* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Part 1
Structural alignment wikipedia , lookup
List of types of proteins wikipedia , lookup
Implicit solvation wikipedia , lookup
Circular dichroism wikipedia , lookup
Rosetta@home wikipedia , lookup
Protein design wikipedia , lookup
Folding@home wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Homology modeling wikipedia , lookup
Protein moonlighting wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Alpha helix wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Protein purification wikipedia , lookup
Western blot wikipedia , lookup
Protein domain wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein–protein interaction wikipedia , lookup
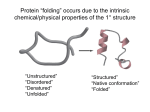
Protein folding and misfolding Folding of proteins into their native conformations occurs spontaneously under physiological conditions and is dictated by the primary structure of the protein. Harini Chandra Affiliations Master Layout (Part 1) 1 This animation consists of 5 parts: Part 1 – Thermodynamics of protein folding Part 2 – Anfinsen’s experiment Part 3 – Amino acid structure determines 3-D folding Part 4 – Molecular chaperones for protein folding Part 5 – Protein misfolding diseases 2 3 Partially folded polypeptide 4 Native state a-helix 5 Source: Biochemistry by Lehninger, 4th edition (ebook) Free energy Unfolded polypeptide chain % residues in native conformation Entropy 1 2 3 4 5 Definitions of the components: Part 1 – thermodynamics of protein folding 1. Entropy: Entropy is a measure of randomness or how disorganized a system is and forms the basis of the second law of thermodynamics, which states that the total entropy of a system cannot decrease without correspondingly increasing the entropy of another system. In other words, the entropy of the universe (system + surrounding) is constantly increasing. Entropy helps in predicting the spontaneity of any process. An unfolded polypeptide chain has high entropy which goes on decreasing as the protein folds into its native state. 2. Free energy: The free energy, also known as Gibbs free energy, is the maximum amount of mechanical work that can be done by a system at constant temperature and pressure. In general, all systems try to attain minimum free energy and a reaction takes place spontaneously only when the associated free energy change is negative. 3. Unfolded polypeptide chain: The amino acids that have been joined together by peptide bonds but have not yet formed their secondary or tertiary structures. This conformation has the highest free energy and entropy. 4. Partially folded polypeptide: The amino acids in the polypeptide chain start interacting by means of hydrogen bonds across the polypeptide backbone in order to initiate the folding process. The free energy and entropy of the system gradually decrease as the folding takes place. 1 2 3 Definitions of the components: Part 1 – thermodynamics of protein folding 5. Native state a-helix: The polypeptide chain assumes its most stable, native conformation in the form of an a-helix with the folding being directed largely by its amino acid residues. This conformation corresponds to minimum free energy and entropy, thereby conferring very high stability. The lowering of entropy is favoured by a corresponding increase in entropy in the surroundings composed of water molecules. 6. Molten globule: Initial collapsed state of a protein with very little thermodynamic stability is known as the molten globule. The amino acid side chains are extremely disordered in this state with several fluctuations being observed. 7. Percentage residues in native conformation: This refers to the number of residues that have assumed their favourable, lowest energy states. The percentage increases gradually as the folding process takes place. 4 5 1 Part 1, Step 1: H2O H2O H2O H2O Beginning of helix formation 2 H2O H2O Unfolded polypeptide chain, high free energy & entropy H2O H2O H2O H2O 4 H2O Helix formation commences, free energy & entropy decrease Action 5 Free energy 3 H2O Entropy The green chain and blue circles shown must assume the different arrangements as shown. Description of the action (Please redraw all figures.) First show the green chain on the top. Then show the blue circles appearing around it . The green chain must then be shown to bend in various ways until it assumes the shape below. And the blue circles must also rearrange themselves as shown. The graph on the right must gradually take shape in the direction indicated. Source: Biochemistry by Lehninger, 4th edition (ebook) Audio Narration An unfolded polypeptide chain has very high free energy and entropy. Protein folding acts to decrease the free energy of the system by forming favorable interactions and assuming a more stable state. The entropy of the polypeptide chain decreases during this process. Part 1, Step 2: H2O H2O H2O H2O H2O Entropy Intrachain hydrogen bonds H2O H2O H O 2 Entropy of water molecules increase, and polypeptide decreases 3 H2O H2O H2O H O 2 H2O H O 2 H2O H2O H O 2 H2O H O 2 Stable native state a-helix 4 Action 5 Free energy 2 % residues in native conformation 1 The black dotted lines must gradually appear on the green chain on top. The chain and blue circles must rearrange themselves. Description of the action (Please redraw all figures.) The blue circles must move around away from the green chain on top and form small clusters. The black dotted lines must appear on as shown on the green chain on top. The figure below must appear as shown and the graph on the right must be gradually completed. Source: Biochemistry by Lehninger, 4th edition (ebook) Audio Narration As the protein continues to fold in order to assume its stable, low energy native state conformation, the entropy also decreases. While this would seem unfavorable for the system, it must be recalled that the entropy of the surrounding water molecules increases during the process, thereby increasing the overall entropy and making it favorable and spontaneous. Master Layout (Part 2) 1 2 This animation consists of 5 parts: Part 1 – Thermodynamics of protein folding Part 2 – Anfinsen’s experiment Part 3 – Amino acid structure determines 3-D folding Part 4 – Molecular chaperones for protein folding Part 5 – Protein misfolding diseases b-mercaptoethanol Disulphide bonds 3 6M urea Noncovalent interaction Native ribonuclease A Remove urea & b-mercaptoethanol Broken disulphide linkages 4 Denatured ribonuclease A 5 1 2 3 Definitions of the components: Part 2 – Anfinsen’s experiment 1. Native ribonuclease A: This is an endonuclease enzyme composed of 124 amino acids that cleaves single-stranded RNA molecules. It has four disulphide bonds in its native state that are essential for conformational folding and enzymatic activity. This was used by Christian Anfinsen to postulate the thermodynamic hypothesis of protein folding, according to which the folded form of a protein represents its free energy minimum. 2. b-mercaptoethanol: b or 2-mercaptoethanol with the formula OHCH2CH2SH is a chemical compound that is used commonly to reduce disulphide linkages in proteins, thereby disrupting the tertiary and quaternary structures. 3. 6M urea: It is an organic compound having two amine groups joined by a carbonyl group and used at concentrations up to 10 M for denaturing proteins by breaking the noncovalent interactions. 4 5 4. Denatured ribonuclease A: On treatment with b-mercaptoethanol and urea, the ribonuclease A loses its native conformation due to breaking of the disulphide and noncovalent linkages. Activity of the enzyme is also lost during this process. However, it was observed by Anfinsen that removal of both urea and b-mercaptoethanol allows the enzyme to fold into its native conformation again with more than 90% enzymatic activity. 1 Part 2, Step 1: b-mercaptoethanol 2 6M urea 3 Native state ribonuclease A Broken disulphide linkages Denatured ribonuclease A 4 Action 5 The green pie shaped objects must break the black lines while the blue pie objects must be break the dotted lines. Description of the action First show the structure on the left. Then show the green pie shaped objects moving towards the black lines and breaking them and simultaneously the blue pie shaped objects breaking the dotted lines. The structure must then unfold and give rise to the open chain displayed below with the green & blue objects remaining bound to it. Audio Narration Ribonuclease A in its native state has four disulphide bonds between its cysteine residues. When treated with bmercaptoethanol and 6M urea, the protein undergoes denaturation and the disulphide linkages are broken. Enzyme activity is lost in the denatured state. 1 Part 2, Step 2: Denatured ribonuclease A 2 Remove urea and b-mercaptoethanol Remove bmercaptoethanol only 3 4 Native state ribonuclease A Action Description of the action 5 The chain must twist itself and reform the structures shown below. Movement in 3D must be shown. First show the structure on top followed by the right arrow & text. The green pieshaped objects must be removed & the structure on the right must appear. Next the left arrow & text must be shown with disappearance of the blue pie-shaped objects & appearance of the figure on the left. Inactive ribonuclease A Audio Narration It was observed by Anfinsen that removal of urea and bmercaptoethanol led to the refolding of the enzyme to assume its native state with more than 90% enzyme activity being intact. However, if only b-mercaptoethanol was removed in presence of urea, the formation of disulphide bonds was random, leading to enzyme with only around 1% activity. Master Layout (Part 3) 1 This animation consists of 5 parts: Part 1 – Thermodynamics of protein folding Part 2 – Anfinsen’s experiment Part 3 – Amino acid structure determines 3-D folding Part 4 – Molecular chaperones for protein folding Part 5 – Protein misfolding diseases 2 Protein folding 1 3 4 2 3 4 Amino acid sequence 1 1 2 3 Protein 1 4 Amino acid sequence 2 Protein 2 5 Source: Biochemistry by Stryer, 5th edition (ebook) 1 2 Definitions of the components: Part 3 – Amino acid structure determines 3-D structure 1. Amino acid sequence 1, 2: These are two completely different amino acid sequences that will give rise to different protein structures. 2. Protein 1, 2: The protein structure corresponding to amino acid sequence 1 and 2 respectively. The first amino acid sequence cannot give rise to the second protein structure & vice versa. 3 4 5 3. Protein folding: The process by which the amino acid side chains in the proteins interact with one another to form energetically favourable bonds with each other thereby allowing regions that are far away from one another to move closer. This process is determined by the amino acid sequence of the proteins and needs to be energetically feasible in order to take place. 1 Part 3, Step 1: Protein folding is governed by distribution of polar & non-polar amino acid residues in proteins Aqueous environment Polar side chains H2O H2O 2 H2O H2O Hydrogen bond interactions Non-polar side chains 3 Folded protein Hydrophobic residues buried inside H2O H2O H2O 4 Action Description of the action Folding of the grey chain. 5 First show the structure on the left with the blue & green projections. Next show this chain folding such that all the green parts come on the inside & blue parts are on the outside as shown in figure on the right. This must be surrounded by water molecules which must move towards the blue regions as depicted in the animations. Audio Narration The process of protein folding is governed by the distribution of polar and non-polar amino acid residues in the protein. Hydrophobic amino acids are driven to interact with one another, a process termed as hydrophobic collapse. They come together andin the process, eliminate water molecules around them. The polar residues remain on the surface and form hydrogen bonds with water molecules while the hydrophobic residues get buried within the core of the protein. 1 Part 3, Step 2: Protein folding is a cooperative process while unfolding is a sharp, quick transition 2 Unfolded state 3 Partially folded Partially folded conformations Folded native state 4 Action Description of the action 5 Show the figures above appearing one at a time followed by the graph. Audio Narration Proteins typically adopt only one characteristic functional native state (Please redraw all figures.)First show conformation which has lowest free energy and is most stable. each of the figures appearing one after another. In each, the chain must become Folding is limited to one conformation due to properties of the amino more compact as it progresses. Once the acid side chains such as hydrophobicity, size, shape etc. Folding is a highly cooperative process wherein there is progressive stabilization last figure has been shown, the graph of the intermediates. Although it is theoretically possible to predict must appear with the blue figure being unfolded to give the red chain. As soon as protein structure from the amino acid sequence, several long-range the structure starts getting modified, there interactions often limit these predictions. must be a rapid increase in the graph curve. 1 Master Layout (Part 4) This animation consists of 5 parts: Part 1 – Thermodynamics of protein folding Part 2 – Anfinsen’s experiment Part 3 – Amino acid structure determines 3-D folding Part 4 – Molecular chaperones for protein folding Part 5 – Protein misfolding diseases 2 2Pi DnaJ Unfolded protein 3 DnaK To GroEL system 4 Folded native conformation protein Partially folded protein GrpE ADP + GrpE + DnaJ 5 Source: Biochemistry by Lehninger, 4th edition (ebook) 1 Definitions of the components: Part 4 – Molecular chaperones for protein folding 1. Unfolded protein: This refers to the protein or polypeptide chain that has not been folded or is in a partially folded state. 2 3 2. DnaJ and Dna K: These are molecular chaperones found in E.coli that are analogous to the eukaryotic heat shock protein (Hsp) chaperone system. These chaperones are proteins that interact with unfolded or partially folded proteins and provide them with suitable microenvironments in which folding can occur. In addition to this chaperone system, the Hsp proteins have also been studied and have been found in abundance in cells that have been stressed by elevated temperatures. 3. ATP: Adenosine triphosphate (ATP) is the energy currency of the cell due to its high energy phosphate bonds. It gets hydrolyzed to liberate adenosine diphosphate (ADP) and a phosphate group (Pi). 4 4. GrpE: This is a nucleotide exchange factor present in bacterial systems that facilitates the release of bound ADP. 5. GroEL system: The GroEL system refers to another group of elaborate protein complexes known as chaperonins that assist the folding of several cellular proteins. 5 1 Part 4, Step 1: ATP DnaJ 2 ATP Unfolded protein ATP DnaK ATP 3 ADP ADP 4 Action Description of the action 5 Pi Blue squares and purple figures must bind to the red ribbon. After this, colour of circle must change as shown below. (Please redraw all figures.) First show the red ribbon binding to the blue squares followed by purple figure (DnaK). The blue squares must interact with the circles which must then change colour as shown in the bottom figure and the arrow stemming out of the downward arrow must appear. th Audio Narration The unfolded protein is bound by DnaJ and then by DnaK which is an ATP bound protein. The hydrolysis of ATP into ADP and Pi by DnaK is stimulated by DnaJ. The resulting DnaK-ADP remains tightly bound to the unfolded protein. 1 Part 4, Step 2: ADP ADP GrpE 2 ADP ADP ADP 3 ADP DnaJ DnaK 4 Action Description of the action 5 The green pie objects must remove the grey circles from the attached rectangle. (Please redraw all figures.) First show the figure on top left. Then show the green pie shapes moving to the grey circles such that they must remove them from the attached rectangle. The blue squares must also get detached resulting in the figure at the bottom. Source: Biochemistry by Lehninger, 4th edition (ebook) Audio Narration The nucleotide exchange factor GrpE present in bacteria facilitates release of ADP along with DnaJ. This leaves the DnaK bound to the partially folded protein which continues to undergo folding to a more favorable low energy conformation. 1 Part 4, Step 3: 2 ATP ATP ATP Partially folded protein 3 DnaK Folded native protein To GroEL system 4 Action Description of the action 5 DnaK regenerated for next round of protein folding The red ribbon must detach from purple square which must again bind the yellow circle. (Please redraw all figures.) First show the red ribbon being detached from the purple squares followed by binding of yellow circles to the purple squares. Audio Narration Once the protein gets completely folded, it gets detached from DnaK which then binds ATP again, thereby completing the cycle and preparing it for the next round of protein folding. Any protein which may not have been folded completely is then taken over by the GroEL chaperonin system which completes the folding. Source: Biochemistry by Lehninger, 4th edition (ebook) 1 2 3 4 5 Master Layout (Part 5) This animation consists of 5 parts: Part 1 – Thermodynamics of protein folding Part 2 – Anfinsen’s experiment Part 3 – Amino acid structure determines 3-D folding Part 4 – Molecular chaperones for protein folding Part 5 – Protein misfolding diseases Alzheimer’s disease Huntington’s disease Creutzfeldt– Jakob disease Cystic fibrosis Pulmonary emphysema Lathyrism 1 Definitions of the components: Part 5 – Protein misfolding diseases 1. Alzheimer’s Disease: 2 3 Structure of certain normal soluble cellular proteins normally rich in alpha helical regions converted into beta strand conformations which further link with each other to form beta sheet aggregates known as amyloids. Insoluble amyloid plaques are essentially made up of a single polypeptide chain or fibrils known as amyloid-b-protein (Ab). Observed in the brain of patients with Alzheimer’s where dead or dying neurons surround plaques. Neurotoxicity believed to be caused by the Ab fibrils before they get deposited as amyloid plaques. The disease presents various symptoms such as memory loss, decreased neuromuscular coordination, confusion and dementia. 2. Huntington’s disease: 4 5 Neurodegenerative disorder of genetic origin affecting muscular coordination. Caused by increased number of trinucleotide repeats, CAG, in Huntingtin gene leading to increased number of glutamine residues incorporated in corresponding protein. This alters the folding of the Huntington protein which has highest concentration in brain and testes. Exact function of the protein is unclear but is known to interact with several other proteins. Mutated protein has also been found to have effects on chaperone proteins which in turn help in folding several other proteins. Prominently affects basal ganglia which plays a key role in movement and behavioural control. 1 2 3 4 5 Definitions of the components: Part 5 – Protein misfolding diseases 3. Creutzfeldt–Jakob disease: Initially believed to be caused by viruses or bacteria. Later discovered to be transmitted by small proteins known as prions. Prion proteins composed of beta sheet structures that have been modified from previously existing alpha helices. Protein aggregates of one abnormal protein sufficient to function as a nuclei for other normal proteins to attach themselves to. Characterized by muscular spasms, loss of muscle control and memory loss. 4. Cystic fibrosis: Autosomal recessive disorder caused by a mutation in gene for the protein cystic fibrosis transmembrane conductance regulator (CFTR) . CFTR regulates components of sweat, digestive juices and mucus. Caused by a deletion of three nucleotides leading to the elimination of a phenylalanine residue from the protein and therefore abnormal folding. Dysfunctional protein gets degraded by the cell. Disorder can affect several body parts such as the lungs, GI tract and reproductive organs. 1 Definitions of the components: Part 5 – Protein misfolding diseases 5. Pulmonary emphysema: 2 3 4 5 Progressive disease of the lung causing shortness of breath. Can be caused by deficiency of the protein alpha-1-antitrypsin (A1AT). A1AT is responsible for protecting the lung tissues from damage by enzyme neutrophil elastase. Abnormally secreted A1AT gets accumulated in the liver thereby allowing lung tissue damage. Causes wheezing, shortness of breath, asthma-like symptoms and also liver cirrhosis. 6. Lathyrism: Regular ingestion of seeds from sweet pea (Lathyrus odoratus) causes disruption of cross-linking in the muscle protein, collagen. Collagen is an important structural protein having a triple helical structure. Cross-links formed are due to the oxidation of lysine residues by the enzyme lysyl oxidase to form allysine. These are essential for proper folding of collagen, giving it the required strength. b-aminopropionitrile, present in abundance in sweet pea, deactivates this enzyme by binding to its active site This prevents cross-linking and proper folding of the protein. Causes muscle fragility and weakness. 1 Part 5, Step 1: Alzheimer’s disease Huntington’s disease Creutzfeldt– Jakob disease Cystic fibrosis 2 Pulmonary emphysema Lathyrism 3 4 Action Description of the action 5 User should be allowed to click on any of the given labels to understand more about it. User should be allowed to click on any of the given labels to understand more about it as given by the definitions in the previous three slides. Audio Narration <As given in the definitions slides.> 1 Interactivity option 1:Step No:1 2 3 Protein folding is an extremely quick process with a time scale of few milliseconds to seconds. Monitoring the process is therefore a formidable task requiring very rapid measurements to be made. One of the suitable techniques for this measurement is the pulsed hydrogen-deuterium (H/D) exchange reaction. This relies on the differential measurement of hydrogen and deuterium which give different signals by a particular spectroscopic technique. Which technique is this? a) UV spectroscopy b) NMR c) ESR d) Mass spectroscopy 4 Interacativity Type 5 Choose the correct option. Options User has to choose one of the four options. If a, c or d are chosen, they must turn red. User can however continue till he gets the right answer (b) which must turn green. User is then directed to step 2. Boundary/limits Results User has to choose one of the four options. If a, c or d are chosen, they must turn red. User can however continue till he gets the right answer (b) which must turn green. User is then directed to step 2. Interactivity option 1:Step No:2 Deuterium labelled peptide N atoms Urea D2O Guanidinium chloride Denatured protein Native protein with with deuterated regular hydrogen peptide nitrogen atoms atoms. Light source Stopping syringe Denatured protein solution Mixer Denaturant solution with H2O Detector Stopped -flow device Computer Stop switch Interactivity option 1:Step No:3 Light source Stopping syringe Denatured protein solution Stop switch Folding initiated in denatured protein by mixing with denaturant diluted H2O and decreasing pH to arrest exchange reaction. Mixer Denaturant solution with H2O Detector Deuterium label Computer Folding for time ‘t’ Not involved in H-bonding Low pH – no exchange reaction Denatured protein pH increased Regular hydrogen from H2O Folding takes place for preset time t, after which pH is rapidly increased again for exchange reaction to occur. Only those D atoms that have not been involved in hydrogen bonding by time t will get exchanged now. H-D exchange reaction pH lowered finally to terminate labelling pulse and H/D ratio at each exchangeable site determined by 2-D NMR. 1 Questionnaire 1. If the free energy of a reaction is negative, the reaction will be 2 Answers: a) Endothermic b) Exothermic c) Non-spontaneous d) Spontaneous 2. b-mercaoptoethanol is responsible for breaking which type of interactions in proteins? Answers: a) Disulphide b) Hydrophobic c) Hydrogen bonds d) Peptide bond 3 3. In the absence of b-mercaoptoethanol but presence of urea what happens to a denatured protein? Answers: a) It gets further denatured b) Random disulphide links are formed c) The native state is restored d) Only non-covalent interactions are restored 4 4. Which of the following is a trinucleotide repeat disorder? Answers: a) Cystic fibrosis b) Alzheimer’s c) Huntington’s d) Lathyrism 5. Which enzyme is inactivated in the disease lathyrism? 5 Answers: a) Hexokinase b) Lysyl oxidase c) Collagenase d) Prolyl hydroxylase Links for further reading Books: Biochemistry by Stryer et al., 5&6th edition Biochemistry by A.L.Lehninger et al., 4th edition Biochemistry by Voet & Voet, 3rd edition