* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Figure 2-1
Gene expression programming wikipedia , lookup
Genomic imprinting wikipedia , lookup
Pathogenomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Ridge (biology) wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genomic library wikipedia , lookup
Genome evolution wikipedia , lookup
Minimal genome wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Genome (book) wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Point mutation wikipedia , lookup
Metagenomics wikipedia , lookup
Helitron (biology) wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome editing wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Microevolution wikipedia , lookup
Designer baby wikipedia , lookup
Gene expression profiling wikipedia , lookup
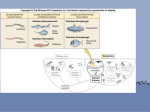
Hi Danielle, good job. I used the Tools:Track Changes option in MSWord to make comments; I focused on the technical aspects of the paper rather than style (I made a couple comments on style). Do you want me to send you pict files or other for your figures rather than you using scanned images? Introduction The cytochrome P450 enzymes compose a well-studied superfamily of monooxygenase hemoproteins involved in the hydroxylation of both endogenous and exogenous substrates. P450 isozymes are found in animals, plants, and bacteria, and in all mammalian cells except mature red blood cells and skeletal muscle cells. Most P450s perform essential physiological functions (e.g. steroid hormone biosynthesis, glucocortoid production, fatty acid hydroxlation) and are thus expressed constitutively (Waterman et al., 1986). Other inducible forms metabolize lipophilic xenobiotics into water-soluble compounds. Induction can occur at either the transcriptional or posttranscriptional level; inducers include anthropogenic pollutants (2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD), phenylhydrazine, furans), barbiturates, and many other aliphatic and aromatic compounds. As terminal oxidases in electron transport chains, P450 enzymes follow the general equation derived by Mason (1957): H+ + XH2 + NADPH + O2 XHOH + NAPD+ + H2O, where X is the xenobiotic substrate. These reactions can release highly reactive free radicals that lead to membrane destruction and carcinogenesis. For this reason, P450 metabolism results in the toxic effects of many common environmental pollutants. Chlorinated dioxins, other planar halogenated aromatic hydrocarbons (HAHs), polynuclear aromatic hydrocarbons (PAHs), and aromatic amines induce cytochrome P450 1A (CYP1A) expression. Located predominantly in the hepatic and extrahepatic tissues, CYP1A is induced through ligand activation of aryl hydrocarbon receptor (AHR). Upon activation in the cytosol, AHR is translocated to the nucleus where it dimerizes with ARNT (AHR nuclear translocator), creating an active transcription factor that initiates transcription of the CYP1A gene. Stegeman (1993) proposes that CYP1A induction is a likely prerequisite for the carcinogenesis that typically results from PAHs and HAHs. YOU CAN MENTION HERE THAT SOME METABOLITES (ex. benzo[a]pyrene:diol epoxide by way of 1A biotransformation) ARE DAMAGING TO DNA BY FORMING FREE RADICALS OR ADDUCTS (Timbrell, 2000; Principles of Biochemical Toxicology, 3rd edition, Taylor & Francis, PA. Although the CYP1A/AHR system efficiently metabolizes PAHs and HAHs, it is unclear whether this is the primary function of CYP1A/AHR, as these toxins do not exist in nature. It is likely that endogenous AHR ligand exists and induces 1A for physiological role (see attached paper) AHR-null mice have deleterious changes in the heart, ovary, liver, and immune system, indicating that CYP1A induction is an adaptive function of AHR (Benedict et al., 2000; Lahvis et al., 2000; Robles et al., 2000; Thurmond et al., 2000; Fernandez-Salguero et al., 1997, 1995; Schmidt et al., 1996). Natural compounds that may act as inducers for CYP1A through AHR or alternate pathways include caffeine, arachidonic acid, and some hormones. Orthologous CYP1A genes have been sequenced and characterized in most vertebrates: birds, mammals, amphibians, and bony fish. Mammals show two types: CYP1A1 and CYP1A2. Hahn et al. (1998) successfully induced CYP1A in cartilaginous fish (little skate, Raja erinacea), but found no evidence for CYP1A induction (there was some immunochemical evidence in lamprey) in jawless fish (sea lamprey, Petromyzon marinus, and Atlantic hagfish, Myxine glutinosa). Photoaffinity labeling studies of more primitive animals (mollusks, annelids, echinoderms, and a urochordate) show no specific binding to AHR proteins (Hahn et al., 1994) see Butler et al (attached). If CYP1A exists in these phyla, it is a variant with a different induction pathway. This indicates that the ancestral gene of CYP1A (as it is known) originated no earlier than 400 million years ago, in the lower Devonian period when the jawed vertebrates diverged from the jawless fish and other primitive vertebrates. A summary of known CYP1A genes is shown in Table 1. <insert Table 1> The cartilaginous and bony fish diverged from each other prior to the divergence of tetrapods in the lower Devonian, 400 million years ago. The elasmobranchs (skates, sharks, and rays) diverged from the chimaeroids in the upper Devonian. Rays and skates together made a final divergence, from the shark lineage, in the Jurassic period, 150 million years ago. At this point, most elasmobranch lineages had become extinct; by the beginning of the Cenozoic, the extant Chondrichthys taxa were represented. On the contrary, bony fish evolution continued at an expontential rate, resulting in more than 24,600 extant species (Nelson, 1994). CYP1A has been cloned and sequenced from numerous teleosts (redundant to paragraph after next) The probable existence of CYP1A and other orthologous P450 genes in Chondrichthys is of particular interest in research of carcinogenesis. It is widely acknowledged that sharks, rays, and other elasmobranchs exhibit an unusually low incidence of cancer (it may seem so, but the fact is that there is really not enough data to say “widely acknowledged”-- it has become a dogma after the publication of a book (I think it was called Sharks don’t get cancer) by a person who later cashed in on using cartilage pills as “natural remedy”. By the way, in my experience sharks do get tumorous growths in their reproductive tract; I did not characterize them to be sure they were a “cancer.” I think you may want to shelve these statements. Some antiangiogenic compounds have been isolated from shark cartilage and clinical trials are on, but it is no panacea. There is evidence to support that the elasmobranch immune system has some features which reduce cancer cell cycling. All this research is still underway in different institutions around the world and hopefully there will be more definitive explanations. It is unknown if there is a connection between this low incidence and CYP1A. (probably no connection, since it seems that their 1A is active (EROD data) this is really too speculative to mention) This study aims to induce, clone, and sequence a novel CYP1A gene in little skate, Raja erinacea and to determine the phylogenetic relationship of skate CYP1A to that of bony fish and tetrapods. Materials and Methods CYP1A Induction A mature female little skate (Raja erinacea) was obtained from the Marine Biological Laboratory, Woods Hole, MA, and kept in constant-flow sea water at 14C. It weighed 0.5kg, had a total length of 45cm, and a disc width of 25cm. It was injected intraperitoneally with the model CYP1A inducer -naphthoflavone (BNF, 50 mg kg-1 body weight) suspended in soybean oil. The skate was anesthetized and euthanized 48 hours post-injection and the liver, pancreas, spleen, spiral valve, stomach, rectal gland, shell gland, gill lamellae, kidney, heart, muscle, and brain were dissected and preserved in RNAlater, a tissue storage reagent (Ambion Diagnostics, Austin, TX) and flash frozen in liquid nitrogen and kept at –80 C until processed. cDNA Preparation Total RNA was isolated from liver tissue with RNA STAT-60 (Tel-test, Friendswood, TX). Liver mRNA (Poly(A)+ )was isolated with Oligotex spin columns (Qiagen, Valencia, CA). Complementary DNA was generated from 1 g poly(A)+ mRNA with oligo-d(T)-primed reverse transcription using the Omniscript RT kit (Qiagen) and a Gene-AMP 2400 thermocycler. Oligonucleotide Primers Degenerate primers were kindly provided by Celine Godard (Woods Hole Oceanographic Institution, Woods Hole, MA). Primer sequences were: 203F1A, 5-GTIGTIWSIGTIGCI AAYGT-3; and 336R1A, 5-GTRTCRAAICCIGCICCRAAIARRTC-3; the primers were synthesized by Life Technologies (Invitrogen, Carlsbad, California). PCR Amplification Amplification of 10 L cDNA was accomplished in 50 L reactions with 10X Gold Buffer, 2mM MgCl2, 0.8 mM of each of four dNTPs, 100 M of each primer, and 5 units of Amplitaq Gold polymerase (Perkin Elmer, Foster City, CA). PCR conditions were: an initial denaturation step of 10 min at 95C, 35 cycles of 15 sec denaturation at 95C and 90 sec annealing/extension at 50C, and a final extension at 50C for 10 min. Cloning and DNA Sequencing A PCR product of the predicted length (400 bp) was visualized on 1% ethidium bromide stained agarose gel, purified with the GeneClean kit (Qbiogene, Carlsbad, CA), cloned into pGEM-T Easy vector (Promega, Madison, WI) and transformed into JM109 competent cells (Promega). Plasmid DNA was checked for insert with EcoRI restriction enzyme digestion; clones containing the 400 bp insert were further purified with the QIAprep Spin Miniprep kit (Qiagen) and sent to the University of Maine-Orono DNA Sequencing Facility for sequencing. Sequence Analysis The NCBI BLASTX search program (www.ncbi.nlm.nih.gov) was used to compare the resulting nucleotide sequence with known genes. Related CYP1A sequences were aligned with ClustalX (Higgins and Sharp, 1988). The PAUP program was used to conduct phylogenetic analyses (Sinauer Assocs., Sunderland, MA; Swofford, 1993). Results Identification of a CYP1A-like Gene in Raja erinacea (You may want to think about refraining from using these types of statements in science writing. A better way of saying it would be in the active affirmative= ex. We amplified the CYP1A ortholog from little skate using RT-PCR with degenerate primers. of finding a CYP1A homolog in little skate (R. erinacea), liver RNA was isolated, then amplified using RT-PCR with degenerate primers CYP1A-203F/CYP1A-336R. The primary PCR product was approximately 400bp as expected. The product was cloned with the pGEM-T Easy vector and JM109 competent cells (Promega). Insert size was confirmed with digestion by EcoRI; two 400bp clones were selected for sequencing. The consensus partial nucleotide sequence was compared to known sequences with a standard nucleotide-nucleotide BLAST search. Instead of verbalizing, you could insert figures?! I.e. Ethidium stained agarose gel with ladder showing PCR product or clones with insert. The highest scoring BLAST results indicate that the fragment is a highly conserved region of a CYP1A gene. The partial cDNA(393 bp), is most similar to the fathead minnow, Pimephales promelas CYP1A . There was high nucleotide identity, 84% to 92%, between among the skate, chicken, rat, eel, and carp CYP1A cDNA. The skate partial nucleotide and corresponding amino acid sequence is shown in Figure 1. Nucleotide identities between the fragment and various CYP1A genes are presented in Table 2. <insert Figure 1, Table 2> PROBABLY BETTER TO LOOK AT AMINO ACID IDENTITIES RATHER THAN DNA Phylogenetic Analysis The skate CYP1A fragment was aligned with fourteen various CYP1 sequences (Table 3) using ClustalX software. Phylogenetic analyses were made with PAUP.( MACCLADE WAS USED JUST TO FORMAT THE FILE, NOT NECESSARY TO MENTION) An unrooted cladogram, presented in Figure 2-1, shows the CYP1 sequences cluster in four distinct monophyletic groups, or clades: CYP1Bs, mammals CYP1As, bony fish CYP1As, and the skate CYP1A, which stands as its own group. A phylogenetic tree (Figure 2-2) was created with monkey CYP2B as an outgroup. The skate CYP1A appears between the mammal and bony fish CYP1As; as in the unrooted cladogram, the plaice gene (PleplA) does not cluster with the other bony fish and has apparently been misidentified. Finally, a maximum parsimony tree, Figure 2-3, was constructed with the plaice CYP1A as the outgroup. Skate CYP1A appears in the same lineage as the bird and mammal CYP1As, in a group divergent from the bony fish. (YOU SHOULD MENTION THAT A MORE COMPLETE ANALYSIS WILL BE (I did) DONE USING THE FULL LENGTH AMINO ACID SEQUENCE) In fact, the trees are a bit different than the trees you show here. The skate clusters within the bony fish 1A . see poster <insert Table 3, Figures 2-1, 2-2, 2-3> ONE THING YOU COULD MENTION (and possibly do yourself) IS THAT THERE ARE WHOLE GENOMES AVAILABLE FOR SOME INVERTEBRATES (C. ELEGANS; DROSOPHILA; ANOPHALES MOSQUITO) AND A SEARCH WITH CYP1A RESULTS IN NO HOMOLOGOUS GENES. THERE IS ALSO A CIONA INTESTINALIS GENOME (tunicate) ….I have not searched it….might be interesting. THERE ARE SOME PLACES IN P450S THAT ARE CONSERVED AMONG OTHER FAMILIES (I-HELIX, HEME-BINDING REGION ETC) WHICH MAY POP UP IF SEARCHING THESE GENOMES WITH A CYP1A SEQUENCE, BUT ACTUALLY REPRESENT OTHER P450 ENZYMES. Table 1 Summary of Known CYP1A Genes Taxa Mammals Birds Reptiles Amphibians Bony Fish Cartilaginous Fish Jawless Fish Invertebrates Induced + + + + + + - Sequenced Selected Species + + + - Mouse (Mus musculus), Rat (Rattus norvegicus), Human (Homo sapiens) Chicken (Gallus gallus), Herring Gull (Larus argentatus) Eastern Painted Turtle (Chrysemys picta picta) Newt (Pleurodeles waltl), Tiger Salamander (Ambystoma tigrinum) Zebrafish (Danio rerio), Mummichog (Fundulus heteroclitus) Skate (Raja erinacea) Atlantic Hagfish (Myxine glutinosa), Sea Lamprey (Petromyzon marinus) Soft-Shell Clam (Mya arenaria) Figure 1 393bp Consensus Partial Nucleotide and translated Amino Acid Sequence for ReCYP1A cDNA GTG GTG TCG GTG GCG AAT GTC ATT TGC GCC CTG TGC TTC GGG AAG CGC TAC AGT CAC GAA GAC CAA GAG CTC V V S V A N V I C A L C F G K R Y S H E D Q E L CTC AAC ATC GTC AAT GTC AGC GAT GAG TTC GGC AAA ATC GTA GCC GCT GGC AAC CCC GCC GAT TTC ATC CCG L N I V N V S D E F G K I V A A G N P A D F I P ATC CTG AGA TTC CTC CCG AAC CAC TCG ATG GAC AAG TTC ATC GCT ATC AAC AAG AGA TTC GCC ACT TTC GTT I L R F L P N H S M D K F I A I N K R F A T F V GAG AAC ATT GTC ATG GAG CAT TAC CGC ACA TTT GAC AAG GAT AAC ATT CGG GAT ATA ACC GAT TCG CTG ATC E N I V M E H Y R T F D K D N I R D I T D S L I GGT CAC TGC CAG GAT AAA AAA GTG GAC GAG AAT GCC AAT ATC CAA ATA TCC GAT GAA AAG ATT GTC GGC ATC G H C Q D K K V D D N A N I Q I S D E K I V G I GTG AAT GAC CTC TTC GGC GCC GGC TTC GAC ACA V N D L F G A G F D T Table 2 Nucleotide Identities between Aligned CYP1A Sequences Minnow 1A Carp 1A Minnow 1A Carp 1A Eel 1A Chicken 1A4 Chicken 1A5 Rat 1A1 Pig 1A1 Skate Fragment 1.00 Eel 1A Chicken 1A4 Chicken 1A5 Rat 1A1 Pig 1A1 Skate Fragment 0.87 0.80 0.85 0.90 0.85 0.88 0.91 1.00 0.81 0.85 0.85 0.89 0.83 0.92 1.00 0.89 0.89 0.86 0.96 0.83 1.00 0.96 0.87 0.79 0.84 1.00 0.87 0.89 0.84 1.00 0.83 0.85 1.00 0.88 1.00 Table 3 GenBank Accession Numbers for CYP1 Genes Used in Phylogenetic Analysis Gene Label Organism Species Accession Number CYP1A Funhe1A Mummichog Fundulus heteroclitus AAD01809 CYP1A Ammma1A Sand lance Ammodytes marinus CAC342 CYP1A Lizau1A Mullet Liza aurata AAB70307 CYP1A Limli1A Dab Limanda limanda CAA0495 CYP1A Angja1A Japanese eel Anguilla japonica BAA8824 CYP1A Plepl1A Plaice Pleuronectes platessa CAB5136 CYP1A1 Oncmy1A1 Rainbow trout Oncorhynchus mykiss A28789 CYP1A1 Micto1A1 Tomcod Microgadus tomcod IS6646 CYP1A1 Ratno1A1 Rat Rattus norvegicus IO4RTMC CYP1A1 Oviar1A1 Sheep Ovis aries AAD1431 CYP1A1 Homsa1A1 Human Homo sapiens NP_00075 CYP1A4 Galdo1A4 Chicken Gallus domesticus X99453 CYP1B1 Musmu1B1 Mouse Mus musculus NP_034 CYP1B1 Homsa1B1 Human Homo sapiens Q16678 Figure 2-1 Unrooted Cladogram with Fish, Bird, and Mammal CYP1 Genes Four distinct clades: CYP1Bs, bird and mammal CYP1s (with plaice CYP1A, “Plepl1A”, as irregular result), skate CYP1A, and bony fish CYP1As. The plaice gene has likely been misidentified or has an incorrect GenBank entry. Figure 2-2 Phylogenetic Tree Showing Divergence of CYP1 Genes The bony fish CYP1A genes clade together, with the exception of the plaice gene (see Figure 4-1 and Results). The skate CYP1A gene appears between the bony fish clade and the chicken and human CYP1As. Figure 2-3 Phylogenetic Analysis of CYP1 Genes using Maximum Parsimony Tree constructed using maximum parsimony with bootstrapping. Most bony fish CYP1A genes are clustered together, as are the bird and mammal CYP1As. The skate CYP1A fragment is most closely associated with the bird and mammal clade. References Benedict, J.C., T.M. Lin, I.K. Loeffler, R.E. Peterson, and J.A. Flaws. (2000). Physicological role of the aryl hydrocarbon receptor in mouse ovary development. Toxicol. Sci. 56:382-388. Carrol, R.L. (1988). Vertebrate paleontology and evolution. W.H. Freedman, NY. Colbert, E.H. (1969). Evolution of the Vertebrates. 2d. ed. Wiley, NY. Fernandez-Salguerno, P.M., T. Pineau, D.M. Hilbert, T. McPhail, S.S.T. Lee, S. Kimura, D.W. Nebert, S. Rudikoff, J.M. Ward, and F.J. Gonzales. (1995). Immune system impairment and hepatic fibrosis in mice kacing the dioxinbinding Ah receptor. Science 268:722-726. Fernandez-Salguerno, P.M., J.M. Ward, J.P. Sundberg, and F.J. Gonzales. (1997). Lesions of aryl hydrocarbon receptor deficient mice. Vet. Pathol. 34:605-614. Garfinkel, D. (1958). Studies on pig liver microsomes. I. Enzyme and pigment composition of different microsomal fractions. Arch. Biochem. Biophys. 77: 493-509. Hahn, M.E., A. Poland, E. Glover, and J.J. Stegeman. (1994). Photoaffinity Labeling of the Ah Receptor: Phylogenetic Survey of Diverse Vertebrate and Invertebrate Species. Arch. Biochem. Biophys. 310(1):218-228. Hahn, M.E., B.R. Woodin, J.J. Stegeman, and D.E. Tillitt. (1998). Aryl hydrocarbon receptor function in early vertebrates: Inducibility of cytochrome P450 1A in agnathan and elasmobranch fish. Comp. Biochem. Phys. C 120:67-75. Hahn, M.E. (2002). Aryl hydrocarbon receptors: diversity and evolution. Chemico-Biological Interactions 141: 131. Higgins, D.G. and P.M. Sharp. (1988). Clustal: a package for performing multiple sequence alignment on a microcomputer. Gene 73:237-244. Klingenberg, M. (1958). Pigments of rat liver microsomes. Arch. Biochem. Biophys. 75:376-386. Lahvis, G.P., S.L. Lindell, R.S. Thomas, R.S. McCuskey, C. Murphy, E. Glover, M. Bentz, J. Southard, and C.A. Bradfield. (2000). Portosystemic shunting and persistent fetal vascular structures in aryl hydrocarbon receptor deficient mice. Proc. Natl. Acad. Sci. USA 97:10442-10447. Lemberg, R. and J. Barrett.(1973). Cytochromes. Academic Press, London. Mason, H.S. (1957). Mechanisms of Oxygen Metabolism. Advances in Enzymology 19:79-233. Nelson, J.S. (1994). Fishes of the World. 3d. ed. Wiley, NY. Robles, R., Y. Morita, K.K. Mann, G.I. Perez, S. Yang, T. Matikainen, D.H. Sherr, and J.L. Tilly. (2000). The aryl hydrocarbon receptor, a basic helix-loop-helix transcription factor of the PAS gene family, is required for normal ovarian germ cell dynamics in the mouse. Endocrinology 141:45-453. Said, M.R. (2000). Identification of cytochrome P450 1B-like genes in these teleost species and phylogenetic analysis of CYP1 genes. M.Sci. Thesis. Mass. Instit. Tech.: Div. Bioengin. Enviro. Health. June 2000. Schmidt, J.V., G.H.-T. Su, J.K. Reddy, M.C. Simon, and C.A. Bradfield. (1996). Characterization of a murine Ahr null allele: Involvement of the Ah receptor in hepatic growth and development. Proc. Natl. Acad. Sci. USA 93:67316736. Swofford, D.L. (1993). PAUP: Phylogenetic Analysis Using Parsimony. Illinois Natural History Survery. Champaign, IL. Shimada, T. H. Yamazaki, M.Mimura, N. Wakamiya, W.-F. Ueng, F.P. Guengerich, and Y. Inui. (1996). Characterization of microsomal cytochrome P450 enzymes involved in the oxidation of xenobiotic chemicals in human and fetal livers and adult lungs. Drug Metab. Dispos. 24:515-552. Stegeman, J.J. (1993). Diversity and Regulation of Cytochrome P450 in Aquatic Species. Proceedings of the NATO Advanced Study Institute: Molecular Aspects of Oxidative Drug Metabolizing Enzymes: Their Significance in Environmental Toxicology, Chemical Carcinogenesis and Health. Held in Kusadasi, Aydin-Turkey, June 20-July 2, 1993. Editors E. Arinc, J.B. Schlenkman, and E. Hodgson. Thurmond, T.S., J.E. Staples, A.E. Silverstone, and T.A. Gasiewicz. (2001). The aryl hydrocarbon receptor has a role in the in vivo maturation of murine bone marrow B lymphocytes and their response to 2,3,7,8tetrachlorodibenzo-p-dioxin. Toxico. Appl. Pharma. 171:157-164. Waterman, M.R., M.E. John, and E.R. Simpson. (1986). Regulation of Synthesis and Activity of Cytochrome P450 Enzymes in Physiological Pathways. Cytochrome P450. Structure, Mechanism, and Biochemistry. First Edition, Edited by Paul R. Ortiz de Montellano. Plenum Press, NY. Chapter 10. Yu, M.-H. (2000). Environmental Toxicology: Impacts of Environmental Toxicants on Living Systems. Lewis, Boca Raton, FL.