* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download PPT File
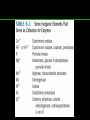
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Lipid signaling wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Ultrasensitivity wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Biochemistry wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Restriction enzyme wikipedia , lookup
Proteolysis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Metalloprotein wikipedia , lookup
Biosynthesis wikipedia , lookup
Catalytic triad wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
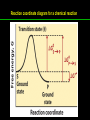
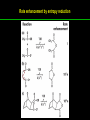
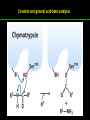
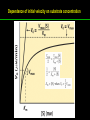
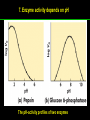
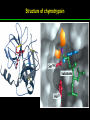
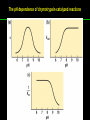
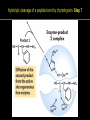
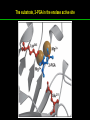
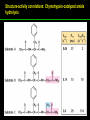
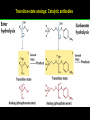
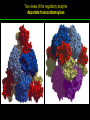
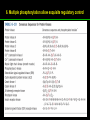
高等生化學 Advanced Biochemistry Enzymes 陳威戎 Preface One way in which this condition might be fulfilled would be if the molecules when combined with the enzyme, lay slightly further apart than their equilibrium distance when [covalently joined], but nearer than their equilibrium distance when free….Using Fischer’s lock and key simile, the key does not fit the lock quite perfectly but exercises a certain strain on it. - J. B. S. Haldane, Enzymes, 1930 Catalysis can be described formally in terms of a stabilization of the transition state through tight binding to the catalyst. - William P. Jencks, article in Advances in Enzymology, 1975 Two fundamental conditions for life 1. The living entity must be able to self-replicate. 2. The organism must be able to catalyze chemical reactions efficiently and selectively. ~ Without catalysis, chemical reactions could not occur on a useful time scale, and thus could not sustain life. Extraordinary catalytic power of enzymes 1. High degree of specificity for their substrates 2. Accelerate chemical reactions tremendously 3. Function in aqueous solutions under very mild conditions of temperature and pH Enzymes 1. An Introduction to Enzymes 2. How Enzymes Work 3. Enzyme Kinetics as an Approach to Understanding Mechanism 4. Examples of Enzymatic Reactions 5. Regulatory Enzymes An Introduction to Enyzmes 1. Late 1700s- digestion of meat by secretions of the stomach. 2. 1800s- conversion of starch to sugar by saliva and plant extracts. 3. Louis Pasteur (1850s)- fermentation of sugar into alcohol by yeast is catalyzed by “ferments”. ~ ferments are inseparable from living yeast cells- “vitalism” 4. Eduard Buchner (1897)- yeast extracts could ferment sugar to alcohol. ~ fermentation was promoted by molecules that continue to function when removed from cells. 5. Frederick W. Kühne- called these molecules “enzymes”. An Introduction to Enyzmes 6. James Sumner (1926)- isolation and crystallization of urease. ~ postulated that “all enzymes are proteins” 7. John Northrop and Moses Kunitz (1930s)- crystallized pepsin, trypsin, and other digestive enzymes. ~ all found to be proteins 8. J. B. S. Haldane (1850s)- wrote “Enzymes” ~ weak bonding interactions between an enzyme and its substrate might be used to catalyze a reaction. 9. Late 20th century- purification and elucidation of the structure and mechanism of many enzymes. 1. Most enzymes are proteins 1. With the exception of a small group of catalytic RNA and DNA molecules, all enzymes are proteins. 2. The catalytic activity depends on the integrity of their native protein conformation. 3. Enzymes have MWs ranging from 12,000 to more than 1 million. 4. Some enzymes require no chemical groups for activity, others require an additional chemical component called a “cofactor” (inorganic ions) or a “coenzyme” (complex organic or metalloorganic molecules). Prosthetic group Holoenzyme vs. Apoenzyme 2. Enzymes are classified by the reactions they catalyze Many enzymes have been named: 1. By adding the suffix “-ase” to their substrate ex: urease 2. For a broad function before the specific reaction catalyzed was known 3. For their sources ex: pepsin ex: trypsin 4. By the Nomenclature Committee of IUBMB ex: ATP:glucose phosphotransferase (E.C. 2.7.1.1.) Enzymes 1. An Introduction to Enzymes 2. How Enzymes Work 3. Enzyme Kinetics as an Approach to Understanding Mechanism 4. Examples of Enzymatic Reactions 5. Regulatory Enzymes Binding of a substrate to an enzyme at the active site How Enzymes Work 1. Enzymes affect reaction rates, not equilibria. 2. Reaction rates and equilibria have precise thermodynamic definitions. 3. A few principles explain the catalytic power and specificity of enzymes. 4. Weak interactions between enzyme and substrate are optimized in the transition state. 5. Binding energy contributes to reaction specificity and catalysis. 6. Specific catalytic groups contribute to catalysis. 1. Enzymes affect reaction rates, not equilibria : Standard free energy change : Biochemical standard free-energy change : Activation energy Ground state: Substrate (S) ; Product (P) Transition state Reaction intermediates (ES or EP) Catalysts enhance reaction rates by lowering activation energies. Reaction coordinate diagram for a chemical reaction Reaction coordinate diagram comparing enzymecatalyzed and uncatalyzed reactions 2. Reaction rates and equilibria have precise thermodynamic definitions For an equilibrium such as Equilibrium constant: Keq ; The relationship between K’eq and G’o: Reaction rate is determined by [reactants] and rate constant: k For a unimolecular reaction S→P, the rate (or velocity), V is expressed by a rate equation: (first-order reaction) For a bimolecular reaction, (second-order reaction) 3. A few principles explain the catalytic power and specificity of enzymes Enzymes are extraordinary catalysts: 1. The rate enhancements that the enzymes bring about are in the range of 5 to 17 orders of magnitude. 2. Enzymes are very specific. The source of energy for the lowering of activation energy: 1. The rearrangements of covalent bonds during an enzymecatalyzed reaction. 2. The noncovalent interactions between enzyme and substrate. 3. A few principles explain the catalytic power and specificity of enzymes : binding energy ; derived from enzyme-substrate interaction. A major source of free energy used by enzymes to lower the activation energies of reactions. How enzymes use noncovalent binding energy? 1. Catalytic power of enzymes is derived from the free energy released in forming many weak interactions with their substrates. This binding energy contributes to specificity and to catalysis. 2. Weak interactions are optimized in the reaction transition state. Enzyme active sites are complementary not to the substrates but to the transition states. 4. Weak interactions between enzyme and substrate are optimized in the transition state 1. Emil Fischer (1894) proposed that enzymes were structurally complementary to their substrates, so that they fit together like a lock and key. (An elegant but misleading idea!) 2. Michael Polani (1921), Haldane (1930), and Linus Pauling (1946) proposed the modern notion of enzymatic catalysis: in order to catalyze reactions, an enzyme must be complementary to the reaction transition state. Weak binding interactions between the enzyme and the substrate provide a substantial driving force for enzymatic catalysis. Complementary shapes of a substrate and its binding site on an enzyme The enzyme dihydrofolate reductase with its substrate NADP+ unbound (left) and bound (right). An imaginary enzyme (stickase) designed to catalyze breakage of a metal stick 5. Binding energy contributes to reaction specificity and catalysis Prominent physical and thermodynamic factors contributing to 1. A reduction in entropy 2. The solvation shell of hydrogen-bonded water 3. The distortion of substrates 4. The need for proper alignment of catalytic functional groups “Induced fit”, a mechanism postulated by Daniel Koshland (1958) Role of binding energy in catalysis Rate enhancement by entropy reduction 6. Specific catalytic groups contribute to catalysis 1. General acid-base catalysis Proton transfers mediated by molecules other than water. Amino acid side-chains in the active site of an enzyme can act as proton donors or acceptors. 2. Covalent catalysis A transient covalent bond is formed between the enzyme and the substrate. Amino acid side-chains and functional groups of coenzymes can serve as nucleophiles in the formation of covalent bonds. 3. Metal ion catalysis Ionic interactions between enzyme-bound metal and substrate. Mediate oxidation-reduction reactions. How a catalyst circumvents unfavorable charge development during cleavage of an amide Amino acids in general acid-base catalysis Covalent and general acid-base catalysis Enzymes 1. An Introduction to Enzymes 2. How Enzymes Work 3. Enzyme Kinetics as an Approach to Understanding Mechanism 4. Examples of Enzymatic Reactions 5. Regulatory Enzymes Enzyme Kinetics as an Approach to Understanding Mechanism 1. Substrate concentration affects the rate of enzymecatalyzed reactions. 2. The relationship between substrate concentration and reaction rate can be expressed quantitatively. 3. Kinetic parameters are used to compare enzyme activities. 4. Many enzymes catalyze reactions with two or more substrates. 5. Pre-steady state kinetics can provide evidence for specific reaction steps. 6. Enzymes are subject to reversible or irreversible inhibition. 7. Enzyme activity depends on pH. 1. Substrate concentration affects the rate of enzymecatalyzed reactions- Initial rate (velocity) One simplifying approach in enzyme kinetics experiments is to measure the initial rate (or initial velocity), designated V0, when [S] is much greater than the concentration of enzyme, [E]. At relatively low concentrations of substrate, V0 increases almost linearly with an increase in [S]. A point is reached beyond which increases in V0 are vanishingly small as [S] increases. This plateau-like region is close to the maximum velocity, Vmax. 1. Substrate concentration affects the rate of enzymecatalyzed reactions- ES complex Victor Henri (1903) proposed that the combination of an enzyme with its substrate to form an ES complex is a necessary step in enzymatic catalysis. Leonor Michaelis and Maud Menten (1913) postulated that the enzyme first combines reversibly with its substrate to form an ES complex in a relatively fast reversible step: The ES complex then breaks down in a slower second step to yield the free enzyme and the reaction product P: 1. Substrate concentration affects the rate of enzymecatalyzed reactions- Steady state The enzyme is first mixed with a large excess of substrate, there is an initial period, the pre-steady state, during which [ES] builds up. The reaction quickly achieves a steady state in which [ES] remains approximately constant over time. The concept of a steady state was introduced by G. E. Briggs and Haldane in 1925. Analysis of these initial rates is referred to as steady-state kinetics. Effect of substrate concentration on the initial velocity of an enzyme-catalyzed reaction 2. The relationship between substrate concentration and reaction rate can be expressed quantitatively [Et]: total enzyme concentration [Et]-[ES]: free or unbound enzyme Since [S] is far greater than [Et], the amount of substrate bound by the enzyme is negligible compared with the total [S]. With these conditions in mind, the following steps lead us to an expression for V0 in terms of easily measurable parameters. 2. The relationship between substrate concentration and reaction rate can be expressed quantitatively The modern derivation of Michaelis-Menten equation: Step 1: the formation and breakdown of ES complex Step 2: Steady-state assumption: the rate of formtion of ES is equal to the rate of its breakdown 2. The relationship between substrate concentration and reaction rate can be expressed quantitatively Step 3: Solve the equation for ES Km= (Michaelis-Menten constant) Step 4: Express Vo in terms of ES Vmax occurs when enzyme is saturated ([ES]=[Et]) Vmax can be defined as k2[Et] Dependence of initial velocity on substrate concentration 3. Kinetic parameters are used to compare enzyme activities All enzymes that exhibit a hyperbolic dependence of V0 on [S] are said to follow Michaelis-Menten kinetics. When k2 is rate-limiting, k2<<k-1 and Km reduces to k-1/k1, which is defined as the dissociation constant, Kd, of the ES complex. Km represents the affinity of the enzyme for its substrate. Define a more general rate constant, kcat = Vmax/[Et], also called “turnover number” Specificity constant, kcat/Km: the best way to compare the catalytic efficiencies of different enzymes or substrates. Transformation of the Michaelis-Menten equation: The double-reciprocal (Lineweaver-Burk) plot Versions useful in the determination of Km and Vmax. 4. Many enzymes catalyze reactions with two or more substrates 5. Pre-steady state kinetics can provide evidence for specific reaction steps A ternary complex is formed in the reaction steps. A Ping-Pong (double-displacement) pathway. 6. Enzymes are subject to reversible or irreversible inhibition Enzyme inhibitors are molecular agents that interfere with catalysis, slowing or halting enzymatic reactions. There are two broad classes of enzyme inhibition: 1. Reversible inhibition (1) Competitive inhibition (2) Uncompetitive inhibition (3) Mixed inhibition (a special case: Noncompetitive inhibition) 2. Irreversible inhibition Suicide inactivators (mechanism-based inactivators) Three types of reversible inhibition- Competitive inhibition Three types of reversible inhibition- Uncompetitive inhibition Three types of reversible inhibition- Mixed inhibition Irreversible inhibition Irreversible inhibitors bind covalently with or destroy functional groups on an enzyme essential for activity or form a particularly stable noncovalent interaction. Suicide inactivators: unreactive until they bind to the active site of an specific enzyme, also called “mechanism-based inactivators”.suitable for “rational drug design” 7. Enzyme activity depends on pH The pH-activity profiles of two enzymes Enzymes 1. An Introduction to Enzymes 2. How Enzymes Work 3. Enzyme Kinetics as an Approach to Understanding Mechanism 4. Examples of Enzymatic Reactions 5. Regulatory Enzymes Examples of Enzymatic Reactions 1. The chymotrypsin mechanism involves acylation and deacylation of a serine residue. 2. Hexokinase undergoes induced fit on substrate binding. 3. The enolase reaction mechanism requires metal ions. 4. Lysozyme uses two successive nucleophilic displacement reactions. 1. The chymotrypsin mechanism involves acylation and deacylation of a serine residue Bovine pancreatic chymotrypsin: a protease that catalyze the hydrolytic cleavage of peptide bonds. Specific for peptide bonds adjacent to aromatic amino acids. General acid-base catalysis and covalent catalysis. Enhance the rate of peptide bond hydrolysis by at least 109-fold. The reaction has two phases: (1) acylation phase ; (2) deacylation phase. Structure of chymotrypsin Structure of chymotrypsin Pre-steady state kinetic evidence for an acyl-enzyme intermediate The pH dependence of chymotrypsin-catalyzed reactions Hydrolytic cleavage of a peptide bond by chymotrypsin- Step 1 Hydrolytic cleavage of a peptide bond by chymotrypsin- Step 2 Hydrolytic cleavage of a peptide bond by chymotrypsin- Step 3 Hydrolytic cleavage of a peptide bond by chymotrypsin- Step 4 Hydrolytic cleavage of a peptide bond by chymotrypsin- Step 5 Hydrolytic cleavage of a peptide bond by chymotrypsin- Step 6 Hydrolytic cleavage of a peptide bond by chymotrypsin- Step 7 Some common nucleophiles and electrophiles in biochemistry 2. Hexokinase undergoes induced fit on substrate binding Yeast hexokinase (Mr 107,862) is a bisubstrate enzyme that catalyzes the following reversible reaction: Hexokinase favors the reaction with glucose by a factor of 106. Reaction rates increase greatly in the presence of the functional phosphoryl group acceptor (glucose). General acid-base catalysis and transition-state stabilization. Induced fit in hexokinase Hexokinase has a U-shaped structure. The ends pinch toward each other in a conformational change induced by binding of D-glucose. 3. The enolase reaction mechanism requires metal ions A glycolytic enzyme, enolase, catalyze the reversible dehydration of 2-phosphoglycerase to phosphoenolpyruvate: Yeast enolase (Mr 93,316) is a dimer with 436 amino acid residues per subunit. Metal ion catalysis, general acid-base catalysis and transitionstate stabilization. Two-step reaction catalyzed by enolase The carboxyl group of 2-PGA is coordinated by two Mg2+ at the active site. A proton is abstracted in step 1 by general base catalysis (Lys345), and the resulting enolic intermediate is stabilized by the two Mg2+ ions. Elimination of the -OH in step 2 is facilitated by general acid catalysis (Glu211). The substrate, 2-PGA in the enolase active site Structure-activity correlations: Chymotrypsin-catalyzed amide hydrolysis Transition-state analogs: Catalytic antibodies 4. Lysozyme uses two successive nucleophilic displacement reactions A natural antibacterial agent found in tears and egg whites. Hen egg white lysozyme (Mr 14,296) is a monomer with 129 a.a. The structure revealed four disulfide bonds and a cleft containing the active site. Substrate: peptidoglycan in bacterial cell walls. Cleaves the b1→4 glycosidic bond between two types of sugar residues in a molecule: N-acetylmuramic acid (Mur2Ac, NAM) and N-acetylglucosamine (GlcNAc, NAG) Key catalytic amino acid residues : Glu35 and Asp52 Ribbon diagram of the hen egg white lysozyme Reaction catalyzed by hen egg white lysozyme Two proposed pathways of lysozyme reaction Two proposed pathways of lysozyme reaction Two proposed pathways of lysozyme reaction Two proposed pathways of lysozyme reaction Ribbon diagram of the covalent lysozyme-substrate intermediate Enzymes 1. An Introduction to Enzymes 2. How Enzymes Work 3. Enzyme Kinetics as an Approach to Understanding Mechanism 4. Examples of Enzymatic Reactions 5. Regulatory Enzymes Regulatory enzymes are modulated in a variety of ways 1. Allosteric enzymes function through reversible noncovalent binding of regulatory compounds called allosteric modulators or allosteric effectors. 2. Other enzymes are regulated by reversible covalent modification. Regulatory Enzymes 1. Allosteric enzymes undergo conformational changes in response to modulator binding. 2. In many pathways a regulated step is catalyzed by an allosteric enzyme. 3. The kinetic properties of allosteric enzymes diverge from Michaelis-Menten behavior. 4. Some regulatory enzymes undergo reversible covalent modification. 5. Phosphoryl groups affect the structure and catalytic activity of proteins. 6. Multiple phosphorylations allow exquisite regulatory control. 7. Some enzymes and other proteins are regulated by proteolytic cleavage of an enzyme precursor. 1. Allosteric enzymes undergo conformational changes in response to modulator binding 1. Allosteric enzymes are significantly different from those of simple nonregulatory enzymes. (Some are structural.) 2. Allosteric enzymes generally have one or more regulatory, or allosteric, sites for binding the modulator. 3. Allosteric enzymes are generally larger and more complex than nonallosteric enzymes. Most have two or more subunits. 4. Aspartate transcarbamoylase (ATCase), which catalyzes an early reaction in the biosynthesis of pyrimidine nucleotides, has 12 polypeptide chains organized into catalytic and regulatory subunits. Subunit interactions in an allosteric enzyme, and interactions with inhibitors and activators Two views of the regulatory enzyme Aspartate transcarbamoylase 2. In many pathways a regulated step is catalyzed by an allosteric enzyme Feedback inhibition Heterotropic allosteric inhibition 3. The kinetic properties of allosteric enzymes diverge from Michaelis-Menten behavior- Sigmoid curve (homotropic) Not Km! 3. The kinetic properties of allosteric enzymes diverge from Michaelis-Menten behavior- Effects of modulators 3. The kinetic properties of allosteric enzymes diverge from Michaelis-Menten behavior- Less common type of modulation 4. Some regulatory enzymes undergo reversible covalent modification 1. Phosphorylation: 1/3 to 1/2 of all proteins in a eukaryotic cell are phosphorylated 2. Adenylylation: 3. Uridylylation: 4. Methylation: Methyl-accepting chemotaxis protein of bacteria 5. ADP-ribosylation: Dinitrogenase reductase of bacteria Some enzyme modification reactions- Phosphorylation Some enzyme modification reactions- Adenylylation Some enzyme modification reactions- Uridylylation Some enzyme modification reactions- ADP-ribosylation Some enzyme modification reactions- Methylation 5. Phosphoryl groups affect the structure and catalytic activity of proteins 1. The attachment of phosphoryl groups is catalyzed by protein kinases; removal by protein phosphatases 2. Addition of a phosphoryl group to a Ser, Thr, or Tyr introduces a bulky, charged group into a moderately polar region. 3. Oxygen atoms can hydrogen-bond with: amide group of peptide backbone at the start of an a-helix charged guanidinium group of Arg 4. Two negative charges can repel neighboring Asp or Glu Regulation of glycogen phosphorylase activity by covalent modification 6. Multiple phosphorylation allow exquisite regulatory control Multiple regulatory phosphorylations 7. Some enzymes and other proteins are regulated by proteolytic cleavage of an enzyme precursor