* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download No Slide Title
Butyric acid wikipedia , lookup
Biosynthesis wikipedia , lookup
Signal transduction wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Genetic engineering wikipedia , lookup
Genomic imprinting wikipedia , lookup
Real-time polymerase chain reaction wikipedia , lookup
Expression vector wikipedia , lookup
Gene therapy wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Gene expression wikipedia , lookup
Gene desert wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Community fingerprinting wikipedia , lookup
Gene nomenclature wikipedia , lookup
Point mutation wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Gene regulatory network wikipedia , lookup
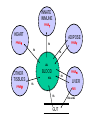
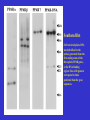
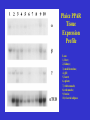
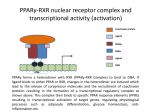
Molecular Control of Fish Lipid Metabolism: Isolation and Characterisation of Peroxisome Proliferator-Activated Receptor (PPAR) Genes from Fish Species 1990 PHARM 0% INDUS 8% FISH OIL USE 2000 Annual production stable at 1.1 to 1.4 million tons PHARM INDUS 2% 10 % AQUA 16 % Actual AQUA 54 % AGRI 34 % AGRI 76 % 2005 PHARM 2% 2010 INDUS 0% INDUS 12 % PHARM 2% AGRI 1% AGRI 9% Projected ?! AQUA 77 % AQUA 97 % Fish Oil Replacement • Fat Deposition? • Nutritional Quality? • Disease Resistance? • Need a better understanding of underlying physiology Peroxisome proliferator -activated receptors • PPARs – Transcription factors – Control genes involved in lipid homeostasis – Activated by PUFA and their eicosanoid derivatives PPARs 9-cisRA PUFA PPAR RXR LIGAND BINDING LIGAND BINDING hinge A/B A/B DNA BINDING AAGTCAnAAGTCA GENE TRANSCRIPTON CHROMOSOME PPRE •PPARs are members of nuclear hormone receptor family •PPARs bind as heterodimer with RXR to PPRE •PPARs are activated by fatty acid (PUFA) ligands •Three forms in mammals, a, b/d and g INNATE IMMUNE PPARg HEART ADIPOSE PPARa PPARg FA FA FA LDL BLOOD OTHER TISSUES PPARb PPARa FA HDL LIVER FA FXR FA GUT Bile acids PPARs and Lipid Homeostasis • Transport – Apolipoprotien AI, AII, CIII, Liver fatty acid binding protein; Fatty acid transport protein; CD36 • Biosynthesis – Acetyl-CoA synthase; Malic enzyme; Stearoyl-CoA desaturase I • Storage – Adipocyte lipid binding protein; Phosphoenolpyruvate carboxylase • Metabolism – Acyl-CoA oxidase; Bifunctional enzyme; Carnitine palmitoyltransferase; CYP4A1, 4A6; Lipoprotein lipase; Medium chain Acyl-CoA dehydrogenase, 3-hydroxy, 3methylglutaryl-CoA synthase; Uncoupling protein I Strategy • Do fish have PPARs? – Construct and screen genomic libraries • What are their ligand activation profiles? – Express fish PPAR genes in cell culture • Diet formulation – Use results to produce a rational framework for fish oil replacement Plaice as a model • Marine species – Highly dependent on fish oil • Small genome- small genes – Facilitates gene isolation from lambda phage libraries • Also salmon, sea bream and sea bass Stategy for PPAR Gene and cDNA Isolation Partial digest Genomic DNA ligate + l bacteriophage arms RT-PCR Isolate and sequence cDNA Isolate and sequence gene Package, plate on lawn of E. coli and screen with hybridisation probe Plaice PPAR Gene Structures 1kb 7kb pPPARa * * 4.5kb pPPARb pPPARg 10kb ** Human PPAR genes are >80kb xlPPARa 100 hsPPARa 100 97 ggPPARa ssPPARa ppPPARa 99 dlPPPARa 100 99 saPPARa ssPPARb1 95 99 ppPPARb dlPPARb 100 99 saPPARb hsPPARb 100 99 ggPPARb 100 xlPPARb xlPPARg 89 hsPPARg 100 Phylogenetic plot of PPAR sequences. ggPPARg ssPPARg ppPPARg 100 dlPPARg 100 81 saPPARg xl. Xenopus laevis; hs, Homo sapiens; gg, Gallus gallus; ss, Salmo salar; pp, Pleuronectes platessa; dl, Dicentrarchus labrax; sa, Sparus aurata. Southern Blot. SstI restricted plaice DNA was hybridised to the probes generated from the first coding exons of the three plaice PPAR genes, or the DNA-binding region. Sizes of fragments correspond to those predicted from the gene sequences. PPAR structure and function Ligand-independent transactivation (phosphorylation?) DNA-binding, Dimerisation, Co-activatorbinding A/B C 20% 90% Ligand-binding, Co-activator-binding D E/F 70% PPAR RXR E/F E/F A/B C C A/B DNA PROMOTER EMSA Performed with in vitro translated fish PPARs and plaice RXR and mouse ACO gene promoter oligo PPAR Transactivation Assays Co-transfect to cells in culture (Multiwell plates) Ligate constitutive gene promoter to PPARgene CMV PPRE PPAR cDNA CMV CAT gene PPAR Ligate a PPAR response element (PPRE) to CAT reporter gene PPAR PPAR cDNA PPRE RXR CAT gene PPRE CAT Measure CAT (Muliwell ELISA) CAT gene Treat cells with potential PPAR activators Plaice PPAR Tissue Expression Profile Lane 1, liver; 2, kidney; 3, small intestine; 4, gill; 5, heart; 6, spleen; 7, white muscle; 8, red muscle; 9, brain; 10, visceral adipose Next Steps • PPAR activators in primary hepatocytes and adipocytes – Determine fatty acid profiles and metabolic indices – Gene expression profiling • Dietary trial with salmon and sea bream – Measure growth, gene expression, fatty acid profiles Dietary Trial • PPARa - Liver and Heart- Fatty acid oxidation– Conjugated linolenic acid (CLA), 16:1, 18:1 ??? • PPARb - All tissues- Function? – 16:1 • PPARg - Adipose - Fat Sorage – No good natural Fatty Acid Ligands • Diet- 16:1 + 18:3 + CLA????