* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download GenomeSequencing_ver3_20040929
Nucleic acid analogue wikipedia , lookup
Multilocus sequence typing wikipedia , lookup
Molecular Inversion Probe wikipedia , lookup
Molecular cloning wikipedia , lookup
Genetic engineering wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Point mutation wikipedia , lookup
Transposable element wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Personalized medicine wikipedia , lookup
Copy-number variation wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Molecular ecology wikipedia , lookup
Non-coding DNA wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
DNA sequencing wikipedia , lookup
Community fingerprinting wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Exome sequencing wikipedia , lookup
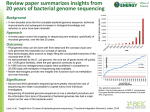
Genome Sequencing: Technology and Strategies Chuong Huynh NIH/NLM/NCBI [email protected] Acknowledgement: Daniel Lawson (Sanger Institute) and Jane Carlton (TIGR) Bioinformatics Flow Chart 1a. Sequencing 1b. Analysis of nucleic acid seq. 2. Analysis of protein seq. 3. Molecular structure prediction 6. Gene & Protein expression data 7. Drug screening Ab initio drug design OR Drug compound screening in database of molecules 4. molecular interaction 8. Genetic variability 5. Metabolic and regulatory networks How to sequence a genome • development of sequencing strategy and source of funding • procurement of DNA and initial library construction • test sequencing • large-scale random sequencing of small (2-3 kb), medium (10 kb) and large (>50 kb) libraries • analysis of raw sequence data by: BLAST, RepeatFinder etc • release of genome data onto sequencing center website • at 8-10 X coverage, random stops • closure of sequence gaps and physical gaps • comparison to physical map • gene model prediction • final gene model annotation • release of data to GenBank and publication Full shotgun sequencing Genomic DNA Marker1 Marker2 large insert library (20 - 500 kb) Minimal tiling path shotgun library: small (2-3 kb) and medium (10 kb) Sequencing (8-10 X) Assembly scaffold contig Gap closure gene prediction, annotation and analysis Partial shotgun sequencing Genomic DNA shotgun library: small (2-3 kb) and medium (10 kb) Sequencing (5X) Assembly contig scaffold Analysis Genome sequencing terms Raw sequence: unassembled sequence reads produced from sequencing of inserts from individual recombinant clones of a genomic DNA library. Finished sequence: complete sequence of a genome with no gaps and an accuracy of > 99.9%. Genome coverage: average number of times a nucleotide is represented by a high-quality base in random raw sequence. Full shotgun coverage: genome coverage in random raw sequence required to produce finished sequence, usually 8-10 fold (‘8-10X’). Partial shotgun coverage: typically 3-6X random coverage of a genome which produces sequence data of sufficient quality to enable gene identification but which is not sufficient to produce a finished genome sequence Paired reads: sequence reads determined from both ends of a cloned insert in a recombinant clone. Contig: contiguous DNA sequence produced from joining overlapping raw sequence reads. Singleton: single sequence read that cannot be joined (‘assembled’) into a contig. Scaffold: a group of ordered and orientated contigs known to be physically linked to each other by paired read information. EST: expressed sequence tag generated by sequencing one end of a recombinant clone from a cDNA library. ESTs are single-pass reads and therefore prone to contain sequence errors. GSS: genome survey sequence generated by sequencing one end of a recombinant clone from a genomic DNA library. The genomic DNA library can in some instances be enriched for the presence of coding regions, for example through use of mung bean nuclease digestion of genomic DNA prior to cloning. SNP: single nucleotide polymorphism ORF: open reading frame, stretches of codons in the same reading frame uninterrupted by STOP codons and calculated from a six-frame translation of DNA sequence. Jan 2003 NCBI Trace Archive Sep 23, 2003 Large-scale genome projects • Sequencing DNA molecules in the Mb size range • All strategies employ the same underlying principles: Random Shotgun sequencing Strategy Libraries Sequencing Assembly Closure Annotation Release Genomic DNA Shearing/Sonication Subclone and Sequence Shotgun reads Assembly Contigs Finishing read Finishing Complete sequence Strategies for sequencing • How big can you go?? Strategy • Large-insert clones Libraries • cosmids 30-40 kb • BACs/PACs 50 - 100 kb • Whole chromosomes • Whole genomes Sequencing Assembly Closure Annotation Release Genome size and sequencing strategies Genome size (log Mb) 0 1 2 3 4 H.sapiens (3000 Mb) D.melanogaster (170 Mb) C.elegans (100Mb) P.falciparum (30 Mb) S.cerevisiae (14 Mb) E.coli (4 Mb) Whole genome shotgun (WGS) Clone-by-clone Whole Chromosome Shotgun (WCS) Whole Genome Shotgun (WGS) with Clone ‘skims’ Genomic DNA Shearing/Sonication Subclone and Sequence Shotgun reads Assembly Contigs Finishing read Finishing Complete sequence Strategies for sequencing • Size and GC composition of genome • Volume of data • Ease of cloning • Ease of sequencing • Genome complexity Strategy Libraries Sequencing Assembly Closure • dispersed repetitive sequence Annotation • telomeres & centromeres Release • Politics/Funding Strategies: Clone by Clone • Simple (0.5 - 2 K reads) Strategy • Few problems with repeats Libraries • Relatively simple informatics • Scalability • Quality of physical map • Fingerprint / STS maps • End sequencing Sequencing Assembly Closure Annotation Release Strategies: Whole Chromosome shotgun (WCS) • Requires chromosome isolation Strategy • Moderate complexity (10’s K reads) Libraries • Problems with repeats • Complex informatics • Inefficient in isolation • Quality of physical map (want good physical map) • Skims of mapped clones Sequencing Assembly Closure Annotation Release Strategies: Whole Genome shotgun (WGS) • Moderate to High complexity (10-100’s K reads) Strategy • Massive Problems with repeats Libraries • Complex informatics • Quality of physical map • Fingerprint map • STS markers • End-sequences • Skims of mapped clones Sequencing Assembly Closure Annotation Release Sequencing my genome Politics Strategy Libraries Production Sequencing Assembly Finishing Closure Annotation Annotation Release TIME MONEY What do you get? DATA!!, DATA !!, and more DATA!! Strategy • Sequence • incomplete complete Libraries Sequencing • First-pass annotation Assembly • Gene discovery Closure • Full annotation Annotation • A starting point for research Release Genome annotation is central to functional genomics ORFeome based functional genomics RNAi phenotypes Gene Knockout Expression Microarray Where is the problem? Most genome will be sequenced and can be sequenced; few problem are unsolvable. Problems lies in understanding what you have: gene prediction annotation Sequencing • Library construction Strategy • Colony picking (random) Libraries • DNA preparation (isolate DNA) • Sequencing reactions • Electrophoresis • Tracking/Base calling Sequencing Assembly Closure Annotation Release Libraries • Essentially Sub-cloning • Generation of small insert libraries in a well characterised vector. • Ease of propagation • Ease of DNA purification • e.g. puc18, M13 Strategy Libraries Sequencing Assembly Closure Annotation Release Libraries - testing • Simple concepts • Insert/Vector ratio (Blue/White ratio) • Real data • Insert size • Sequence …. • Simple analysis Strategy Libraries Sequencing Assembly Closure Annotation Release Sequence generation • Pick colonies growth medium •Template preparation (DNA isolation) • Sequence reactions • Standard terminator chemistry • pUC libraries sequenced with forward and reverse primers •Tracking and noise Strategy Libraries Sequencing Assembly Closure Annotation Release Sequence generation • Electrophoresis of products • Old style - slab gels, 32 > 64 > 96 lanes • New style - capillary gels, 96 lanes • Transfer of gel image to UNIX • Sequencing machines use a slave Mac/PC • Move data to centralised storage area for processing Strategy Libraries Sequencing Assembly Closure Annotation Release Gel image processing • Light-to-Dye estimation • Lane tracking • Lane editing • Trace extraction • Trace standardisation • Mobility correction • Background substitution Strategy Libraries Sequencing Assembly Closure Annotation Release Pre-processing • Base calling using Phred • modifies SCF file format • Quality clipping from Phred • Vector clipping • Sequencing vector • Cloning vector • Screen for contaminants • Feature mark up (repeats/transposons) Strategy Libraries Sequencing Assembly Closure Annotation Release Finishing • Assembly: Process of taking raw single-pass reads into contiguous consensus sequence (Phred/Phrap) • Closure: Process of ordering and merging consensus sequences into a single contiguous sequence • Finished is defined as sequenced on both strands using multiple clones. In the absence of multiple clones the clone must be sequenced with multiple chemistries. The overall error rate is estimated at less than 1 error per 10 kb Strategy Libraries Sequencing Assembly Closure Annotation Release Genome Assembly Strategy • Pre-assembly (assembly algorithm) • Assembly • Automated appraisal • Manual review Libraries Sequencing Assembly Closure Annotation Release Pre-Assembly Strategy • Convert to CAF format • flatfile text format • choice of assembler • choice of post-assembly modules • choice of assembly editor Libraries Sequencing Assembly Closure Annotation Release www.sanger.ac.uk/Software/CAF Assembly Strategy • Assemble using Phrap • Read fasta & quality scores from CAF file • Merge existing Phrap .ace file (previous assembly) as necessary • Adjust clipping (where vector, quality start) Libraries Sequencing Assembly Closure Annotation Release Assembly appraisal • auto-edit • removes 70% of read discrepancies of seq. assembly (highlight misassembly); manually • Remove cloning vector • Mark up sequence features (for finisher) • “Finish” Program (or Program “AutoFinish”) • Identify low-quality regions • Cover using ‘re-runs’ and ‘long-runs’ • Compare with current databases • plate contamination Strategy Libraries Sequencing Assembly Closure Annotation Release Manual Assembly appraisal Strategy • Use a sequence editor (GAP/consed) • Tools to identify Internal joins • Tools to identify and import data from an overlapping projects • Tools to check failed or mis-assembled reads for inclusion in project Libraries Sequencing Assembly Closure Annotation Release Manual editing • Sanger uses 100% edit strategy Strategy • Where additional data is required: Libraries • Check clipping • Additional sequencing • Template / Primer / Chemistry • Assemble new data into project • GAP4 Auto-assemble • Repeat whole process Sequencing Assembly Closure Annotation Release Manual Quality Checks • Force annotation tag consistency • All unedited data is re-assembled using Phrap Strategy • All high-quality discrepancies are reviewed Libraries • Confirm restriction digest (clones) • Check for inverted repeats • Manually check: • Areas of high-density edits • Areas with no supporting unedited data • Areas of low read coverage (need to confirm) Sequencing Assembly Closure Annotation Release Gap closure • Read pairs Strategy • PCR reactions (long-range / combinatorial) Libraries • Small-insert libraries Sequencing • Transposon-insertion libraries Assembly Closure Annotation Release Gap closure - contig ordering • Read pair consistency Strategy • STS mapping Libraries • Physical mapping Sequencing • Genetic mapping • Optical mapping • Large-insert clone • skims • end-sequencing Assembly Closure Annotation Release Annotation • DNA features (repeats/similarities) Strategy • Gene finding Libraries • Peptide features • Initial role assignment • Others- regulatory regions Sequencing Assembly Closure Annotation Release Annotation of eukaryotic genomes Genomic DNA ab initio gene prediction transcription Unprocessed RNA RNA processing Mature mRNA Gm3 AAAAAAA translation Nascent polypeptide Comparative gene prediction folding Active enzyme Functional identification Function Reactant A Product B Genome analysis overview: C.elegans DNA features • Similarity features • mapping repeats • simple tandem and inverted • repeat families • mapping DNA similarities Strategy Libraries Sequencing Assembly • EST/mRNAs in eukaryotes Closure • Duplications, Annotation • RNAs Release • mapping peptide similarities • protein similarities Gene finding • ORF finding (simple but messy) Strategy • ab initio prediction Libraries • Measures of codon bias • Simple statistical frequencies • Comparative prediction • Using similarity data • Using cross-species similarities Sequencing Assembly Closure Annotation Release Peptide features • Peptide features • low-complexity regions • trans-membrane regions • structural information (coiled-coil) • Similarities and alignments • Protein families (InterPro/COGS) Strategy Libraries Sequencing Assembly Closure Annotation Release Initial role assignment • Simple attempt to describe the functional identity of a peptide • Uses data from: • peptide similarities • protein families • Vital for data mining • Large number of predicted genes remain hypothetical or unknown Strategy Libraries Sequencing Assembly Closure Annotation Release Other regulatory features • Ribosomal binding sites Strategy • Promoter regions Libraries Sequencing Assembly Closure Annotation Release Data Release • DNA release • Unfinished Strategy • Finished Libraries • Nucleotide databases • GENBANK/EMBL/DDBJ • Peptide databases • SWISSPROT/TREMBL/GENPEPT • Others Sequencing Assembly Closure Annotation Release Real World Example: Malaria Genome Project If time permits. Sequencing the Plasmodium genomes Four species of malaria infect man: Plasmodium falciparum P. vivax P. malariae P. ovale Four species of malaria infect rodents: P. yoelii P. berghei P. chabaudi P. vinckei Plasmodium falciparum ~30 million base pairs (Mb) 80% (A+T) 14 chromosomes DNA “unstable” in E. coli No large insert DNA clones suitable for sequencing Too large for whole genome shotgun (‘96) Whole chromosome shotgun strategy was selected Comparison of genome features Feature P.falciparum P.y.yoelii Size (Mb) No. chroms Coverage (fold) No. gaps (G+C) content (%) No. genes Mean gene length (bp) Gene density (bp/gene) Genes with introns (%) Genes with ESTs (%) Genes with proteomic data (%) Exons: Mean no./gene (G+C) content (%) Introns: (G+C) content 23.1 14 5 5,812 22.6 5,878 1,298 2,566 54.2 48.9 18.2 2.0 24.8 21.1 22.9 14 14.5 93 19.4 5,268 2,283 4,338 53.9 49.1 51.8 2.4 23.7 13.5 Intergenic sequences: (G+C) content RNAs: no. tRNAs no. 5s rRNAs no. rRNA units 20.7 39 3 4 13.6 43 3 7 P. falciparum genome status Chr Size (bp) No. gaps Fold coverage 1 643,293 0 13.3 2 (TIGR) 947,102 0 11.1 3 1,060,087 0 10.9 4 1,204,112 0 16.8 5 1,343,552 0 15.1 6 1,377,956 8 16.8 7 1,350,452 14 15.8 8 1,323,195 24 16.2 9 1,541,723 0 17.9 10 (TIGR) 1,694,445 4 15.6 11 (TIGR) 2,035,250 3 11.3 12 (Stanford) 2,271,477 0 16.3 13 2,747,327 37 17.2 14 (TIGR) 3,291,006 3 9.2 0 22,788 0 ND 22,853,764 93 14.5 Eukaryotic annotation - TIGR Project DB Annotation Station/Manatee Annotation DB DDS/DPS EGC Gene finders Gene models BLAST PFAM/TIGRFAM SignalP/TMHMM Alignments of genomic to Functional proteins and ESTs assignments PFB0680w The P. falciparum genome P. falciparum S. pombe Size (bp) S. cerevisiae D. discoideum A. thaliana 22,853,764 12,462,637 12,495,682 19.4 36.0 38.3 22.2 34.9 No. of genes 5,268 4,929 5,770 2,799 25,498 Mean gene length* (bp) 2,283 1,426 1,424 1,626 1,310 Gene density† 4,338 2,528 2,088 2,600 4,526 Percent coding 52.6 57.5 70.5 56.3 28.8 Genes with introns (%) 53.9 43 5.0 68 79 43 174 ND 73 ND No. 5S rRNA genes 3 30 ND NA ND No. rRNAs units 7 200-400 ND NA 700-800 (G+C) content (%) No. tRNA genes *excluding introns; †bp per gene 8,100,000 115,409,949 Distribution of gene lengths 3000 P. falciparum S. pombe Number of genes 2500 S. cerevisiea 2000 1500 1000 15.5% 3.0-3.6% 500 0 < 300 300-999 1000-1999 2000-2999 3000-3999 Gene length (bp, excluding introns) >4000 The P. falciparum proteome Feature Number Per cent Total predicted proteins 5,268 Hypothetical proteins 3,208 60.9 InterPro matches 2,650 52.8 PFAM matches 1,746 33.1 Process 1,301 24.7 Function 1,244 23.6 Component 2,412 45.8 Targeted to apicoplast 551 10.4 Targeted to mitochondrion 246 4.7 Transmembrane domain(s) 1,631 31.0 Signal peptide 544 10.3 Signal anchor 367 7.0 Non-secretory proteins 4,357 82.7 Gene Ontology™ Structural features Florens et al. Nature 419:520-526 52% of predicted gene products detected by proteomics Metabolism and transport Analysis based on similarity searches with sequences of known enzymes 14% (733) of genes encoded enzymes Lower than in bacterial genomes (25-33%) Enzymes more difficult to identify due to AT-rich genome and evolutionary distance between P.f. and other sequenced organisms Or P.f. has smaller proportion of genome devoted to enzymes, reduced metabolic potential A T P A DP (13) H+ C a2+ H+ Zn2+ A T P A DP A T P A DP A T P A DP P Pi (16) N OVEL INHIBITORS N OVEL INHIBITORS PROTEASE INHIBITORS Large peptides H+ H+ H+ glucosamine riboflavin dephosphoCoA aspartate CoA oxaloac etate CO 2 malate L-LACTATE MITOCHONDRION DHF Sulfonamides or NADH Purines and Pyrimi dine s NAD+ ATP N OVEL INHIBITORS orotate RNA DNA ornithine N-acetyl-glutamate cysteine alanine spermidine methionine salvagepathway dTMP Cy tc Fe2+ O2 UQH 2 Cy tc Fe3+ H2O THF Pyrimethamine Cy cloguany l Shikimic Acid Pathway chor ism ate pABA DOXP Pathway pyruvate acetyl-CoA UQ deoxy xy lulose-5P Fos midomy cin 2C-methy lerythrose-4P Haem C Haem A c is -ac onitate Pr oto hae m (FPIX 2+) s uc cinate oxoglutarate s uc ciny l-CoA ubiquinonepool DHF Pu rine s alvag e, Pyr im id ine syn thesis APICOPLAST Tricarboxylic acid fumarate cycle isocitrate UQ Methy lene THF PRPP pyruvate malate Atovaquone or dihydroorotate CDP dUMP AMP proline serine PEP acetyl-CoA oxaloac etate c itrate malate pABA 7,8-dihydropteroate IMP (6) putrescine N OVEL x ylulos e-5P INHIBITORS + erythros e-4P 3-deoxy arabinoheptulos anate7-phos phate acetate oxaloac etate dCDP glutamate ribose-5P N OVEL INHIBITORS Pyrimethamine Cy cloguany l hypox anthine aspartate glutamine NOVEL INHI BITOR S THF XMP Pi Folate biosynthes is CO 2 Folate Biosynthesis x anthine asparagine ornithine ribulose-5P dihydrox yacetone-P + gly ceraldehy de-3P oxo acid amino acid FAD 6-phos phogluconate fructose-1,6-bis P GLYCEROL GMP Pi fructose-6P glucosamine-6P Haemozoin FMN (2) gly cine glucose-6P glucosamine-1P guanine ? ? Amino ac ids Chloroquine Artemes inin Quinine GDP di /t ri A T P A DPcarboxylate s (4) Pentose Phosphate Pathway GLUCOSE Small peptides (3) Glycolysis glucose-1P GTP H+ Pi Am ino Co m pou nds my o-inositol-1P FPIX 2+ O2- FPIX 3+ (2) glycosyl ph ophatidylin osito l (GPI an cho rs ) FOOD VACUOLE O2 H+ Pi (2) N OVEL INHIBITORS Haem o glob in H+ SO 2-4 H+ ? H+ Pi P (2) PROTEASE INHIBITORS H+ Mn2+ sugar phosphates V H+ N a+ water/ glycero l PEP F ? H+ mitochondrial/plastidcarriers sugar H+ nucleosid e/base glucose H+ nucleotide ornt-sugar? metabolites H+ dru gs? 2+, P -l ipi ds, Cu oth ercations? carboxylates? F, V, & P- type ATPases drugs? ABC tr ansporter s Haem Biosynthesis malony l-CoA acetoacety l-ACP + malonyl-ACP Glyce ro lipids acy l- ACP Tric losan Glycerolipid Metabolism gly cerol triac ylglyc erol c holine phosphatidylcholine m o difie d tRNAs FattyAcid Biosynthesis Thiolac tomyc in ALA ALA gly cine ethanolamine isopenteny l-PP porphobilinogen enoyl-ACP N OVEL INHIBITORS phosphatidylethanolamine 3-ox oacy l-ACP Fatty acid elonga tion 3-hydrox yacy l-ACP Analysis of transporters in P. falciparum Organization of multi-gene families in P. falciparum P. falciparum Genome Summary Feature Value Comments Genome size 24 million base pairs 1% of the human genome Number of chromosomes 14 23 pairs Number of gaps 93 (0-37 per chr) Genome >98% complete (A+T) content ~ 80.6% Number of genes ~5,300 Proteins of unknown function 60% Most (A+T) rich genome sequenced to date Yeast: 5,770 Human: ~35,000 More than other genomes Possible surface proteins ~900 Test for use in vaccines Gene products detected 52% by proteomics Genes conserved in rodent 60% malaria P. yoelii yoelii See Florens et al. See Lasonder et al. See Carlton et al. Extra Slides