* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Chapter 3 - Proteins
Mitogen-activated protein kinase wikipedia , lookup
Paracrine signalling wikipedia , lookup
Peptide synthesis wikipedia , lookup
Signal transduction wikipedia , lookup
Gene expression wikipedia , lookup
Expression vector wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Magnesium transporter wikipedia , lookup
Point mutation wikipedia , lookup
Protein purification wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Interactome wikipedia , lookup
Metalloprotein wikipedia , lookup
Western blot wikipedia , lookup
Genetic code wikipedia , lookup
Biosynthesis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Two-hybrid screening wikipedia , lookup
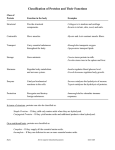
Chapter 3 - Proteins Test Your Knowledge About Basic Protein Structure • Name one polar and one nonpolar amino acid, then make a list of all the additional amino acids that you remember. • What are the four weak (noncovalent) interactions that determine the conformation of a protein? • (True/False) A protein is at a near entropy minimum (point of lowest disorder, or greatest order) when it is completely stretched out like a string and when it is properly folded up. Explain. • (True/False) Loops of polypeptide that protrude from the surface of a protein often form the binding sites for other molecules. Explain. • (True/False) For a family of related genes that do not match genes of known function in the sequence database, it should be possible to deduce their function using “evolutionary tracing” to see where conserved amino acids cluster on their surfaces. Explain. Also Pro, Phe, Met, Trp, Gly, Cys Reactions that promote protein folding Adapted from L. Wu et al., 1995, Nature Struc. Biol. 2:281; courtesy of J. Harris and P. S. Kim Molecular chaperones Chaperones Video – chaperone-aided protein folding • Average length ≈ 10 residues ≈ 15 Å. • Minimum length = 4 residues: how many H-bonds? • Maximum length = 40 residues. • Pleated - look edge-on • Strands ~ 5 Å apart • Note direction of H-bonds will differ in antiparallel & mixed sheets Loops & Turns •Connect secondary structural elements. •Loops often carry the functional groups. •Hairpin turns: Shortest possible loops (2 residues). •Gly often in tight turns. Motifs (protein folds) Beta-Hairpins I Beta-Hairpins II Beta Corners Beta Barrels Helix Hairpins Alpha-Alpha Corners E-F Hand Helix-Turn-Helix Beta-Alpha-Beta Motifs Greek Key Motifs Domains/Modules Pyruvate kinase • B sheet core with protruding loops • Loops for binding interactions • N and C terminals at opposite poles or form “plug-ins” Domain shuffling Families/Clans Pyruvate kinase This is a member of the Pyruvatekinase-likeTIM barrel superfamily Other families GP120 Family Envelope glycoprotein GP120 RVT_1 Family Reverse transcriptase (RNA-dependent DNA polymerase) COX1 Family Cytochrome C and Quinol oxidase polypeptide I Oxidored_q1 Family NADH-Ubiquinone/plastoquinone (complex I), various chains MFS_1 Family Major Facilitator Superfamily HCV_NS1 Family Hepatitis C virus non-structural protein E2/NS1 Serine Protease family Other Important Protein Structural Features • Subunits – Dimers, tetramers, large assemblies of monomers • Filamentous proteins • Globular proteins Video – clathrin assembly Table 1. Proportions of amino acids that are inaccessible to solvent in a study of twelve proteins. Amino Acid Side Chain Proportion Buried I = Ile 0.60 V=Val 0.54 C=Cys 0.50 F=Phe 0.50 L=Leu 0.45 M=Met 0.40 A=Ala 0.38 G=Gly 0.36 W=Trp 0.27 T=Thr 0.23 S=Ser 0.22 E=Glu 0.18 P=Pro 0.18 H=His 0.17 D=Asp 0.15 Y=Tyr 0.15 N=Asn 0.12 Q=Gln 0.07 K=Lys 0.03 R=Arg 0.01 Test Your Knowledge Now Small proteins may have only one or two amino acid side chains that are totally inaccessible to solvent. Even in large proteins, only about 15% of the amino acids are fully buried. A list of buried side chains from a study of twelve proteins is shown in Table 1. The list is ordered by the proportion of amino acids of each type that are fully buried. What types of amino acids are most commonly buried? Least commonly buried? Are there any surprises? If so, why?