* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download How does the cell regulate arsenate respiration and
Metagenomics wikipedia , lookup
Genomic imprinting wikipedia , lookup
RNA interference wikipedia , lookup
Microevolution wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Primary transcript wikipedia , lookup
Non-coding RNA wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
RNA silencing wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Epitranscriptome wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene expression programming wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression profiling wikipedia , lookup
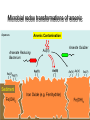
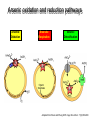
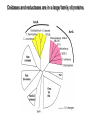
Microbiology of arsenic redox transformations UCSC Chad Saltikov and collaborators Environment Molecular Genetics Public Health Microbial redox transformations of arsenic Aqueous Arsenate Reducing Bacterium Sediment Arsenic Contamination As(III) Arsenite Oxidizer Phylogenetic relationship of arsenic metabolizing microbes Arsenic oxidation and reduction pathways Arsenite Oxidation Arsenate Respiration Arsenic Detoxification Adapted from Silver and Phung 2005. Appl. Env. Micro. 71(2):599-608 Oxidases and reductases are in a large family of proteins arrA genes from other bacteria Akaliphilus metalliredigenes: 66%* Geobacter uraniumreducens: 65%* Desulfosporosinus sp. Y5: 66%* *aa similarities to ArrA of ANA-3 The arsenic island of Shewanella: ANA-3, CN-32, and W3-18-1 CN-32 W3-18-1 What environmental conditions trigger arsenate reduction? ArsC (detoxify) vs. ArrA (respire) As(V) As(III) Monitor two arsenate reduction pathways in our model arsenate reducer Shewanella sp. ANA-3 (an)aerobic vs. As Monitor the transcription of arrA and arsC Saltikov and Newman 2003 PNAS 100(19):10983-10988 Quantify gene specific mRNA: Grow culture to mid log Extract/Purify RNA Reverse Transcribe RNA Quantify genespecific mRNA by real time PCR Threshold Cycle (Ct) Standard curve for 29 y = -3.356x + 19.857 27 R2 = 0.9982 25 23 21 19 0.04 ng 17 15 -3 -2 -1 log ng DNA 0 1 Dynamic expression in various growth phases with As(V) as electron acceptor Saltikov et al. J. Bacteriology 2005. 187 (21): 7390-7396 How does phosphate influence As(V) respiration and arr/ars expression As(V)+Pi Saltikov et al. J. Bacteriology 2005. 187 (21): 7390-7396 Fumarate+Pi Expression What about other electron acceptors? arrA arsC expressiona 0.012 0.004 inductionb - expressiona 0.015 0.005 inductionb - 0.015 0.002 0.013 0.002 - 0.10 0.02 0.37 0.06 7 25 Fumarate As(V) As(III) 0.09 0.03 0.72 ± 0.09 0.92 ± 0.06 7.8 10.0 0.04 ± 0.01 0.44 ± 0.08 0.71 ± 0.04 10.3 16.9 Nitrate As(V) As(III) 0.05 ± 0.02 0.08 ± 0.01 0.07 ± 0.01 - 0.04 ± 0.00 0.21 ± 0.11 0.68 ± 0.12 5.1 16.2 TMAO As(V) As(III) 0.02 ± 0.00 0.43 ± 0.17 0.29 ± 0.03 20.1 13.5 0.02 ± 0.00 0.26 ± 0.09 0.61 ± 0.08 13.6 32.7 Substrate: Oxygen As(V) As(III) As(III) = inducer O2 and NO3- inhibit a. Expression represents the ratio of the relative quantity of arrA or arsC transcripts to that of the housekeeping gene gyrB. b. Induction was determined by normalizing the expression value for arrA or arsC to no As conditions. Saltikov et al. J. Bacteriology 2005. 187 (21): 7390-7396 What are the sensitivities of arrA and arsC expression to As? A. B. ∆arrA, ∆arsC Wild-type No As No As Log [arsenite] µM Saltikov et al. J. Bacteriology 2005. 187 (21): 7390-7396 Log [arsenate] µM Are cytochromes required for arsenate respiration in our model organism? factoid: Shewanella has 39 reading frames encoding ctype cytochromes Iron containing proteins similar to heme of a red blood cell. CymA--tetraheme cytochrome is required for respiring arsenate periplasm As(V) As(III) ArrAB UQH2 UQ dehydrogenase cytoplasm Adapted from Schwalb et al. 2003 Biochemistry 42(31):9491-9497 cymA is required for respiring As(V) in CN-32 Growth on nitrate Growth on arsenate cymA restores growth on As(V) 0.2 ANA-3+pBBR1MCS2 AN-CymA+pBBR1MCS2 AN-CymA+ANpCymA Growth (OD 600 nm) 0.15 AN-CymA+CNpCymA 0.1 0.05 0 0 2 4 6 Time (hours) 8 10 Secondary structure prediction of CymA Inner Membrane Model for microbial arsenate reduction Respiration Detoxification Environmental significance of arsenate respiratory reduction Can arrA be used to monitor and track As(V) reduction? A. ArrA protein and primer design B. Detection in various strains arrA primers 16S rDNA primers 16S rDNA primers Malasarn et al. 2004 Science 06: 455 L. Ladder: 100 bp 1. ANA-3 arrA deleted 2. Shew. oneidensis MR-1 3. Desulfitobacterium dehalogenens 4. E. coli 5. Pseudomonas chloraphis 6. Shewanella sp. ANA-3 7. Desulfito. hafniense 8. Desulfito. frappieri 9. D. strain GBFH 10. Wolinella sp. 11. Wolinella succinogenes 12. Citrobacter sp. 13. Bacillus str. E1H 14. Bacillus, str. MLS10 15. Sulfur. barnesii SES-3 16. Shewanella sp. HAR-4 17. Chrysiogenes 18. OREX-4 19. Pyrobac. arsenaticum Pore water concentrations of total As, Fe, and Mn in Fe/As rich reservoir * Mn Fe As Kneebone et al. 2002 ES&T 36(3):381-386 New primers for detecting arrA-like genes in As-enriched sediments Increasing depth in core 1517 1200 1000 1 2 3 4 5 6 7 8 Blanks 500 400 300 200 100 Primer Dimmers Conclusions Two genetic pathways for arsenate reduction – Respiratory by arr and detoxification by ars Respiration pathway triggered by As: – As(III) > As(V) – Repressed by nitrate and oxygen The mechanism for arsenate reduction involves other components in the cell. The arrA gene is a useful marker for As redox – Gene copy number seems to correlate with redox gradients of As … more work to be done. Acknowledgments/Collaborators UC Santa Cruz – Julie Nilsen Caltech – Prof. Dianne Newman, Prof. Janet Hering, Rich Wildman USGS – Dr. Ron Oremland, Dr. Thomas Kulp, Dr. Larry Miller, Shelly Hoeft