* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Energetics and kinetics of protein folding Comparison to other self
G protein–coupled receptor wikipedia , lookup
Bottromycin wikipedia , lookup
Theories of general anaesthetic action wikipedia , lookup
Protein moonlighting wikipedia , lookup
List of types of proteins wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Genetic code wikipedia , lookup
Western blot wikipedia , lookup
Protein (nutrient) wikipedia , lookup
Interactome wikipedia , lookup
Homology modeling wikipedia , lookup
Biosynthesis wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Circular dichroism wikipedia , lookup
Expanded genetic code wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Proteolysis wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Protein domain wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Biochemistry wikipedia , lookup
Protein adsorption wikipedia , lookup
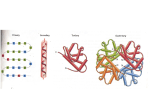
Energetics and kinetics of protein folding QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Comparison to other self-assembling systems? The search for the conformational energy minimum is global not a combination of multiple independent local searches Traveling salesman problem a classic example of a global search problem The conformational energy landscape describes the relative energy of all possible conformational states of a molecule QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Two-dimensional energy landscape Frustrated systems QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. The thermodynamic hypothesis Quic kTime™ and a TIFF (Unc ompres sed) decompress or are needed to see this picture. The native structure of a protein is determined solely by the sequence of amino acids in its poly-peptide chain and represents the state of the lowest conformational energy under native conditions. Christian Anfinsen QuickT ime ™an d a TIFF ( Uncomp res sed) deco mpre ssor ar e need ed to see this pictur e. Nobel Prize in Chemistry 1972 Solving the energetic problem Make more stabilizing interactions than you break 5-15 kcal + Hydrophobic effect Conformational entropy Hydrogen bonds Electrostatic interactions Van der Waals interactions very small Number With increasing size the ratio of surface to buried residues decreases 5 hydrophilic 0 hydrophobic 11 hydrophilic 4 hydrophobic 16 hydrophilic 14 hydrophobic Entropic cost per amino acid 0.6kcal/mol*ln10=1.3 kcal/mol Average number of methyl groups per hydrophobic amino acid 5 Hydrophobic effect per buried amino acid = 5*0.8 kcal/mol=4 kcal/mol ---> ~ 1/3 of amino acids need to be buried V=4/3 r3 x=2 ; y=4 n= 64 ratio = 8 x=3 ; y=5 n=125 ratio = 4.6 x=4 ; y=6 n=216 ratio = 3.4 4/3 x3 = hydrophobic total 4/3 y3 QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Using the hydrophobic effect to evade immune Detection: Neisseria gonorrhoeae pilin Parge et al. 1995 Nature Using the hydrophobic effect to evade immune Detection: Neisseria gonorrhoeae pilin QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Levinthal’s paradox QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Cyrus Levinthal Proteins have an incredible number of possible conformational states, yet they are able to fold very quickly. Levinthal’s paradox 150 AA domain 10 different conformations per side chain -------> 10150 possible conformations. Atomic vibrations occur on fsec. (10-15 sec) time scale Time to search all possible conformations >> age of the universe Number of particles in the universe ~1085 Levinthal’s paradox Random protein sequences typically do not fold, neither do most other polymers. There must be a code / mechanism That allows proteins to fold within a reasonable Period of time. Two models: Frame work Global collapse Framework model Secondary structure elements form first Packing of secondary structure elements leads to molten globule Repacking of the core in the molten globule leads to native structure Global collapse model QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. - Protein collapses due to hydrophobic effect - Hydrophobic environment drives formation of secondary structure to form molten globule - Repacking of molten globule leads to native structure Phi value analysis Alan Fersht Phi value =1 interaction already exists in transition state Phi value =0 interaction is not part of transition state Current state of debate A detailed understanding of protein folding remains illusive because we still lack experimental information on many of the states along the folding trajectory The transition state of a two-state folder tends to be very compact. Proteins with similar folds tend to fold following a similar mechanism.