* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Weldon_McVean - Wellcome Trust Centre for Human Genetics
Minimal genome wikipedia , lookup
Human–animal hybrid wikipedia , lookup
X-inactivation wikipedia , lookup
Genomic library wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Pathogenomics wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Neocentromere wikipedia , lookup
SNP genotyping wikipedia , lookup
Metagenomics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genome (book) wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Copy-number variation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Public health genomics wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Microevolution wikipedia , lookup
Designer baby wikipedia , lookup
Helitron (biology) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Gene desert wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Human genome wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Genome evolution wikipedia , lookup
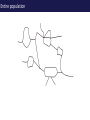
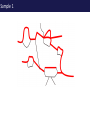
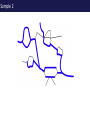
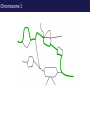
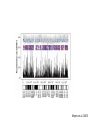
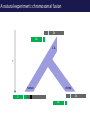
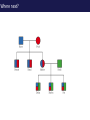
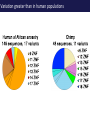
The tangled genome Gil McVean The real heroes PanMap – Genome sequencing of 10 Western Chimpanzees • Patterns of small insertion and deletion are quite different and reveal details of DNA repair pathways • Patterns of recombination in humans and chimpanzees are highly diverged at the fine-scale, but largely conserved at broad scales • There are a surprising number (6+ now ‘confirmed)’) of transspecific polymorphisms, probably maintained through hostpathogen interactions A tangle of sequence Difficulties of working with an incomplete reference Using de novo assembly to find variants EntireEntire population population Sample 1 Sample 2 Chromosome 1 Prop. sites MIC Pat Marlon Dylan Marlies Ruud Dennis Using Cortex leads to a high quality set of variants 0.6 0.6 0.5 0.5 0.4 0.4 0.3 0.3 0.2 0.2 0.1 0.1 Mendel consistency 0.0 1.0 0.8 0.6 0.4 0.2 ●●●● ●● ●●● ●●●●● ●●●● ●●●●●●●●●●●●●●●● ●●● ●● ● ●● ● ●●● ● ● ● ● ● ● ● ● ● ● ●●● ● ● ● ● ● ● ● ● ●●●●●●●●●● ● ●●●●●● ●●●●●●●●●●● ● ● ●●● ●●●● ●●●● ● ●● ● ●●● ●● ●●●●●●●●●● ● ●●●●●●●●●●●●●●●●●●●●●●●●● ● ●●●● ●●●●●●● ●●●●●●●●●●●● ●●●●●●●●● ●●●● ●●●●●●●● ●●●● ●● ●●● ●●●●●●●●●● ● ●●●●●●●●●●●●●●● ● ● ●●●●●●● ● ● ● ● ● ●●● ●●● ●● ●●● ●● ●● ●●●●● ● ● ● ● ●● ● ● ●●● ● ● ● ● ● ● ● ●●●● ● ● ● ● ● ● ● ● ● ● ● ●● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ● ●● ● ● ● ● ● ● ●● ● ● ● ● ●● ● ● ● ● ●● ● 0.0 ● 20 50 40 100 60 Chunk Chunk 150 80 200 100 Diversity in Western Chimpanzees • Similar diversity as humans of European origin (0.06%-0.08%) • Excess of common variants • 1% variants shared with humans Non-slippage indels are strongly biased to deletions 13:1 bias toward deletions. Unexpected peak at 4bp Indels as indicators of DNA repair processes Insertions deletions 25 25 20 20 Longest 15 word agreement 15 10 10 5 5 10 15 Indel size 20 25 5 5 10 15 Indel size 20 25 TGACGAACTTAT ACTGCTTGAATA TGACGA AT AC TGAATA TGACTTAT TGAC--AT ACTGAATA Losing GAAC A tangle of trees Myers et al. 2005 The zinc-finger protein PRDM9 determines hotspot location Myers et al. 2010 PRDM9 Zinc fingers are radically different between humans and chimps Perhaps the most diverged gene between humans and chimpanzees Repeatedly hit by adaptive evolution across mammals Only known ‘speciation gene’ in mammals Polymorphic in humans – leads to variation in hotspots and genome instability Questions • We know from previous work in a few regions that hotspot locations tend not to be shared between humans and chimpanzees • Calculations suggested that only 40% of human hotspots were driven by PRDM9 binding • But.. – Is there any hotspot sharing? – Do we conservation of recombination rates at any scale? – What features determine hotspot location in chimpanzees? The first genome-wide fine-scale map of recombination for a non-reference organism Auton et al. 2012 Chimpanzee recombination is dominated by hotspots in a manner similar to humans But the hotspots are not in the same locations Fine-scale profiles around genes are similar As is rate variation around CpG islands Substantial PRDM9 diversity, but overlap in predicted binding sequences No signal for predicted binding sequences Similarities at 1Mb scale Human and chimp recombination rates are correlated at the chromosomal scale Human and chimp recombination rates are only correlated at broad scales Lower correlation in structural rearrangements • All, bar one, of the inverted regions are pericentric so change in position wrt to centromere does not contribute • Change in proximity to telomere is important A natural experiment: chromosomal fusion 2b 2a C.A. t human chimp 2b 2 2a Fusion region shows 3-fold decrease in recombination rate Fusion region shows 3-fold decrease in recombination rate A tangle of histories Distribution of sickle allele Of malaria How many variants are shared through descent? Human polymorphism Chimpanzee polymorphism 9.4 million autosomal and 261,000 X chromosome SNPs from 1000 genomes Pilot 1 YRI (59 individuals) 3.8 million autosomal and 102,000 X chromosome SNPs from PanMap Pan troglogdytes verus (10 individuals) SNPs shared by humans and chimpanzees (33,906 autosomal and 527 X chromosome) reduce recurrent mutation identify potentially functional coding variants Human-chimpanzee shared coding SNPs Human-chimpanzee shared haplotypes At least two shared SNPs in 4kb with the same LD reduce artifactual sharing due to known or cryptic paralogs by filtering out SNPs with low 50 bp mappability, with high read depth, or not found in 1000 Genomes Phase 1 135 shared non-synonymous SNPs 1 shared premature stop SNP 200 shared synonymous SNPs 130 regions with shared haplotypes outside the MHC outside the MHC 7 resequenced using Sanger sequencing 8 with more than two pairs in LD Outside of the MHC, six clear-cut cases of trans-species polymorphisms FREM3/GYPE MTRR All non-coding and putatively regulatory IGFBP7 In intron of IGFBP7 20kb IGFBP7 gene structure 4kb Human-Chimpanzee shared SNPs Regulatory region in HUVEC Chromatin state segmentation by HMM DNaseI hypersensitive sites Weak enhancer Strong enhancer Regulatory region in NHEK and HMEC Strong enhancer Weak enhancer Weak enhancer Open chromatin by FAIRE TFBS conserved in human/mouse/rat TFBS identified by ChIP-seq SRF ISGF-3 GATA-2 Average pairwise differences Primate phastCons score CUTL1 RelA Bach1 STAT3 • In total, 130 regions with shared human-chimpanzee haplotypes. Six clear-cut cases of ancient balanced polymorphisms. • None are protein-coding. Eleven occur in non-coding genes (e.g., 7 in lincRNAs). Eleven compelling cases of regulatory regions. • What do these regions have in common? SNPs shared by humans and chimpanzees Shared haplotypes Glycoproteins Closest gene within 20 kb of a human-chimp shared haplotype (n=26, p=2x10-5, FDR=0.03) Shared coding SNPs Glycoproteins Genes human-chimp coding shared SNP (n=99, p=0.017, FDR=0.20) Enrichment of membrane glycoproteins -> host-pathogen interactions Project Participants • • University of Oxford Adam Auton Rory Bowden Peter Humburg Zam Iqbal Gerton Lunter Julian Maller Simon Myers Susanne Pfeifer Isaac Turner Oliver Venn Peter Donnelly (PI) Gil McVean (PI) Biomedical Primate Research Centre Ronald Bontrop • University of Chicago Adi Fledel-Alon Ryan Hernandez (UCSF) Ellen Leffler Cord Melton Laure Segurel Molly Przeworski (PI) • Funders Howard Hughes Medical Institute National Institute of Health Royal Society Wellcome Trust Where next? Remarkable structural and sequence diversity in chimp PRDM9 Variation greater than in human populations Little correlation in fine-scale structure around DNA repeat elements No activating motif discovered in chimp CCTCCCT