* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Glucose/Galactose Binding Protein (GGBP)
Gene expression wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
Biochemical cascade wikipedia , lookup
Lipid signaling wikipedia , lookup
Biochemistry wikipedia , lookup
Ultrasensitivity wikipedia , lookup
Metalloprotein wikipedia , lookup
Signal transduction wikipedia , lookup
Expression vector wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Magnesium transporter wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Paracrine signalling wikipedia , lookup
Interactome wikipedia , lookup
Structural alignment wikipedia , lookup
Point mutation wikipedia , lookup
Western blot wikipedia , lookup
Protein purification wikipedia , lookup
Proteolysis wikipedia , lookup
Homology modeling wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein–protein interaction wikipedia , lookup
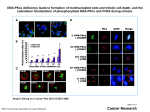
Molecular modeling of casein kinase-1 to determine relevance of conserved potential phosphorylation sites Paige R. Pritchett1, Troy C. Messina1, and Cynthia J. Brame2 1Centenary College of Louisiana, Department of Physics, 2911 Centenary Boulevard, Shreveport, LA 71104 2Center for Teaching, Vanderbilt University, Nashville, TN 37212 Eukaryotic protein kinases transfer a phosphate from a nucleoside triphosphate to a protein substrate. The eukaryotic protein kinases are homologous and therefore exhibit significant structural similarity but can be divided into eight subfamilies with closer structural and functional relationships. The CK1 subfamily is ubiquitous and highly conserved within Eukarya, and has members that are relevant to regulating cell proliferation and differentiation, chromosomal segregation, and circadian rhythm. In humans, CK1s have been linked to neurodegenerative diseases and cancer. We are investigating regulation of CK1 activity by phosphorylation, using yeast CK1 protein kinases as models. We have previously identified phosphorylation sites that negatively regulate activity through in vitro and in vivo studies of phosphorylation-mimicking and -preventing mutations. In this study, we have used NAMD to investigate the effect of mutations on protein structure in silico. We have identified structural changes large enough to modify kinase activity, for example S179 shifts towards a positively charged RD pocket by 3-4 Å upon phosphorylation or mutation to a glutamic acid. The phosphorylated amino acid is also subject to a much more restrictive potential of mean force over the coordinate between S179 and the RD pocket. The simulation results will be further contextualized in terms of the laboratory work. Overview Protein kinases are highly conserved in form and function within the domain Eukarya. The Casein kinase-1 family of protein kinases has been implicated in controlling neural processes and in the normal development of yeast buds. This second consequence suggests CK1 may be important in the proliferation and development of cells in eukaryotes. In yeast cells, phosphorylation leads to deactivation of the protein. Blast alignments show conserved regions across many CK1 orthologs. Experiments mutating phosphorylatable amino acids in the conserved regions have been performed and implicate a particular serine as important to the activity of CK1. Mutations to this amino acid was performed using molecular modeling software VMD and NAMD. We then performed molecular dynamics simulations and steered molecular dynamics to better understand the structural relation to CK1 activity. Methodology/Procedure Conclusion and Ongoing Work The crystal structure of the wildtype, non-phosphorylated (active) version of casein kinase-1 was obtained from the protein data bank (PDB ID: 1CSN). NAMD 2.9 (Win32 version) and VMD 1.9.1 (Windows OpenGL version) were used to mutate, solvate, visualize, and simulate the wildtype and phosphorylated versions of the protein. The current results show that there is a movement of the lowest energy conformation for the phosphorylated version that holds the phosphorylated S179 closer to the RD pocket at R130. The graph at left displays the movement of the phosphorylated S179 in relation to the wildtype. The serine at position 179 (S179) was mutated or phosphorylated to see what structural changes resulted. Molecular dynamics (MD) simulations were run on the solvated enzyme to provide structural information for determining change in structure as a result of the changes to S179. The structures are both tightly held at those points and are unlikely to cross any energy barriers that involve a force of more than ~3 kcal/mol due to thermodynamic hindrance. Adaptive biasing force simulations are still being run to obtain a complete graph of the potential of mean force. The distance between S179 and a conserved, arginine (R130) was calculated from the simulation. R130 is a putative charge interaction partner for pS179. A distance histogram was generated from the distance data and used to support the investigation of the phosphorylated S179 and its possible role in activation/deactivation of CK1. Acknowledgements • Funding – – Centenary College Biophysics Department Centenary College Gus S. Wortham Endowed Chair of Engineering • Dr. Lucy Robinson References The above left image shows the relation of protein kinase families. Casein kinase-1 is located at the top center of the image. The above right image displays the location of labeled CK1 during the budding of a yeast cell. The highlighted portions correspond to the position of the CK1 and make it clear that it is consistently associated with the site where the bud forms. Serine Phosphoserine Because MD at physiological temperatures only allows for low energy structures that are near the original starting structure to be observed, another way of determining lowest energy structures that may be separated by large barriers had to be used. To determine any other differences in conformation as a result of phosphorylation adaptive biasing force simulations were run. This simulation directed the S179 to move closer and further from the conserved RD pocket at residue 130 and calculated the force it took to hold the protein at each position from 0-20 Å. The above center image shows the protein solvated in water for simulation. The above lower image displays the protein with S179 and R130 highlighted in blue and yellow, respectively. [1] Brame, C. J., Pruitt, W. M., and Robinson, L. C. (2008) A Molecular Genetics Laboratory Course Applying Bioinformatics and Cell Biology in the Context of Original Research, CBE Life Sci Educ 7, 410–421. [2] Phillips, J. C., Braun, R., Wang, W., Gumbart, J., Tajkhorshid, E., Villa, E., Chipot, C., Skeel, R. D., Laxmikant, K., and Schulten, K. (2005) Scalable molecular dynamics with NAMD, J. Comp. Chem. 26, 1781–1802. [3] Case, D. A. (2012) AMBER 12. [4] Brooks, B. R., Bruccoleri, R. E., Olafson, B. D., States, D. J., Swaminathan, S., and Karplus, M. (1983) CHARMM: A program for macromolecular energy, minimization, and dynamics calculations, Journal of Computational Chemistry 4, 187–217. [5] Schulten, K. VMD online script library, VMD Script Library. [6] Xu, R. M., Carmel, G., Sweet, R. M., Kuret, J., Cheng, X., Carmel, G., Leichus, B., and Cheng, X. (1995) Crystal structure of casein kinase-1, a phosphate-directed protein kinase., EMBO J. 14, 1015–1023. [7] Schulten, K. NAMD Tutorials, NAMD Tutorials. [8] Jeffrey, P. D. et al. (1995) Mechanism of CDK activation revealed by the structure of a cyclinA-CDK2 complex, Nature 376, 313-320. [9] Mok, J. et al. (2011) Deciphering Protein Kinase Specificity through Large-Scale Analysis of Yeast Phosphorylation Site Motifs, Sci. Signal 3(109), 1-29. [10] Knight J. D. R., Qian, B., Baker, D., and Kothary, R. (2007) Conservation, Variability, and the Modeling of Active Protein Kinases, PLoS 10(e982), 1-15. [11] Robinson, L. C., Bradley, C., Bryan, J. D., Jerome, A., Kweon, Y., and Panek, H. R. (1999) The Yck2 Yeast Casein Kinase 1 Isoform Shows Cell Cycle-specific Localization to Sites of Polarized Growth and Is Required for Proper Septin Organization, Mol Biol Cell. 10(4), 1077–1092.