J. (1978) The accumulation of deleterious genes in a population - Muller's ratchet. Theor. Popul. Biol., 14, 251-267. Hastings, I.M. (1992) Population genetic aspects of deleterious cytoplasmic genomes and their effects on the evolution of sexual reproduction. Genet. Res., 59, 215-225. Hastings, I.M. (1999) The cost of sex due to deleterious intracellular parasites. J. Evol. Biol., 12, 177-183. Hoekstra, R.F. (1987) The evolution of sexes. In Stearns, S.C. (ed.) The Evolution of Sex and its Consequences. Birkhaeuser Verlag, Basel, Switzerland. Hoekstra, R.F. (1990) Evolution of uniparental inheritance of cytoplasmic DNA. In Maynard Smith, J. and Vida, J. (eds.) Organisational Constraints on the Dynamics of Evolution. Manchester University Press, Manchester, UK. Hurst, L.D. and Hamilton, W.D. (1992) Cytoplasmic fusion and the nature of sexes. Proc. R. Soc. Lond. B, 247, 189-194. Kaneda, H., Hayashi, J.I., Takahama, S. et al. (1995) Elimination of paternal mitochondrial DNA in intraspecific crosses during early mouse embryogenesis. Proc. Natl Acad. Sci. USA, 92, 4542^546. Kearsey, S.E., Munro, E. and Craig, I.W. (1985) Studies of heterogeneous mitochondrial populations in a mouse cell line: The effects of selection for or against mitochondrial genomes that confer chloramphenicol resistance. Proc. Roy. Soc. Lond. B., 224, 315-324. Kurland, C.G. (1992) Evolution of mitochondrial genomes and the genetic code. Bioassays, 14, 709714. Uniparental mitochondrial inheritance Lansman, R.A. and Clayton, D.A. (1975) Selective nicking of mammalian mitochondrial DNA in vivo: Photosensitization by incorporation of 5-bromodeoxyuredine. J. Mol. Biol, 99, 761-776. Law, R. and Hutson, V. (1992) Intracellular symbionts and the evolution of uniparental inheritance. Proc. R. Soc. Lond. B, 248, 69-77. Lynch, M. (1996) Mutation accumulation in transfer RNAs: molecular evidence for Muller's ratchet in mitochondrial genomes. Mol. Biol. Evoi, 13,209-220. Lynch, M. (1997) Mutation accumulations in nuclear, organelle, and prokaryotic transfer RNA genes. Mol. Biol. EvoL, 14, 914-925. Lynch, M. and Blanchard, J.L. (1998) Deleterious mutation accumulation in organelle genomes. Genetica 102/103, 29-39. Maynard Smith, J. (1989) Evolutionary Genetics. Oxford University Press, Oxford, UK, pp. 241-243. Muller, H.J. (1964) The relation of recombination to mutational advance. Mutat. Res., 1, 2-9. Nachman, M.W., Brown, W.M., Stoneking, M. and Aquadro, C.F. (1996) Nonneutral mitochondrial DNA variation in humans and chimpanzees. Genetics, 142, 953-963. Pamilo, P., Nei, M. and Li, W.-H. (1987) Accumulation of mutations in sexual and asexual populations. Genet. Res., 49, 135-146. Rand, D.M., and Kann, L.M. (1996) Excess amino acid polymorphism in mitochondrial DNA: Contrasts among genes from Drosophila, mice and humans. Mol. Biol. EvoL, 13, 735-748. Takahata, N. and Slatkin, M. (1983) Evolutionary dynamics of extranuclear genes. Genet. Res., 42, 257-265. Wallace, D.C. (1992) Mitochondrial genetics: a paradigm for aging and degenerative disease? Science, 256, 628-632. Yang, D., Oyaizu, Y., Oyaizu, H. et al. (1985) Mitochondrial origins. Proc. Natl Acad. Sci. USA, 82, 4443^447. Ill