* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Plant proteomics workshop_final072114
Survey
Document related concepts
Gene regulatory network wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Protein moonlighting wikipedia , lookup
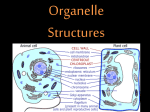
Cell membrane wikipedia , lookup
Protein adsorption wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Paracrine signalling wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Signal transduction wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Western blot wikipedia , lookup
Degradomics wikipedia , lookup
Transcript
Sample Preparation and Perspective on Proteomics Dhileepkumar Jayaraman Ané lab Department of Agronomy University of Wisconsin - Madison Plant proteomics workshop 07/21/2014 Proteomics “PROTEin complement expressed by a genOME” Identification all the proteins in a cell or organism Includes any PTMs, cellular localization, functions, and interactions 2/43 http://www.chem.purdue.edu/Tao How complex is the proteome? 3/43 http://www.piercenet.com Proteomic approaches Experimental material 4/43 (Gosh &Xhu, 2014) A decade of plant proteomics and mass spectrometry 5/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.21365 Where we stand? Articles Reviews Why? 6/43 (Pubmed data) Plant cells are enclosed within a rigid cell wall Cellulose microfibrils, an electron microscope view Phospholipids of the plasma membrane are amphipathic, containing both a polar (hydrophilic) head and a nonpolar (hydrophobic) tail. 7/43 Sample preparation considerations Compatible with subsequent steps leading to protein identification by MS. Extraction of proteins from a cell, tissue or organelle should be as complete as possible. Maximum no. of proteins in minimum no. of biochemical steps and minimum time. 8/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Extraction Buffer Composition Purposes of the Extraction Buffer 1. Dissolve cellular membranes 2. Inactivation of protease 3. Assist in the removal of contaminants Class of additive example concentration purpose Detergents 0.1-1% Solubilization of poorly soluble proteins 50-150mM Maintain ionic strength of medium Stabilize lysosymal membranes, reduce protease release Salts Deoxycholate, Triton X-100, SDS NaCl, KCl, (NH4)2SO4 Glucose or sucrose 25 mM Metal chelators EDTA, EGTA 1 mM Reducing agents DTT, DTE 1-10 mM 2-Mercaptoethanol 0.05% Reduce oxidation damage, chelate metal ions Reduce oxidation damage 9/43 www.embl.de Detergents and inhibitors Use of Detergents to Lyse Cells: Like Dissolves Like Mixed micelle Plasma membrane (phospholipid bilayer) Detergent molecules + SDS Protease inhibitor inhibition of Aprotinin serine proteases Protease inhibitors function by reversibly or irreversibly binding to the protease. Leupeptin cysteine and serine proteases PMSF serine proteases Pepstatin A aspartic proteases “Cocktail” is generally used Phosphatase inhibitor inhibition of Sodium Fluoride Ser/Thr and acidic phosphatase Sodium Othovanadate Tyr and alkaline phosphatase Sodium Pyrophosphate Ser/Thr phosphatase 10/43 www.embl.de Grinding methods Step 1: Disruption of cell walls by grinding Step 1+2: mechanical disruption and homogenization in extraction buffer Grind sample into a fine powder to shear cell walls and membranes Step 2: Lysis of cells in extraction buffer Mix thoroughly with extraction buffer to dissolve cell membranes and inhibit nuclease activity 11/43 A homogenizer allows cells to be mechanically disrupted within the extraction buffer Crude lysate Grinding methods considerations Disruption of cell walls by grinding Localized heating- leading to protein denaturation and aggregation. Pre-chill equipment and samples on ice at all times. Reproducibility. Cells disrupt at different times. 12/43 keep Grind sample into a fine powder to shear cell walls and membranes Plant organelle proteomics 13/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Organelle isolation Characterization of proteomes in different sub-cellular locations. understanding of plant functions, biosynthetic and signaling pathways. Sub-cellular fractionation Simplification of the proteome 14/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Organelle isolation....... (i) disruption of the CW and membrane (ii) fractionation of the crude homogenate. Fractionation-physical differences between organelles. Differential centrifugationenriched fractions of the organelle. Purification by density gradient centrifugation. 15/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Purity assessment of organelle Microscopy – sensitive and expensive Biochemical -less sensitive and cheap 16/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Organelle and structure specific chemical dyes (Hirayama etal.,2013, Nature) 17/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Organelle specific protein markers 18/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Proteomics of cell wall Less abundant and physically embedded in an insoluble polysaccharide matrix. Lithium chloride and CaCl2. Phosphotungstic acid (PTA) staining-free from contamination, Vandate sensitive H+ATPase activity. Cytosolic contamination: Catalase as marker enzyme. 19/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Proteomics of plasma membrane PM-associated proteins display a large diversity of physico-chemical properties. Salt treatments - integral membrane proteins. Alkaline treatments -lipid-anchored PM proteins. Organic solvents -hydrophobic proteins. 20/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Proteomics of cytosol Physically disrupt the plant cell wall but to maintain organelle integrity. Protoplasts isolation followed by disruption of protoplasts. Differential centrifugation. Purity testing- using markers for other organelles. Cytosolic markers cFBPase, TRXh3 and UGPase. 21/43 (Ito et al., 2011, J. proteome Res) Proteomics of nucleus Optimal nuclear preparations consist of homogenous membrane-bound forms with intact nucleoli. Contamination - DAPI staining. The nuclear proteins -high salt buffers. The enrichment for the nuclear-resident proteins-histone as marker. Further purity-antibodies directed against marker proteins for other cell compartments. Phase‐contrast image DAPI‐stained nuclear fraction 22/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Proteomics of mitochondria The majority of mitochondrial proteins encoded by the nucleus and synthesized as precursors in the cytosol before being targeted to mitochondria. Low abundance proteins identification is a challenge but they play a very important role. Mitotracker. 23/43 (Plant proteomics, Mtds&Protocols, 2014) Purification of protein complexes Most proteins exist as complexes. These macromolecular complexes should be purified with out losing their confirmations. Co-immunopurification. Affinity tag purification. 24/43 http://onlinelibrary.wiley.com/doi/10.1002/mas.20301 Our proteomics projects, past & present……… 25/43 Plant-microbe mutualisms Arbuscular mycorrhization Symbiotic mutualisms Legume nodulation 26/43 • • • • • Fungi: Glomeromycota More than 80% of higher plants Very ancient symbiosis (460 MYA) No organogenesis Phosphorous, potassium and nitrogen • Protection against pathogens • Low level of host specificity • • • • • • Rhizobia: gram negative bacteria Restricted to legumes More recent symbiosis (60 MYA) Nodule organogenesis Nitrogen fixation Highly specific Legume nodulation Medicago truncatula-Sinorhizobium meliloti model Legume root nodules 27/43 Mutual recognition of chemical signals Genetic analyses of symbiotic signaling 28/43 Rose*, Venkateshwaran* et al., 2012 MCP Isobaric tags for relative and absolute quantification (iTRAQ) 29/43 Ross, Huang, Pappin, et al. MCP 2004 Quantitative phosphoproteomics workflow 30/43 Unique peptides 63290 Unique phosphopeptides 15335 Localized phosphosites 13506 Proteins 7739 Phosphoproteins 3926 Applications of large scale approaches 1. Phosphoproteomics time course experiments 2. Phosphoproteomics combined with genetics 3. Medicago, proteomic, phosphoproteomic and acetylomic atlas 31/43 Global analysis of the time Course experiments 32/43 Global analysis of the time Course experiments 33/43 Rose*, Venkateshwaran* et al., 2012 MCP Tracking individual phosphorylation events Dynamin-related protein 2B (DRP2B) 34/43 Tracking individual phosphorylation events Selected Reaction Monitoring (SRM) Genetics 35/43 Applications of large scale approaches 1. Phosphoproteomics time course experiments 2. Phosphoproteomics combined with genetics 3. Medicago, proteomic, phosphoproteomic and acetylomic atlas 36/43 Genetic analyses of symbiotic signaling 37/43 Rose*, Venkateshwaran* et al., 2012 MCP Global Changes in Mutants Global Phosphoisoform Changes Global Transcript Changes 38/43 Cross-talk between different symbiotic signaling pathways in Medicago 39/43 Rose*, Venkateshwaran* et al., 2012 MCP Applications of large scale approaches 1. Phosphoproteomics time course experiments 2. Phosphoproteomics combined with genetics 3. Medicago, proteomic, phosphoproteomic and acetylomic atlas 40/43 Medicago, proteomic, phosphoproteomic, acetylomic atlas Leaves Seeds Stems Flowers 10, 14 &28 dpi nodules Roots 41/43 http://more.biotech.wisc.edu/ 42/43 Rose*, Venkateshwaran* et al., 2012 MCP Thank you 43/43