* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download MICROEVOLUTION
Therapeutic gene modulation wikipedia , lookup
Gene therapy wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Gene desert wikipedia , lookup
Genomic imprinting wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Genome (book) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genetic engineering wikipedia , lookup
Gene nomenclature wikipedia , lookup
Koinophilia wikipedia , lookup
Gene expression programming wikipedia , lookup
Pharmacogenomics wikipedia , lookup
History of genetic engineering wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Human leukocyte antigen wikipedia , lookup
Designer baby wikipedia , lookup
Human genetic variation wikipedia , lookup
Population genetics wikipedia , lookup
Genetic drift wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
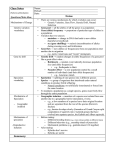
MICROEVOLUTION Materials: 3 colors of beans, totaling 4000 for each group. Each group has 49% one color, 42% the second color and 9% the 3rd color. Purpose: To simulate the microevolution model with populations of colored beans, illustrating random mating and the effects of selection and genetic drift. Background: Populations, not individuals, evolve by gradual changes over time in the frequency of alleles that are found at genetic loci. These changes result from mutation, selection, migration, or genetic drift. Collectively, these processes comprise microevolution. Mechanisms of microevolution are well understood. In fact, one of these mechanisms, selection, has been used for centuries to increase the productivity of crops and livestock. According to the microevolution model, a population of organisms can be considered to be a gene pool, which is composed of all the copies of every allele in the population at a given moment. In diploid organisms, the genes in the gene pool occur in pairs in each individual, and individuals may be homozygous or heterozygous for a particular trait. In 1908, G.H. Hardy and G. Weinberg showed that no matter how many generations elapsed, sexual reproduction in itself couldn’t change the frequencies of alleles in a gene pool. Changes if they occur, must be due to the action of other factors---the real agents of evolution. Example….a population of plants with red and white flowers. Red flowers have the dominant allele and white flowers are found in individuals with only the recessive alleles. A study of 10,000 plants indicated that 84% of the plants had red flowers and 16% had white flowers. Of the red flowered plants, 36% of the total plants were homozygous for the red allele and 48% were heterozygous. The genotypic frequencies were: 36% AA Where A = red allele 48% Aa a = white allele 16% aa 100% The frequencies of alleles can be obtained by the following: Every individual with the genotype AA contributes two A alleles to the gene pool, and every individual with the genotype Aa contributes one A allele so given 10000 individuals, the frequency of the A allele in the gene pool equals: 2 x 36% x 10,000 + 48% x 10,000 = 12,000 A alleles In the same population, the frequency a in the gene pool can be calculated by similar reasoning (aa individual contributes two a, and Aa contributes 1 a. Therefore, 2 x 16% x 10,000 + 48% x 10,000 = 8,000 a alleles On a percentage basis, 60% (12,000/12,000 + 8,000) of the alleles of the gene pool are A and 40% (8,000/ 12,000 + 8,000) are a. This information describes the gene pool at the time of the study. But, what happens to the allele frequencies when this population reproduces sexually? The frequency of the dominant allele is p and that of the recessive allele is q, in any population where there are only two alleles at one gene locus: P + q = 1 and p = 1 – q or q=1–p Sooo, a population in Hardy-Weinberg equilibrium, the homozygote dominant will be found p2% of the time, the heterozygote 2 pq% of the time and the homozyous recessive q2% of the time. (p2% … 36% AA ) (q2% … 16% aa) (2 pq% … 48% Aa) Lab: Your lab population is 4000 beans in a coffee can. There are 3 colors: 49% are one color, 42% another color and 9% a third color. These percentages correspond to genotypes in hypothetical populations in which there are two alleles at a given locus. Indicate what colors correspond to the genotypes below: Genotype % Color Number (% x 4000) AA 49 . Aa 42 . aa 9 . Determining starting allele frequency-Accounting method # of AA individuals x 2 =3920 +# of Aa individuals x 1 =1680 # of A alleles in gene pool =5600 Do the same for the a allele Determining starting allele frequency-Hardy-Weinberg Calculation method To use this there is the major assumption that the genotypes are in proportion to what is expected by the Hardy-Weinberg equilibrium. (which is that AA=p2 and aa=q2 ) Calculate the p and q values for the population in the can and enter it below. P=________ q=________ Remember p+q=1 Are the relative frequencies of A and a the same by the accounting and calculation methods?_____ 1. To simulate random mating in your population, mix up the can and withdraw two beans. These are your breeding pair. 2. Repeat this 100 times...recording the colors drawn each time. RETURN BEANS TO CAN AFTER EACH DRAW! (Why?) 3. Record the info on the table below and list the Genotypes of the new breeding group. AA______ Aa_______ aa_______ Total 200 Are these frequencies the same as the total population of the can? Compare them using p and q Mating Pairs (color x color) 1 2 3 4 5 6 Corresponding pair Tallys Total of tallys