* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download (reversed and/or heterotaxic) phenotype in SWV mice
Neuronal ceroid lipofuscinosis wikipedia , lookup
Genome (book) wikipedia , lookup
Genetic engineering wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
X-inactivation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Genomic imprinting wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene therapy wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Gene desert wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Gene nomenclature wikipedia , lookup
Public health genomics wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression programming wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
History of genetic engineering wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
TEKATOLOGY 47:595-602 (19931
Expression of the IV (Reversed and/or Heterotaxic) Phenotype
in SWV Mice
W.M. LAYTON, M.W. LAYTON. MICHAEL BINDER,
DAVID M. KURNIT, ANDRZEJ J HANZLIK,
MARGARET VAN KEUREN, AND FRED G. BIDDLE
Department of Anatomy, Dartmouth Medical School, Hunocw, Neu'
Humpsh,ire iW.M.L., M,W.I,,, M.R.) 03755; Departments of Pediatrics
(A.J.H..D.M.K., M.V.K.) and Human Genetics (D.M.K.I.Howard Hughes
Medicat Institute ('D.M.K., A.J.H., M.V.K.1, Urtiucrsity of Michignri Medical
School, Ann Arbor, Michtgan 48109; and Depurtnrt.nt uf Pcdiutr
University of Calgary, Calgary, Alberta, Canada T2N 4N1 IF.G.B.1
ABSTRACT
Approximately 50% of iuliu mice have situs incersus (mirror
image reversal of viscera) and 40% have heterotaxia (anomalous arrangement
of viscera). The occurrence of heterotaxia is independent of situs. Using the
cross-intercross breeding system to put the iv gene on the SWV background, an
occasional presumed i u l i mouse was found that had a n IV (situs inucrsus and/or
heterotaxic) phenotype. Testcrosses of these reversed animals indicated an iui t
genotype. Since iv is linked tightly t o Igh-C on chromosome 12, we inferred the
genotype with a polymorphism of Igh-C demonstrated using the polymerase
chain reaction (PCR). This confirmed them to be ivi + . The expression of the IV
phenotype in animals heterozygous for the tu gene may be due t o an interaction
of iv with a n autosomal recessive gene found in SWV. We have not found the IV
phenotype in heterozygous iui + mice following placement of the it, gene on six
other inbred strains. Rarely, we also found that presumed SWV + + mice had
the IV phenotype. Test matings of these phenodcviants, corroborated by PCR,
have confirmed them to be + I + . Although the phenotypes of the affected SWV
+ I + and ivl + mice resembled those found in iviiu mice, the occurrence of situs
inversus and heterotaxia were not independent of each other, and most, of the
SWV mice with the IV phenotype had heterotaxia with situs solitus.
This infrequent. dominant expression of the iv gene has so far only been seen
when iv is on the SWV background. These findings are consistent with the idea
that this phenomenon is due to the interaction of the iv gene with another gene
found so far only in the SWV strain.
1993 Wiley-Liss, Inc.
(
The autosomal recessive gene iu (Hummel
and Chapman, '59), which maps to chromosome 12 (Brueckner et al., '89), causes situs
inuersus (mirror image reversal of placement of organs) in mice. One half of iuliu
mice have situs inuersus and approximately
40% have some sort of heterotaxia, a n abnormal arrangement of organs in relation to
each other. Heterotaxia occurs independently of visceral situs in ivliv mice, so that
the frequency of heterotaxia in mice with
situs inversus is equal to its frequency in
those with situs solitus (normal placement
of organs). In mouse strains examined previously, ivl + heterozygotes have demonstrated normal laterality indicating that
Q 1993 WILEY-LISS, INC
~
8
the iu gene is a loss-of-function allele: in the
absence of iu gene activity in the ii)!ii$
mouse, laterality determination does not occur resulting in the random determina t ion
'
of visceral .situs (Layton, '76; Kurnit et al.,
'87).
Heterotaxia in the context used here can
be considered as residual situs inriersus in a
mouse with situs solitu's, or residual s i t u s
solitus in one with situs inr?ersus.Occasionally, thc heterotaxia is so marked that the
predominant situs cannot be determined;
Received January 30, 1992: accepted Januar? 12 1993
596
W.M. LAYTON ET AL.
iv donor
swv
Generation
Cycle
i1
discard
-
+/+
iv/+
iv/iv
iv/iv
112 SOL 1/2 IV
+I+
l2
discard
Fig. 1. Crossintercross mating plan used to place the iu gene on the SWV background.
The solid black circles indicate iuiiu animals with situs inuersus and the hatched circles indicate iuliu animals with situs solitus.
these cases are categorized as situs ambiguus. Most cases of situs ambiguus are in
mice with the situs of the thorax discordant
with that of the abdomen. Heterotaxia occurs in a number of reproducible patterns,
the majority of which have been described
by Hummel and Chapman ('59) and Layton
('78).
During a n attempt to put the iu gene on
various inbred backgrounds using a crossintercross system of mating (Green, '81; Fig.
l ) , we encountered apparent partial dominant expression of the iu gene on the SWV
background. In the cross-intercross mating
system, a G1 (Fl) generation is produced
from the cross iuliu X + I + to yield iul+
heterozygotes. Members of G1 are crossed
among themselves to produce a G2 generation. Reversed G2 mice, presumably reflecting a n iuliu genotype, are then crossed with
+I+ mice to produce G3 iui+ mice for a
second cycle. This process is repeated, usually for 10 cycles (20 generations). The even
numbered generations contain one-quarter
iuliu mice, approximately one half of which
should have situs inuersus. The odd numbered generations are all iul+ and should
have the wild-type phenotype. This expected result was the case with six of the
seven inbred strains; however, for the SWV
strain, a reversed pup (based on location of
the stomach) was found in G7. Since the albino SIV (iuliu) mice were housed in the
same room as the albino SWV, this unexpected finding was attributed to a mating
error and the study was begun again. However, in the repeat study another reversed
pup was found in G3 and still others in succeeding odd-numbered generations, which
we shall call the G-odd effect.
There are two possible explanations for
these findings: 1)One of the parents was a
SIV (iuiiu)mouse instead of SWV (+I + 1, so
t h a t the reversed G-odd mice were actually
iuliu. 2) The iu gene is dominant on the SWV
background, but with reduced penetrance.
IV (REVERSED) PHENOTYPE IN SWV MICE
Breeding tests to infer genotype are difficult
to interpret in these animals because of reduced penetrance of the IV phenotype and
the high rate of lethal congenital heart malformations associated with the IV phenotype (Layton, '78). Recently, we have demonstrated that SWV and SIV are
polymorphic at the Zgh-C locus on chromosome 7 2 (Hanzlik e t al., '90). Since theZgh-C
locus is linked tightly to zu, this has made i t
possible to distinguish if the reversed G-odd
mice are id+ or iuliu. Below are summarized genetic analyses that support the hypothesis that SWV has a n unlinked autosoma1 recessive gene that interacts with iv to
cause expression of the IV phenotype in
iul + heterozygotes with reduced penetrance.
MATERIALS AND METHODS
Non-inbred iuliv mice were obtained originally from Dr. Katherine Hummel of The
Jackson Laboratory. These were kept in a
small closed colony and eventually an inbred strain was produced by brother-sister
mating for 20 generations. This strain is
called SIViLay provisionally (SIV in this paper). Inbred strains AiJ, AKRIJ, BALHlcJ,
CBAiJ, C57BLi6J, and DBAl2J were obtained from The Jackson Laboratory and
SWV from F.G.B. At weaning, all mice were
classified and tagged by ear punch. Newborn pups were examined for the location of
the milk spot, which is the white milk-filled
stomach viewed through the translucent abdominal wall. Those mice with a right-sided
milk spot (and thus situs irzuersus) were
marked by clipping the tip of the tail.
The attempt to inbreed the iv gene on the
SWV background using a cross-intercross
system (Fig. 1)was started three times (experiments 1 , 2 , and 3). Although none of the
experiments went the 20 generations required for inbreeding, all experiments were
consistent with the unanticipated results
reported herein. During experiment 1, the
identification of IV phenotype was based on
the position of the milk spot. During experiment 2, examinations became more thorough. In addition to using the milk spot location to identify the IV phenotype in living
pups, most of the animals were autopsied
excepting those that were too autolyzed or
lost due to maternal cannibalism. These two
methods of identifying affected mice are not
equivalent. The use of the milk spot only
identifies animals with a right-sided stom-
597
ach and thus misses cases that have a leftsided stomach with heterotaxia. However,
because of the selective loss of pups with the
IV phenotype prior to weaning, autopsies
done after the perinatal period miss mice
with the IV phenotype that have died (usually as a result of heart malformations).
Since G5 of experiment 2, all wild-type
SWV ( + i + 1 mice have been autopsied.
In order to test the hypothesis that the iv
phenotype in G-odd iui+ mice is due to interaction of the iu gene with another autosoma1 gene, two sets of experiments were
done:
1. If the reversed G-odd mice were genetically different from the non-reversed G-odd
mice, the incidence of reversal and heterotaxia and their association wit,h each other
in the offspring of rkversed G-odd parents
would be different from that in the offspring
of non-reversed G-odd parents. This experiment was set up prospectively and G-odd
parents of generations G-5 to G-7 were used.
Results of 102 of 805 autopsies of offspring
of non-reversed G-odd parents were not used
in the tabulated results of this experiment
because the parents of these mice, although
not reversed, were found a t autopsy t o have
the iv phenotype.
2. Two backcrosses were set up (see Fig.
3). BC1 should show the G-odd effect since
approximately li4 of its members are heterozygous for iu and homozygous for the putative "hi" gene that is hypothesized t o interact with i u i + t o result in the ii,
phenotype. None of these offspring can be
iuizu. RBCl should not show the G-odd effect
since no animals would be iui + , hi/hi. However, approximately half of them will be
iuliv so that about 114 should be reversed. To
infer the genotype of these mice a t t,he iv
locus, we took advantage of the tight linkage of Igh-C with iu (Brueckner et al., '89;
Hanzlik et al., '90) and of a polymorphism of
Igh-C that enabled us to differentiate SWV
from SIV. This was based on the difference
in the length of a segment ofIgh-C, which in
t u r n was due t o a variable number of tandem dinucleotide repeats of (A,C) and/or
(C,T) in this segment (Hanzlik et al., '90).
This difference was demonstrated using
PCR. The segment from SWV was longer
than that from SIV (Hanzlik et a]..'90). This
"PCR test" first became available during G9
of the third experiment and was used subsequently. Details about the primers and
598
W.M. LAYTON ET AL.
the conditions used for the PCR, as well as
the mapping of iu close to Igh-C ( 1 recombinant out of 201 animals examined) are
given in Hanzlik et al. (’90).
RESULTS
In all three experiments attempting to
place the iu gene onto the SWV background,
we continued to find mice with the IV phenotype in odd numbered generations. Hereafter, we shall call this the “G-odd effect.”
This unexpected finding does not appear to
be the result of a breeding error; testcrosses
of two reversed G5 mice with C57BLl6-ivliu
mice resulted in 4123 and 4/24 reversed
pups, respectively. This is consistent with
the 25%incidence of reversal expected in a n
ivi+ x iuiiv mating and not with the 50%
expected if both G5 animals were iuliu. The
PCR test established the i d + genotype of
16 G9 animals with the IV phenotype from
experiment 3; one animal was homozygous
for the upper (SWV) allele, presumably representing a recombinant between the iu and
Igh-C loci (Fig. 2; Hanzlik et al., ’90).
The incidence of the IV phenotype a s
shown by reversal a t birth and the phenotype at autopsy is given in Table l. The incidence of this G-odd effect was independent
of the sex of the reversed parent and was
found with approximately equal frequency
in males and females. The proportion of
mice with the IV phenotype that had heterotaxia alone was much higher in the
G-odd mice than in ivliv mice as most of the
affected G-odd mice had situs solitus (Table
2). This is in contrast to SWV iuliu mice, in
which situs inversus and situs solitus are
equally frequent in heterotaxic mice. In 424
G1 mice, there were no cases of reversal at
birth and only a single instance of IV phenotype. Note that this finding rules out a n
anomaly a t the iv locus in the SWV mouse,
as the compound heterozygote created in G1
essentially shows no IV phenotype. Assuming that the single case of the IV phenotype
in G1 was a phenodeviant (which is seen in
SWV mice; vide infra), we hypothesized that
the increasing frequency of the G-odd effect
with G-number was the result of interaction
of iv with another recessive gene carried by
SWV which we shall call “hi.”
In a n effort to evaluate the h i gene hypothesis further, we performed three breeding experiments that were consistent with
this hypothesis: 1) We found that there was
a significantly higher frequency of IV phe-
4 2 7 0 bp
4 2 4 0 bp
Fig. 2. Determination of status a t the iu locus using
the closely-linked Zgh-C locus. We demonstrated previously that thelgh-C locus is closely linked to iu and that
the SIViLay (original) genotype could be distinguished
from the SWV genotype at the Zgh-C locus (Hanzlik et
al., ’90).Using the oligonucleotides described in Hanzlik et al. (’901,we amplified via PCR (Saiki et al., ’85)
the intervening region of the lgh-C locus that includes
a n (A,C)- (T,C)-repeat stretch. Following arnplification, the product was electrophoresed through a 2%
Nusieve + 0.54 agarose (Seakem) gel, stained with
ethidium bromide, and photographed. This protocol distinguished between the SWV + and the SIV/Lay (original iu) chromosomes, with the SWV allele being larger
than the SIViLay allele. Using this analysis, we confirmed that a rare reversed SWV mouse indeed had a
SWV + / i genotype (lane 2). Using this analysis, we
also confirmed that three heterozygous reversed F9
mice were id+ (lanes M),as expected from the breeding analysis. Other mice were subjected to this analysis
with the same result (data not shown). This analysis
was consistent with the breeding experiments.
+
notype in the offspring of reversed G-odd
mice than in the offspring of non-reversed
G-odd mice (Table 2, cf. rows labeled “G-Odd
Inverted x G-Odd Inverted” and “G-Odd
Solitus x G-Odd Solitus”). This difference
was consistent with the hypothesis that
such reversed animals were homozygous for
a second (“hi”) gene. 2 ) Offspring of the
backcross BC1 [(SWV x SIV) x SWV] (Fig.
3), which should contain no iuiiv mice but
N
( R E V E R S E D ) PHENOTYPE IN SWV
599
MICE
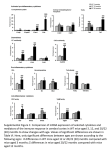
TABLE 1. Incidence of situs inversus at birth and the iv phenotype at autopsy in odd and even gen.erations of
S W V 1+ I+ 1 X SIV livliv) cross-intercross matings (Fig. 1)’
Generation
G3
G5
G7
G9
Odd generations total
G2
G4
G6
G8
E v e n generations total
Sctus inuersus at birth
Number affectedltotal
(R)
Litters
17
36
38
28
119
3/176
101283
241399
101271
4711129
(1.7)
13.5)
(6.0)
(3.7)
(4.2)
25
63
48
46
182
401276
831674
551530
411279
21911759
(14.5)
(12.3)
(10.4)
(14.7)
Phenotype at autopsy
Number affectedkotal
(‘XI
(50)
(17 11
61121
33/193
311339
211226
911879
(9 1)
(9 31
(10 41
(16 8 )
(13 6)
(15 11
111 1)
(13 9)
38!226
791581
731482
441395
234/1684
(11 2)
’These data are from experiments 2 and 3.
TABLE 2. Patterns of N nhenotwe’
Phenotvue of offsurine
Situs inuersus
IV phenotype
IV phenotype
Autopsies
Group
Non inbred-iuliu’
C57BLl6 Lay-iuiiu
SIVLay-iuliu
Total-iuiiu
G-Odd ( G 3 4 9 )
G-Even (G2-G8)
G-Odd INV X G-Odd INV
G-Odd SOL X G-Odd SOL
Backcross (BC1)
Backcross (RBCI)
Heterotaria
IV phenotype
#
#
(91
#
I%)~
#
(%)%
950
842
290
2082
754
1632
94
703
944
258
638
582
206
1426
78
231
24
85
(67.2)
(69.1)
(71.0)
(68.51
484
359
150
20
127
328
355
113
796
73
167
(51.41
(61.01
(54 91
(10.3)
19
64
(2.0)
(24.8)
(75.91
(61.7)
(72 8 )
(69.61
125.6)
155.0)
(58.3)
(48.2)
(21.1)
(14.21
(25.51
(12.1)
993
14
41
4
39
160.9)
17
65
19
40
I55 81
(93 61
(72 3 1
I70 8,
(76 5)
(10001
(62 51
’Results of third experiment only. G-Odd are litters in odd-numbered generations (G-Odd all have an tv + genotype1 G-Even are
litters in even-numbered generations (G-Even are 114 iuku, 1/2 i d + , and 114 f: + genotype).G-Odd INV x G-Odd INV are mice that
are the offspring of parents with situs inversus of G-Odd generations (the parents’ genotype was zu! - I, G-Odd SOL i G-Odd SOL are
mice that were offspring of parents with situs solitus of G-Odd generations (the parents’ genotype was m + I . Backcross IBC11 were
V and 112 + /
genotype. Reciprocal backcross
mice that were the product of (SWV x SIV!Lay) x SWV and thus BC1 are 112 ~ L +
(RBC1)were mice that were the product of ISWV x SIV/Lay) x SIVLay and thus RBCl are 1!2 iu!+ and 1!2 i r h genotype
‘From data in Hummel and Chapman (‘59) and Layton (’761.
3Number reversed a t autopsy as a percentage of number with IV phenotype.
‘Number with heterotaxia as a percentage of number with IV phenotype.
-
114 of which should be ivl + hilhi, showed a
low incidence of IV phenotype (Table 2). It is
noteworthy that all of the affected mice had
heterotaxia and virtually all had situs solitus. I n this respect, they resembled the affected G-odd mice rather than the “conventional” iviiu mice (non-inbred iuliv, C57BLi
6Lay iviiv, and SIViLay iuliu). These two
backcrosses were originally done before the
PCR test was available. Because of the importance of BC1, this was repeated and all
seven of the mice with the IV phenotype
that resulted from this backcross were iui +
as indicated by PCR. 3) Affected offspring of
the reciprocal backcross [(SWV x SIV) x
SIVl; (RBC1 in Table 2 and Fig. 3) resembled affected iviiv mice: when compared
with affected G-odd mice they had a relatively high incidence of situs inversus and
low incidence of heterotaxia (Table 2).
We also encountered a few instances of
the IV phenotype (most frequently heterotaxia with situs solitus) in putative SWV
+i+ mice, We started our SWV colony in
1976; we examined 1,351 newborn pups cursorily and autopsied a relatively small number of weanlings and adults without finding
the IV phenotype. However, since 1983,
when we realized that there might be a n
association between the SWV background
and reversal, we have examined all newborn SWV pups carefully and have done autopsies on virtually all SWV mice. During
this period, we have encountered three in-
600
W.M. LAYTON ET AL.
swv
SIV
+ I + , hilhi
iv/iv,+l+
I
Fl
DISCUSSION
iv/+, hi/+
*iv/+
{
hilhi
backgrounds (AIJ,AKWJ, BALBlcJ, CBAIJ,
C57BL/6J, and DBA/2J), we examined 209
G-odd litters containing 1,179 pups at birth.
None of these pups had the IV phenotype as
demonstrated by a right-sided milk spot.
iv/+
hi/+
{
hi/+
ivliv
i
+/+
Fig. 3. Mating plan for backcrosses BC1 and RBC1.
In BC1, 114 of the mice are hypothesized to be i u l + hiihi
and thus to include the mice with the IV phenotype
(asterisked). However, none of them are iuiiu, so the IV
phenotype cannot be the result of homozygosity for iu.
In RBC1, 1/2 of the resultant animals are iviiu and 1/2
iui + . However, none of them are hilhi, so that none of
the iv/ i- mice would be expected to show the IV phenotype.
stances of situs inversus in 1,587 newborn
SWV pups and six cases of heterotaxia with
situs solitus in 1,250 autopsies. This gives
a n 0.3% incidence of the IV phenotype in
SWV i I + mice. Recently, using PCR to detect the Zgh-C allele, we tested two of these
SWV +I+ animals with the IV phenotype
and have confirmed that these cases are not
due to accidental crosses to SIV (Fig. 2). Unfortunately, those + / + mice that have heterotaxia with situs solitus cannot be ascertained prior to autopsy, and thus cannot be
used for breeding tests. Recently, however,
a male SWV + / + mouse (identification
number 10VU41) was born with a reversed
milk spot. His parents had another pup with
heterotaxia and situs solitus and a pair of
littermates of his parents also had a pup
with situs solitus and heterotaxia. Breeding
tests of this male and his close relatives
have been uniformly negative. Matings
with his mother and sisters produced only
normal offspring. Matings between his siblings have also failed to produce any pups
with the IV phenotype. At autopsy, this
mouse had situs ambiguus and the PCR test
was consistent with a + i + genotype a t the
iu locus.
Putting the iv gene on six other inbred
Tests for polymorphism at the Zgh-C locus
demonstrated that the instances of 1V phenotype found in SWV ivl+ and + i + mice
were real and not the result of mistaking
the ivliu albino parent for one that is + i + .
In contrast to iviiv mice, those showing the
G-odd (n greater than or equal to 3) effect
and phenodeviant SWV + / + mice most often have heterotaxia with situs solitus (Table 2). In G1, the genotype of all animals is
heterozygous a t both the IV and HI loci, so
little reversal is expected. This appears to
represent a graded response; the mildest expression of the phenotype is heterotaxia
alone. Next most frequent is heterotaxia
with situs inversus. Situs inversus without
heterotaxia is rare in SWV mice.
The G-odd effect is due to a n interaction of
the iu gene with the SWV genome, possibly
with a hypothetical mutant gene that we
shall call “hi.” We propose t h a t both iu and
hi are loss of function mutations. The iv
gene product enforces normal coordination
of the sense of asymmetry among asymmetrical structures and sets a switch that determines global situs. Without the iv gene
product the coordination of asymmetry is
imperfect, resulting in heterotaxia, and the
situs switch is set randomly. Although the
hi gene product acts in a manner similar to
that of iu i t is weaker and acts mostly t o
enforce coordination of asymmetry. In its
absence (in the SWV mouse) there are rare
instances of heterotaxia, a few of which
have situs inversus. However, in the absence of the hi gene product, iv acts as a
dominant trait due to haploinsufficiency . In
such a case the resulting phenotype is most
likely to consist of heterotaxia, less often of
heterotaxia with situs inversus, and rarely
of situs inversus alone.
A number of phenodeviant traits, such a s
the cases of IV phenotype described here,
have been found in various inbred mouse
strains. For example, lateral cleft lip, open
eyelids at birth, and atrial septa1 defects
are found in the A strain (Kalter, ’68; Nora
et al., ’68).In the C57BLi6 strain, microphthalmia or anophthalmia is relatively common (Kalter, ’79). This strain also has a
IV (REVERSED) PHENOTYPE IN SWV MICE
low incidence of ventricular septa1 defects
(Nora et al., '68). As we have documented,
stochastic effects may play a n important
role in these cases (Kurnit et al., '87).
The penetrance and expression of single
gene-determined phenotypes can be
changed by genetic background (Juriloff et
al., '87). For example, the first arch (fur)
mutation is lethal when homozygous due to
abnormal development of structures derived
from the maxillary process of the first branchial arch (Juriloff and Harris, '83). The far
mutation occurred in the BALBicGaBc
strain; heterozygotes (far/ ) in this strain
are normal, but in the ICRiBc strain most
heterozygotes have some expression of far
such a s disrupted patterns of mystacial
vibrissae (8081, hemifacial deficiencies
(40%1),and cleft palate (20%) (Juriloff et al.,
'87). Breeding studies suggest the change
from recessive to partial dominance of the
fur mutation is a n effect of genetic background due to a small number of loci rather
than due to isolalleles or a difference between BALBicGaBc and ICR/Bc in the normal wild-type ( + ) alleles at the fur locus
(Harris and Juriloff, '89). Another mutation, Dactylaplasia (Due)is a dominant mutation that causes absence of the median
digits of all four limbs and is a recessive
lethal due to unknown causes (Chai, '81).
The phenotype of dactylaplasia in D a d +
heterozygotes depends on homozygosity for
a n unlinked recessive modifier (mdacl
rnduc); a dominant supressor ( + / + or
mdad + ) causes Dad + heterozygotes to develop phenotypically normal digits.
The incidence of some phenodeviant traits
can be changed by altering the intrauterine
environment. The incidence of situs inuersus, heterotaxia, and heart malformations
in the Non Obese Diabetic (NOD) strain of
mice is affected by the presence and severity
of diabetes in the dam during early pregnancy (Morishima et al., '91). The incidence
of the IV phenotype varied from .05% in the
offspring of non-diabetic dams to 31% in
those from dams that were diabetic during
the first 3 days of pregnancy. Similar to the
findings reported here for SWV, in the NOD
mouse situs inversus without heterotaxia
was rare. Most cases of the IV phenotype
consisted of heterotaxia with situs solztus.
There is some indication of clustering of the
IV phenotype in the SWV iuiiu mouse,
which suggests a n environmental effect.
Thus stochastic, genetic, and/or environ-
+
60 1
mental factors may play a role in the expression of the IV phenotype.
Because the discovery of heterotaxia requires a detailed post mortem examination,
it may be more prevalent in laboratory mice
than is generally recognized. For example,
the WBiReJ strain of mice have a variety of
azygous drainage patterns of the thorax,
some of which are similar to those found in
the IV phenotype iBiddle e t al., '91).
Determination of laterality therefore involves at least two loci, viz., IV and HI. The
finding that putative + i + h i!hi SWV mice
only rarely show the IV phenotype but that
i d + hi/hi SWV mice more often show the
IV phenotype indicates that the HI locus interacts with the IV locus to effect laterality.
ACKNOWLEDGMENTS
This work was supported by a grant from
the NIH (IIL 377031, by the Howard Hughes
Medical Institute, and by the New Hampshire AHA. DMK is an Investigator, AJH is
an Associate, and MVK is a Research Specialist of the Howard Hughes Medical Institute.
LITERATURE CITED
Biddle, F.G., J.D. Jung, and B.A. Eales (19911 Genetically-determined variation in the azygous vein in the
mouse. Teratology, 44:675-683.
Brueckner, M., P. D'Eustachio, and A.L. Horwich i 19891
Linkage mapping of a mouse gene, iu, that controls
left-right asymmetry of the heart and viscera. Proc.
Natl. Acad. Sci. U.S.A., 86:5035-5038.
Chai, C.K. (1981) Dactylaplasia in mice: A two-locu::
model for development anomalies. J. Hered., 72.234237.
Green, E.L. (1981) Genetics and Probability in Animal
Breeding Experiments. New York, Oxford University
Press.
Hanzlik, A.J., M. Binder, W.M. Layton, L. Rowe. 11.
Layton, B.A. Taylor, M.M. Osemlak, J.E. Richards.
D.M. Kurnit, arid G.D. Stewart (19901 The murine
situs inversus viscerum iiv) gene responsible for visceral asymmetry is linked tightly to the fgh-Ccluster
on chromosome 12. Genomics, 7:389%393.
Harris, M.J., and D.M. Juriloff (1989)Test of the isoallele hypothesis at the mouse first arch ( f a r )locus. J.
Hered., 80:127-131.
Hummel K.P., and D.B. Chapman (1959) Visceral inversion and associated anomalies in the mouse. J .
Hered., 502-13.
Juriloff, D.M., and M.J. Harris (19831Abnormal facml
development in the mouse mutant first arch. J. Craniofac. Genet. Dev. B i d ?3:317-337.
Juriloff, D.M., M.J. Harris, and U. Froster-Iskenius
(1987) Hemifacial deficiency induced by a shift in
dominance of the mouse mutation far: A possihie genetic model for hemifacial microsomia. J . Craniofac.
Gen. Dev. Riol.; 7:27--44.
Kalter, H. (1968) Sporadic congenital malformations of
newborn inbred mice. Teratology, 1.193-200
Kalter, H. (19791 The history of the A family ot'inbred
602
W.M. LAYTON ET AL.
mice and the biology of its congenital malformations.
Teratology, 20t213-232.
Kurnit, D.M., W.M. Layton, and S. Matthysse (1987)
Genetics, chance, and morphogenesis. Am. J. Hum.
Genet., 41t979-995.
Layton, W.M. (1976) Random determination of a developmental process. Reversal of normal visceral asymmetry in the mouse. J. Hered., 67:336-338.
Layton, W.M. (1978) Heart malformations in mice homozygous for a gene causing situs inuersus. Birth Defects Orig. Art. Series, 14r277-293.
Morishima, M., M. Ando, and A. Takao (1991) Vis-
ceroatrial heterotaxy syndrome in the NOD mouse
with special reference to atrial situs. Teratology, 44:
91-100.
Nora, J.J., R.J. Sommerville, and F.C. Fraser (1968)
Homologies for congenital heart diseases: Murine
models, influenced by dextroamphetamine. Teratology, lt413-416.
Saiki, R.K., S.Scharf, F. Faloona, K.R. Mullis, G. Horn,
H.A. Erlich, and N. Arnheim (1985) Enzymatic amplification of p-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia.
Science, 230:1350-1354.