* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Complex III
Multi-state modeling of biomolecules wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Western blot wikipedia , lookup
Biochemistry wikipedia , lookup
Mitochondrial replacement therapy wikipedia , lookup
Photosynthesis wikipedia , lookup
Microbial metabolism wikipedia , lookup
Mitochondrion wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Citric acid cycle wikipedia , lookup
Metalloprotein wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Electron transport chain wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
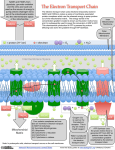
Functions Complex I •Transport of electrons from NADH to ubiquinone •Electron source: NADH •Co-factor: Flavin mononucleotide • Transport: via eight redox groups, iron–sulphur clusters Complex I Functions Cont. Electron acceptor: Ubiquinone Ubiquinone function: Transfers of electrons to next complex in the chain (Complex III) Simultaneous shunting of protons across inner mitochondria membrane to intermembrane space Stoichiometry: 4H+/2e- In the first step of electron transfer, the hydride ion is transferred to FMN, forming is then oxidized in two steps via a semiquinone intermediate. The two electrons are transferred one at a time to the next oxidizing agent, an iron–sulfur cluste These Fe–S clusters provide a channel for electrons, directing them to the membrane-bound portion of the complex where ubiquinone (Q) accepts electrons one at a time passing through a semiquinone anion intermediate before reaching its fully reduced state, ubiquinol Q and are lipid soluble cofactors. They remain within the lipid bilayer and can diffuse freely in two dimensions. One of the reasons for the complicated electron transport chain within complex I is to carry electrons from an aqueous environment to a hydrophobic environment within the membrane. In the first step of electron transfer, the hydride ion is transferred to FMN, forming is then oxidized in two steps via a semiquinone intermediate. The two electrons are transferred one at a time to the next oxidizing agent, an iron–sulfur cluster. These Fe–S clusters provide a channel for electrons, directing them to the membrane-bound portion of the complex where ubiquinone (Q) accepts electrons one at a time passing through a semiquinone anion intermediate before reaching its fully reduced state, ubiquinol In complex I, there are 4 protons translocated across the membrane for every pair of electrons that pass from NADH to QH 2. . These do not include the protons required for ubiquinone reduction. The proton pump is probably an antiporter located in the membrane-bound module. The mechanism of proton translocation is not clear Complex II Mitochondrial Electron Transport The entry point for electrons from FADH2 of flavoproteins is Ubiquinol (QH2). Succinate-Q reductase complex Complex II •Also called succinate dehydrogenase: – A component of the TCA cycle in mitoch. in both eukaryotic cells and prokaryotic organisms Complex II •Complex II has 4 or 5 polypeptides, all encoded by the nuclear genome •In mitochondria has 4 subunits. 2 integral membrane proteins: the large cyto. b, cybL or C, subunit and the small cybS or D subunit and the iron-sulfur protein •5 mitochondrial complexes, I to V, complex II the only one with no subunits encoded by the mitochondrial genome. Complex II This protein provides two centers for oxidation/reduction reactions •FAD FADH2 •Fe3+ Fe2+ Complex II •Function: Mitochondrial respiratory chain • Composition: 4 Subunits; All nuclear encoded •Flavoprotein: FAD (SDHA; Fp) •Functions: Catalytic site; Covalently bound FAD cofactor •Iron-Sulfur protein: SDHB (Ip) •Function: Electron transfer between FAD and membrane-bound quinone •Structure: Contains three different iron-sulphur clusters •Cytochrome b subunits: SDHC ; SDHD Integral membrane proteins: Bind Fp & Ip to matrix Location of Complex II Matrix side of mitochondrial inner membrane Binding to membrane is dependent on 2 small (15.5 and 13.5 kDa) proteins SDHC & SDHD Complex II contains three identical multisubunit enzymes that associate to form a trimeric structure that is firmly embedded in the membrane The overall shape resembles a mushroom with its head projecting into the interior of the membrane compartment Most of them have a bound heme b molecule and this subunit is often called cytochrome b. All of the membrane subunits have a Q binding site positioned near the interior surface of the membrane at the point where the head subunits are in contact with the membrane subunits. The sequence of reactions for the transfer of two electrons from succinate to Q begins with the reduction of FAD by a hydride ion. This is followed by two one electron transfers from the reduced flavin to the series of three iron–sulfur clusters In those species with a cytochrome b anchor, the heme group is not part of the electron transfer pathway. Very little free energy is released in the reactions catalyzed by complex II. This means that the complex cannot directly contribute to the proton concentration gradient across the membrane. Instead, it supplies electrons from the oxidation of succinate midway along the electron transport sequence. • Q can accept electrons from complex I or II and donate them to complex III and thence to the rest of the electron-transport chain. • Reactions in several other pathways also donate electrons to Q. one of them, the reaction catalyzed by the glycerol 3-phosphate dehydrogenase complex Complex III COMPLEX III (CYTOCHROME REDUCTASE) Location of Complex III: Inner mitochondrial membrane •Composition •Nuclear subunits •Number: 10 •Components Cytochrome c1 (CYC1) Ubiquinone-binding protein (UQPC) Ubiquinol-cytochrome c reductase core protein II (UQCRC2) Rieske FeS protein •Mitochondrial subunits •Number: 1 •Component: Cytochrome b •Complex III Functions •Transfers electrons from ubiquinol to cytochrome c •Coupled with transfer of electrons across inner mitochondrial membrane •Contains 3 redox centers •Cytochrome b •Cytochrome c •Rieske FeS protein Associations •Supercomplex of Complexes I, III, IV • Association of Complexes III & IV may be stabilized by cardiolipin Complex III: PDB 1BE3 membrane Half of the homodimeric structure is shown. Approximate location of the membrane bilayer is indicated. Not shown are 2 CoQ binding sites, one near heme bH & the other near heme bL. The b hemes are positioned to provide a Fe-S pathway for electrons across the membrane. Complex III (bc1 Complex) heme bH heme bL heme c1 The Rieske iron-sulfur center (Fe-S) has a flexible link to the rest of the complex. PDB 1BE3 Complex III (bc1 Complex) It changes position during e- transfer. membrane Fe-S extracts an e- from CoQ, & then moves closer to heme c1, to which it transfers the e-. (Fe-S protein in green.) Fe-S heme bH heme bL heme c1 Complex III is an obligate homo-dimer. Fe-S in one half of the dimer interacts with bound CoQ & heme c1 in the other half of the dimer. Arrows point at: Fe-S in the half of complex colored white/grey heme c1 in the half of complex with proteins colored blue or green. PDB-1BGY Complex III homo-dimer Fe-S heme c1 Matrix H+ + NADH NAD+ + 2H+ 2 e Q I 2H+ + ½ O2 H2O –– III IV ++ 4H + + 4H cyt c 2H+ Intermembrane Space Complex III (bc1 complex): H+ transport in complex III involves coenzyme Q (CoQ). O O CH3O CH 3 CH3O CH 3 CH3O (CH 2 CH O C CH 3 e CH 2)nH CH 3 CH3O (CH 2 CH O coenzyme Q C CH 2)nH coenzyme Q • e + 2 H+ OH CH3O CH 3 CH 3 CH3O (CH 2 CH OH C CH 2)nH coenzyme QH2 The “Q cycle” depends on: mobility of CoQ in the lipid bilayer existence of binding sites for CoQ within the complex that stabilize the semiquinone radical, Q·. matrix Q Q Cycle: 2 H+ Q. QH2 QH2 cyt bH Complex III cyt bL Q e e Q· Fe-S 2 H+ intermembrane space cyt c1 cyt c As depicted above, electrons enter complex III via coenzyme QH2, which binds at a site on the positive side of the inner mitochondrial membrane, adjacent to the intermembrane space. matrix Q 2 H+ Q. QH2 QH2 QH2 gives up cyt bH one e that is Complex III transferred via e hemes bL & bH to cyt bL a bound Q on e Q Q· Fe-S cyt c1 the other side of + 2 H the membrane. intermembrane space cyt c Loss of one e- to the b hemes, and release of 2 H+ to the intermembrane space, generates a Q·- radical in the site adjacent to the intermembrane space. Q·- becomes Q as it gives up a second e- to the Rieske iron-sulfur center (Fe-S). matrix 2 H+ Fe-S is reoxidized by Q Q. QH2 QH2 electron cyt bH transfer to Complex III cytochrome c1, which e cyt bL passes the e Q Q· Fe-S cyt c1 electron out of the complex to 2 H+ cyt c cytochrome c. intermembrane space Some evidence suggests instead a concerted reaction in which e- transfer from QH2 to Fe-S & cytochrome bL is essentially simultaneous. But there is agreement about the overall reaction cycle. It takes 2 matrix 2 H+ cycles for . Q Q QH2 QH2 CoQ bound at a site cyt bH near the Complex III matrix to e be reduced cyt bL e to QH2, as Q Q· Fe-S cyt c1 2e- are 2 H+ transferred cyt c intermembrane space from the b hemes , and 2H+ are extracted from the matrix compartment. In 2 cycles, 2 QH2 enter the pathway & one is regenerated. matrix Overall reaction catalyzed by complex III, including net inputs & outputs of the Q cycle : Q 2 H+ Q. QH2 QH2 cyt bH Complex III cyt bL Q e e Q· Fe-S 2 H+ intermembrane space cyt c1 cyt c QH2 + 2H+(matrix) + 2 cyt c (Fe3+) Q + 4H+(outside) + 2 cyt c (Fe2+) Per 2e- transferred through the complex to cyt c, 4H+ are released to the intermembrane space. Complex III contains two copies of the enzyme and is firmly anchored to the membrane by a large number of that span the lipid bilayer . The functional enzyme consists of three main subunits: cytochrome c1 cytochrome b, and the Rieske iron–sulfur protein (ISP). Other subunits are present on the inside surface but they do not play a direct role in the ubiquinol:cytochrome c oxidoreductase reaction. The reaction begins when QH2 binds to the Q o site in the cytochrome b subunit. QH2 is oxidized to the semiquinone and a single electron is passed to the adjacent Fe–S complex in the ISP subunit. From there, the electron transfers to the heme group in cytochrome c1 This transfer is facilitated by movement of the head group of ISP Soluble cytochrome c is reduced by transfer of an electron from the membrane-bound cytochrome subunit of complex III. In this reaction, the terminal electron acceptor is cytochrome c. This molecule serves as a mobile electron carrier transferring electrons to complex IV, the next component of the chain. The oxidation of QH2 at the Q0 site is a two-step process with a single electron transferred at each step. The path of electrons from the second step, oxidation of the semiquinone intermediate, follows a different route than the first electron. In this case, the electron is passed sequentially to two different b-type hemes within the membrane portion of the complex. The first heme group bL has a lower reduction potential and the second heme bH has a higher reduction potential The bH heme is part of the Qi site where a molecule of Q is reduced to QH2 in a two-step reaction that involves a semiquinone intermediate. A single electron is transported from bL(at the site) to bH(at the site) to Q to produce the semiquinone. Then, a second electron is transferred to reduce the semiquinone to QH2 The second electron is derived from the oxidation of a second molecule of QH2 at the site Q0 . This second oxidation of QH2 also results in the reduction of a second molecule of cytochrome c since the two electrons from the second follow separate paths. The net result is that the oxidation of two molecules of QH2 at the Q0site produces two molecules of reduced cytochrome c and regenerates a molecule of QH2 at the Qi site. Four protons are produced during the oxidation of two molecules of QH2 at the Q o site. These protons are released to the exterior of the membrane compartment and they contribute to the proton gradient that is formed during membrane associated electron transport. Complex IV COMPLEX IV, CYTOCHROME c OXIDASE SUBUNIT I; MTCO1 Alternative titles; symbols CYTOCHROME c OXIDASE I; COI • 1 of 3 mitochondrial DNA (mtDNA) encoded subunits (MTCO1, MTCO2, MTCO3) of respiratory Complex IV •located within the mitochondrial inner membrane and is the third and final enzyme of the ETC of MOP COMPLEX IV •It collects electrons from reduced cyto. C and transfers them to O2 to give water and The energy released is used to transport protons across the mitochondrial inner membrane •Complex IV is composed of 13 polypeptides. Subunits I, II, and III (MTCO1, MTCO2, MTCO3) are encoded by mtDNA while subunits IV, Va, Vb, VIa, VIb, VIc, VIIa, VIIb, VIIc, and VIII are nuclear encoded Complex IV Composition & Related Proteins Nuclear subunits Number: 10 Presumed to play a structural & regulatory role in COX Mitochondrial subunits Number: 3 Largest COX subunits Form catalytic core of COX Contain the 3 copper atoms & 2 heme A molecules Serve as prosthetic groups in holoenzyme Directly involved in electron transfer This complex catalyzes the oxidation of the reduced cytochrome c molecules produced by complex III. The reaction includes a four-electron reduction of molecular oxygen to water and the translocation of four protons across the membrane. The total mass of mammalian complex IV is greater than 400 kDa. Additional subunits in the eukaryotic complexes play a role in assembling complex IV and in stabilizing the structure. The core structure of cytochrome c oxidase is formed from the three conserved subunits—I, II, and III. These polypeptides are encoded by mitochondrial genes in all eukaryotes. Subunit I is almost entirely embedded in the membrane. The bulk of this polypeptide consists of 12 transmembrane There are three redox centers buried within subunit I—two of them are a-type hemes (heme-a and a3 ), and the third is a copper atom The copper atom is in close to the iron atom of forming a binuclear center where the reduction of molecular oxygen takes place. Subunit II has two transmembrane helices that anchor it to the membrane. Most of the polypeptide chain forms a domain located on the exterior surface of the membrane. This domain contains a copper redox center 1CuA2 The external domain of subunit II is the site where cytochrome c binds to cytochrome c oxidase. Subunit III has seven transmembrane helices and is completely embedded in the membrane. There are no redox centers in subunit III and it can be artificially removed without loss of catalytic activity. Its role in vivo is to stabilize subunits I and II and help protect the redox centers from inappropriate oxidation–reduction reactions. Electrons are transferred one at a time from the site to the heme a prosthetic group in subunit I. From there they are transferred to the heme binuclear center. The two heme groups (a and ) have identical structures but differ in their standard reduction potentials One oxygen atom is bound to the iron atom of the group and the other is bound to the copper atom. Subsequent protonation and electron transfer results in the release of a water molecule from the copper site followed by release of a second water molecule from the iron ligand. The overall reaction requires the uptake of four protons from the inside surface of the membrane Cytochrome c Oxidase ATP Synthase ATP Synthase Proton diffusion through the protein drives ATP synthesis! Two parts: F1 and F0 Racker & Stoeckenius confirmed Mitchell’s hypothesis using vesicles containing the ATP synthase and bacteriorhodopsin ATP synthase, embedded in cristae of the inner mitochondrial membrane, includes: ADP + Pi ATP F1 + 3H matrix Fo intermembrane space F1 catalytic subunit, made of 5 polypeptides with stoichiometry a3b3gde. Fo complex of integral membrane proteins that mediates proton transport. F1Fo couples ATP synthesis to H+ transport into the mitochondrial matrix. ADP + Pi ATP F1 3 H+ matrix Fo intermembrane space Transport of least 3 H+ per ATP is required, as estimated from comparison of: DG for ATP synthesis under cellular conditions (free energy required) DG for transfer of each H+ into the matrix, given the electrochemical H+ gradient (energy available per H+). In spite of their name, F-type ATPases are responsible for synthesizing, not hydrolyzing, ATP. They are membranebound and have a characteristic knob-and-stalk structure Rotation of the γ subunit inside α β the hexamer alters the conformation of the subunits, opening and closing the active sites. The a, b, and γ subunits form an arm that also attaches the component to the oligomer. This unit is termed the “stator.” Passage of protons through the channel at the interface between the a and c subunits causes the rotor assembly to spin in one direction relative to the stator. The entire structure is often called a molecular motor. Rotation of the subunit within the component takes place in a stepwise, jerky manner where each step is 120° of rotation. As the c-ring rotates it twists the shaft until enough tension builds up to cause it to snap into the next position within the hexamer. If the c-ring has 10 subunits then a complete rotation requires the translocation of 10 protons and results in the production of 3 ATP molecules. Enzyme that actually makes the ATP The mechanism of ATP synthesis from ADP In 1979 Paul Boyer proposed the binding change mechanism based on observations suggesting that the substrate and product binding properties of the active site could change as protons moved across the membrane. The oligomer of ATP synthase contains three catalytic sites. At any given time, each site can be in one of three different conformations. The three conformations are: (1) open: newly synthesized ATP can be released and can bind, (2) loose: bound cannot be released, and (3) tight: 1. One molecule of ADP and one molecule of bind to an open site. 2. Rotation of the shaft causes each of the three catalytic sites to change conformation. The open conformation (containing the newly bound ADP and Pi) becomes a loose site. The loose site, already filled with ADP and becomes a tight site. The ATP-bearing tight site becomes an open site. 3. ATP is released from the open site, and ADP and condense to form ATP in the tight site. ADP + Pi ATP Matrix H+ + NADH NAD+ + 2H+ 2 e Q I 2H+ + ½ O2 H2O –– III IV Fo ++ 4H+ F1 4H+ cyt c 2H+ 3H+ Intermembrane Space The Chemiosmotic Theory of oxidative phosphorylation, for which Peter Mitchell received the Nobel prize, states that coupling of ATP synthesis to respiration is indirect, via a H+ electrochemical gradient. ADP + Pi ATP Matrix H+ + NADH NAD+ + 2H+ 2 e Q I 2H+ + ½ O2 H2O –– III IV Fo ++ 4H+ F1 4H+ cyt c 2H+ 3H+ Intermembrane Space Chemiosmotic theory - F1Fo ATP synthase: Non-spontaneous ATP synthesis is coupled to spontaneous H+ transport into the matrix. The pH & electrical gradients created by respiration are the driving force for H+ uptake. H+ return to the matrix via Fo "uses up" pH & electrical gradients. Matrix H+ + NADH NAD+ + 2H+ 2 e Q I Simplified depicting: 2H+ + ½ O2 H2O –– III IV ++ 4H + + 4H cyt c 2H+ Intermembrane Space Ejection of a total of 20H+ from the matrix per 4etransferred from 2 NADH to O2 (10H+ per ½O2). Not shown is OH- that would accumulate in the matrix as protons, generated by dissociation of water (H2O H+ + OH-), are pumped out. Also not depicted is the effect of buffering. Transport of ATP, ADP, & Pi ATP produced in the mitochondrial matrix must exit to the cytosol to be used by transport pumps, kinases, etc. ADP & Pi arising from ATP hydrolysis in the cytosol must reenter the matrix to be converted again to ATP. Two carrier proteins in the inner mitochondrial membrane are required. The outer membrane is considered not a permeability barrier. Large outer membrane VDAC channels are assumed to allow passage of adenine nucleotides and Pi. MITOCHONDRIAL TRANSPORT Electroneutral Transport •Inner membrane is highly impermeable, it contains many transport proteins to control the movement of substances into and out of the matrix. Most of the transport systems require the exchange of molecules, and most of these exchanges occur with molecules having the same charge; this is termed electroneutral transport. Electroneutral Transport cont. -For instance pyruvate (-1 charge), which enters the matrix for further metabolism via pyruvat dehydrogenase or pyruvate carboxylase, exchanges with a negatively charged hydroxyl ion (OH ). - Phosphate (PO4 ), which enters the matrix for synthesis of ATP) also can exchange with OH . - Citrate, a tricarboxylic acid, has a negative 3 (3 ) charge at physiological pH, but exchanges with malate, a dicarboxylic acid (2 ). To maintain electroneutrality of this exchange, a proton accompanies the citrate thus neutralizing one of + its negative charges (citrate3 + H exchanges for malate2 ). Electrogenic Transport •There are some transport systems in which the exchange of molecules involves unequal charges. •When there is the net movement of charge across the membrane, this is termed electrogenic transport. •In mitochondria, net negative charges move out of the matrix while net positive charges must move into the matrix by following the charge gradient •Recall that the pumping of protons creates a gradient of negative inside to positive outside. So that the more negatively charged matrix will favor the net outward movement of negative charge. •The most important electrogenic transporter is the adenine nucleotide transporter •ATP produced in the mitochondrial matrix by oxidative phosphorylation is needed in the cytoplasm for energy-requiring processes such as Muscle contraction Lipogenesis Cholesterol synthesis Gluconeogenesis •Hence, it is obligatory that ATP only be transported into the cytoplasm while ADP moves into the mitochondria matrix where it can be phosphorylated to ATP Electrogenic transport system in mitochondria. The adenine nucleotide exchange ensures that ATP4- produced in the mitochondrial matrix is transported to the cytoplasm where it is needed. Adenine nucleotide transporter •ADP bears a negative three charge whereas ATP has a negative four charge at physiological pH •Thus, the exchange of ATP4- moving out with ADP3- moving in creates a net charge of negative one moving into the intermembrane space. •The charge gradient facilitates this exchange of ADP for ATP. The reverse exchange cannot occur in functional mitochondria because it would require the more negative ATP molecule to move towards the more negative environment of the matrix, a process that cannot occur Phosphate Can Be Obtained by H+ Symport or OH– Antiport Active Transport of ATP, ADP, and Pi Across the Mitochondrial Membrane Because the inner mitochondrial membrane is impermeable to charged substances, a transporter is required to allow ADP to enter and ATP to leave mitochondria. • This transporter is called the adenine nucleotide translocase. • Some of the free energy of the proton concentration gradient is expended to drive this transport process. The phosphate carrier does not draw on the electrical component of the protonmotive force, but does draw on the concentration difference (pH). The combined energy cost of transporting ATP out of the matrix and ADP and into it is approximately equivalent to the influx of one proton. ADP + Pi ATP ATP4 matrix lower [H+] __ Adenine nucleotide ++ translocase 3 H+ ATP4 ADP3 H2PO4 H+ energy requiring reactions ADP + Pi higher [H+] cytosol (ADP/ATP carrier) is an antiporter that catalyzes exchange of ADP for ATP across the inner mitochondrial membrane. At cell pH, ATP has 4 () charges, ADP 3 () charges. ADP3/ATP4 exchange is driven by, and uses up, membrane potential (one charge per ATP). The P/O Ratio The P/O ratio is the ratio of molecules phosphorylated to atoms of oxygen reduced. Four protons are translocated by complex I, four by complex III, and two by complex IV. Thus, for each pair of electrons that pass through these complexes from NADH to a total of ten protons are moved across the membrane. Matrix H+ + NADH NAD+ + 2H+ 2 e Q I 2H+ + ½ O2 H2O –– III IV ++ 4H + Intermembrane Space + 4H cyt c + 2H The P/O ratio for succinate is only since electrons contributed by succinate oxidation do not pass through complex I. The ratio of protons translocated intermembrane space per pair of electorns transferred by coupled electron- transport complexes is 4:1 for complex I, 4:1 for complex III, and 2:1 for complex IV.These measurment can be used to calculate the ratio of molecules of ADP phosphorylated to atoms of oxygen reduced, called P:O ratio. • • • 1. 2. When mitochondriaNADH is the substrate for the respiratory electron chain, 10 protnsare exported to the cytosol, and the P:O ratiois 10÷4=2.5 The P:O ratio for succinateis 6÷4=1.5 These P:O values should be consider maximum values- the effeciency of energy conservation is not 100% because: protons can slowly leak back into moitochondria the adenine nucleotide translocase and other transport processes consume the energy of the proton concentration gradient. Contributes its electrons at a lower energy level Uncouplers and Inhibitors Much of our knowledge of mitochondrial function results from the study of toxic compounds. Specific inhibitors were used to distinguish the electron transport system from the phosphorylation system and helped to define the sequence of redox carriers along the respiratory chain. If the chain is blocked then all the intermediates on the substrate side of the block become more reduced, while all those on the oxygen side become more oxidised. It is easy to see what has happened because the oxidised and reduced carriers often differ in their spectral properties. If a variety of different inhibitors are available then many of the respiratory carriers can be placed in the correct order. There are six distinct types of poison which may affect mitochondrial function: 1) Respiratory chain inhibitors (e.g. cyanide, antimycin, rotenone & TTFA) block respiration in the presence of either ADP or uncouplers. 2) Phosphorylation inhibitors (e.g. oligomycin) abolish the burst of oxygen consumption after adding ADP, but have no effect on uncoupler-stimulated respiration. 3) Uncoupling agents (e.g. dinitrophenol, CCCP, FCCP) abolish the obligatory linkage between the respiratory chain and the phosphorylation system which is observed with intact mitochondria. 4) Transport inhibitors (e.g. atractyloside, bongkrekic acid, NEM) either prevent the export of ATP, or the import of raw materials across the the mitochondrial inner membrane. 5) Ionophores (e.g. valinomycin, nigericin) make the inner membrane permeable to compounds which are ordinarily unable to cross. 6) Krebs cycle inhibitors (e.g. arsenite, aminooxyacetate) which block one or more of the TCA cycle enzymes, or an ancillary reation. Inhibitors of ETC: 1- Antimycin A: Antimycin A binds to the Qi site of Complex III (the enzyme cytochrome c oxidoreductase), in the cytochrome b subunit. The inhibition of Complex III by Antimycin A result in the formation of large quantities of the toxic free radical, Superoxide. Antimycin blocks the flow of electrons from semiquinone to ubiquinone in the Q-cycle of complex III in oxidative phosphorylation. By doing so it inhibits the electron transport pathway thus preventing the consumption of oxygen (which occurs at Complex IV) and disrupting the proton gradient across the inner membrane. It is the disruption of the proton gradient that prevents the production of ATP as protons are unable to flow through the ATP synthase complex. 2- Rotenone: works by interfering with the electron transport chain in mitochondria. Specifically, it inhibits the transfer of electrons from Fe-S centers in Complex I to ubiquinone. This prevents NADH from being converted into usable cellular energy (ATP). 3- 2,4-Dinitropheno Ionophores that disrupt the proton gradient by carrying protons across the membrane. This uncouples proton pumping from ATP synthesis.[3] Ionophores that disrupt the proton gradient by carrying protons across the membrane. This uncouples proton pumping from ATP synthesis.[3] 4- Malonate: is a powerful inhibitor of cellular respiration, because it binds to the active site of the succinate dehydrogenase in the citric acid cycle but does not react, since it does not have the -CH2-CH2- group (as in succinate) which is required for dehydrogenation. It is the only enzyme that participates in both the citric acid cycle and the mitochondrial electron transport chain (in this role it is often called Complex II). 5- oligimycin: inhibits ATP synthase by blocking its proton channel, which is necessary for oxidative phosphorylation of ADP to ATP (energy production). 6- Cyanide, Carbon monoxide: Inhibit the electron transport chain by binding more strongly than oxygen to the Fe–Cu center in cytochrome c oxidase, preventing the reduction of oxygen. 7- C-Ceramide: Ceramide is a lipid second messenger that mediates the effects of tumor necrosis factor and other agents on cell growth and differentiation. Ceramide is believed to act via activation of protein phosphatase, prolinedirected protein kinase, or protein kinase C An investigation of the site of ceramide action revealed that the activity of respiratory chain complex III is reduced by C2-ceramide with halfmaximum effect at 5-7 µM. In contrast, N-acetylsphinganine (C2dihydroceramide) 10th home work Name a poison that can keep ADP from exchanging with ATP. • What happens to the energy change in the cellwhen this poison is working • Does electron flow speed up , stay the same or what? 11th home work The C subunits of the Fo component of Fo F1 ATP synthase form an ion channel across the inner mitochondrial membrane. When certain glutamate aspartate residues of a C subunit react eith dicyclohexylcarbodiimide (DCCD), the subunit is unable to participitate in proton transport. (a) What is the effect of DCCD on electron transport and respiration in suspentions of intact mitochondria? (b) What happens when dinitrophenol is subsequently added to DCCD treated mitochondria? Electron Shuttle System ELECTRON SHUTTLE SYSTEMS •In our discussion of glycolysis, you learned that under aerobic conditions, NADH produced during glycolysis must be oxidized by the mitochondria. •In this way, the cell regenerates NAD+ for the glyceraldehyde-3-phosphate dehydrogenase reaction and produces additional energy by feeding electrons to the respiratory chain. •Since there is no transporter to move NADH directly into the matrix, oxidation of cytoplasmic NADH by mitochondria must occur indirectly either via the malate-aspartate or the αglycerol phosphate electron shuttle. The malate – aspartate shuttle The malate-aspartate shuttle (1)Reduces oxaloacetate (OAA) to malate. The αketoglutarate (KG) transporter (2)Exchanges malate for KG. Mitochondrial malate dehydrogenase (3)Generates intramitochondrial NADH by oxidation of malate to oxaloacetate. Mitochondrial aspartate aminotransferase (4) catalyzes the transfer of an amino group from glutamate (glu) to oxaloacetate to produce KG and aspartate (asp). KG is transported out on its translocase (2) and aspartate is transported out on the unidirectional aspartate translocase (5). Cytoplasmic aspartate aminotransferase (6) regenerates oxaloacetate for reaction (1) by transferring the amino group from aspartate to KG producing glutamate, which is transported into the matrix in exchange for aspartate (5). Inner Membrane Intermembrane Space Matrix Side ATP4- ADP3- Glutamate1- + H+ Adenine nucleotide translocase Aspartate1- Aspartate translocase Electrogenic Translocases Electrogenic transport systems of mitochondria. The malate-aspartate shuttle Cont. •Electrons from cytoplasmic NADH are used to reduce oxaloacetate to malate, which is then transported into the matrix. •Thus malate carries the electrons from NADH into the mitochondria. •In the matrix, NADH is produced by the oxidation of malate to oxaloacetate in the citric acid cycle via malate dehydrogenase. •This shuttle is irreversible because the transport of aspartate out of the matrix in exchange with glutamate moving into the matrix is electrogenic just as is the exchange of ATP with ADP The malate-aspartate shuttle Cont. •While these amino acids normally carry the same charge, the exchange is electrogenic because a proton neutralizes to zero the negative charge on the glutamate. •Thus aspartate1- exchanges for glutamate0 so that there is a net outward movement of a negative charge. •It is important that this transporter operate in only one direction (unidirectional) to ensure that electrons from mitochondrial NADH do not move to the cytoplasm in these tissues. Glycerol Phosphate Shuttle Glycerol Phosphate Shuttle •Thus, electrons are fed directly to coenzyme Q. Since complex I is bypassed, this shuttle produces one fewer ATP from glycolytic NADH than occurs with the malate-aspartate shuttle. •Also unlike the other shuttle, the electron carrier, glycerol 3-phosphate, never permeates the inner membrane but interacts instead with the transmembrane glycerol 3-phosphate dehydrogenase. Glycerol Phosphate Shuttle •The glycerol phosphate (GP) shuttle occurs almost exclusively in the liver. •Electrons from cytoplasmic NADH reduce DHAP to (GP), which in turn carries the electrons to the respiratory chain. •Electrons reach the respiratory chain via glycerol phosphate dehydrogenase that is bound to the inner membrane. •This enzyme contains a FAD prosthetic group, as we showed for succinic dehydrogenase of the citric acid cycle. Glycerol Phosphate Shuttle Cytoplasmic glycerol 3-phosphate dehydrogenase (1) oxidizes NADH. Glycerol 3-phosphate dehydrogenase in the inner mitochondrial membrane (2) reduces bound FAD to FADH2. Superoxide Anions the superoxide radical hydroxyl radical and hydrogen peroxide All of these species are highly toxic to cells. They are produced by flavoproteins, quinones, and iron– sulfur proteins. Almost all of the electron transport reactions produce small amounts of these reactive species, especially If superoxide radical is not rapidly removed by superoxide dismutase it will cause the breakdown of proteins and nucleic acids. The rapidity of this process is typical of electron transfer reactions. In this case, a copper ion is the only electron transfer agent bound to the enzyme. The copper ion is reduced by superoxide anion and it then reduces another molecule of The hydrogen peroxide formed can be converted to and by the action of catalase Some bacteria species are obligate anaerobes. They die in the presence of oxygen because they cannot deplete reactive oxygen species that arise as a by-product of oxidation– reduction reactions. These species do not have superoxide dismutase. All aerobic species have enzymes that scavenge reactive oxygen molecules. سبحانك اللهم وبحمدك أشهد ان ال إله اال انت استغفرك واتوب اليك أسأل هللا أن يهبكم قطرة من مغفرته ال تبقي لكم ذنبا.. ونظرة من رضاه ال تترك لكم عيبا.. وقبضة من خزائن رحمته ال تبقي لكم هما وال غما..