* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Protein Folding
Photosynthetic reaction centre wikipedia , lookup
Size-exclusion chromatography wikipedia , lookup
Biochemical cascade wikipedia , lookup
Point mutation wikipedia , lookup
Gene expression wikipedia , lookup
Signal transduction wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Paracrine signalling wikipedia , lookup
Magnesium transporter wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Expression vector wikipedia , lookup
Biochemistry wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Metalloprotein wikipedia , lookup
Homology modeling wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Interactome wikipedia , lookup
Western blot wikipedia , lookup
Protein purification wikipedia , lookup
Two-hybrid screening wikipedia , lookup
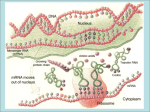
Self-Organizing Biostructures NB2-2007 L.Duroux Lecture 6 1. Protein Folding (Proteins 2nd Ed., T.E. Creighton) 2. Protein Quaternary Structure 1. Protein Folding Another case of essential self-assembly process Protein folding is essential to life Why is the “Protein Folding” so Important? Proteins play important roles in living organisms. Some proteins are deeply related with diseases. And structural information of a protein is necessary to explain and predict its gene function as well as to design molecules that bind to the protein in drug design. Today, whole genome sequences (the complete set of genes) of various organisms have been deciphered and we realize that functions of many genes are unknown and some are related with diseases. Therefore, understanding of protein folding helps us to investigate the functions of these genes and to design useful drugs against the diseases efficiently. In addition to that, the understanding opens the door to designing of proteins having novel functions as new nano machines. 1a. Examples Protein (mis)folding can lead to fatal diseases Mad cow disease, or bovine spongiform encephalopathy (BSE), is a fatal brain disorder that occurs in cattle. Abnormal protein folding is considered crucial to the onset of the disease. What causes mad cow? To illustrate the concept of protein folding we chose villin, a protein which exists in the stomach and intestine of animals (including homo sapiens). Why do proteins fold? What causes mad cow disease? Bovine epidemic in UK (1986): 170 000 cows died Symptoms: “mad”, aggressive, nervous, spongiform encephalopathy Other examples: scrapie (sheep), CreutzfeldJacob Disease (humans) S. Prusiner (1982): Infectious agent are “proteinaceous infectious particles” = prions Prions: proteins found in the nerve cells of all mammals. Abnormally-shaped prions found in BSE-infected cows The difference in normal and infectious prions may lie in the way they fold Brain surface of CJD patient on autopsy showing sponge-like appearance Prions, infection and folds. 1. 2. Native Infectious Contamination: Ingestion / Genetics Bloodstream nervous system. 3. Molecular interaction Infectious / Native change in conformation of native ( Infectious) 4. Accumulation of Infectious form in fibrillates (self-assembly) 5. Internalization/vesicles clogging cell death 6. Release Infectious form 7. Large, sponge-like holes : spongiform encephalopathy Villin headpiece sub-domain: a study case for protein folding Villin’s function: structure to intestinal villi stabilizes bundles of actin filaments folds recognized by specific receptor point of actin filaments Folding Simulated by distributed dynamics (Folding@home) one and only one way of folding is the correct way. 1b. Folding mechanisms Proteins Can Fold into 3D Structures Spontaneously The three-dimensional structure of a protein is self-organized in solution. The structure corresponds to the state with the lowest free energy of the protein-solvent system. (Anfinsen’s dogma) If we can calculate the energy of the system precisely, it is possible to predict the structure of the protein! Anfinsen experiment: Spontaneous renaturation of Ribonuclease A Primary structure contains sufficient information to allow formation of secondary and tertiary structures Fig. 4.29 Levinthal Paradox We assume that there are three conformations for each amino acid (ex. -helix, β-sheet and random coil). If a protein is made up of 100 amino acid residues, a total number of conformations is 3100 = 515377520732011331036461129765621272702107522001 ≒ 5 x 1047 If 100 psec (10-10 sec) were required to convert from a conformation to another one, a random search of all conformations would require 5 x 1047 x 10-10 sec ≒ 1.6 x 1030 years However, folding of proteins takes place in msec to sec order. Therefore, proteins fold not via a random search but a more sophisticated search process. Is it possible to watch the folding process of a protein using molecular simulation techniques? Time Scales of Protein Motions Permeation of an ion in Porin channel Elastic vibrations of proteins α-Helix folding β-Hairpin folding Bond stretching Protein folding 10-15 10-12 10-9 10-6 10-3 100 (fs) (ps) (ns) (μs) (ms) (s) Time Forces Involved in the Protein Folding Electrostatic interactions van der Waals interactions Hydrogen bonds Hydrophobic interactions (Entropy driven, role of water) Protein folding hierarchy a) Formation of secondary structure elements b) Hydrophobic colapse – molten globule – compact intermediate with high content of secondary structure elements c) Native contacts formation d) In case of multi-domain proteins: interdomain organization. e) Out of pathway intermediates: misfolded proteins – formation of nonative disulfide bonds - Proline cis-> isomerisation: Protein folding mechanisms The next few slides show four different protein folding mechanisms currently known These mechanisms describe different possible sequences and paths, shown with arrows, that the chains of amino acids can follow to go from the unfolded state to the final protein form, called the native state Diffusion/Collision • First form secondary structure by diffusion/collision • Hierarchical: form helices & hairpins, then microdomains, decrease entropy unfolded state formation of microdomains diffusion and collision of microdomains native state Nucleation unfolded state Nucleation Form nucleus of structure, then grow (ala 1st order phase transition) formation of a nucleus native state Collapse Collapse first Hydrophobically driven: remove water to form hydrogen bonds unfolded state collapse native state Topomer search unfolded state Form rough native shape first (topomer search) "topomer" Find the right “topology” first, then pack side chains native state Evolution will use any mechanism that works! No single mechanism is observed, different examples appear in nature Form secondary structure first (BBA5) Collapse first (protein G Hairpin) Hierarchical: form alpha-helices & beta-sheets Hydrophobically driven: remove water to form hydrogen bonds first Form rough native shape first (Villin) 1c. Energetic Considerations Importance of kinetic factors during folding Observed folded conformation not necessarily the most thermodynamically stable Folded conformation = the most kinetically accessible Not necessarily a pathway to lowest potential energy Energy landscapes in protein folding pathways Many paths lead to the lowest energy state that represents the native protein. Protein folding dictated by primary structure Multiple intermediate steps Important driving forces: Hydrophobic effect Hydrogen bonding Van der Waals Charge-charge The pathways for protein folding On these pathways, the protein molecules would pass through welldefined partially structured states, some of which could be transient, but others would be populated significantly Similar to Reaction of small molecules: specific pathway and small region of conformational space, so Levinthal paradox is avoided Supported existence of partially folded intermediates formed both during folding and under partially denaturing conditions Recent studies: the behavior of different proteins often appears quite distinct: some involves well-defined compact intermediates, whilst others are effectively a two-state reaction Energy Surfaces, Energy Landscapes Based on A description of statistical ensembles and emphases the difference between the folding reactions A major distinguishing feature of PF is the extreme heterogeneity of reaction and the complex interplay between the entropic and elthalpic contributions to the free energy of system Denatured protein usually resembles a “random coil”, in which local interactions dominate the conformational behavior. Extremely heterogeneous, both globally and at the level of individual residues. Nearly Levinthal Paradox The enthalpies difference of the denatured and folded protein are on the order of 30-100kcal/mol 1eV=22.9kcal/mol=96.32kJ/mol~11560K; H-bond 20kJ/mol A schematic energy landscape for protein folding. The surface is derived from a computer simulation of the folding of a highly simplified model of a small protein. The surface 'funnels' the multitude of denatured conformations to the unique native structure. The critical region on a simple surface such as this one is the saddle point corresponding to the transition state, the barrier that all molecules must cross if they are to fold to the native state. Superimposed on this schematic surface are ensembles of structures corresponding to different stages of the folding process. The transition state ensemble was calculated by using computer simulations constrained by experimental data from mutational studies of acylphosphatase. Molten Globule An intermediate state in the folding of protein pathway of a protein that has some secondary and tertiary structure, but lacks the well packed amino acid side chains that characterize the native state of a protein. Observed for many protein under both equilibrium and non-equilibrium conditions. By contrast, for fast folding proteins without intermediates, the search for a core or nucleus is likely to be the ratedetermine step; once the core is formed, folding to the native state is fast A Unified Mechanism of Protein Folding? The mechanism developed by considering the free energy surfaces for reaction provide immediate insight into how the Levinthal paradox is overcome. Each folding trajectory is different: depending both on starting point and on the stochastic nature of the folding process The overall folding behavior can be changed drastically by relatively small changes in the model parameter Simulations shows that: Fast 2-states folding can occur when collapse involves only a small subset of highly stabilizing native contacts in a core region or nucleus for large protein, long range contacts are important; cooperativity between the shortrange initiation and long range contacts lead to efficient folding. (In fact, helical protein tend to fold faster than b sheet protein) A core in large systems may occur independently in different regions, resulting additional complexities in folding, including the formation of partially structured intermediates and the possibility of extreme heterogeneity in the folding kinetics Uniform (Hydrophobic) residues often rapidly collapse to a disorganized globule with the slow step in folding corresponding to reorganization events within a compact ensemble of states, especially in large lattices. Some core residues are important and have been conserved during evolution 1d. Molecular Chaperones A case of natural kinetic control in protein folding Molecular chaperones Increase the rate of correct folding of nascent polypeptide chains Aid in the assembly of multisubunit proteins Protect proteins from stress-induced damage (eg. Heat shock) Chaperonin GroEL/GroES Chaperonine from E. coli Multisubunit protein comples GroEL – cis and trans ring 7 fold symetry, cis ring binds 7 molecules of ATP Cis ring hydrolyses ATP and undergoes conformatinal changes resulting in increase of cis ring cavity GroES – dome like hectameric ring GroEL/GroES – assists only sa subset of protein folding these proteins contains /b secondary structures Gro ES Gro EL Cis-ring Gro EL Trans-ring Molecular chaperones assist protein folding Mechanism of chaperon action 1. ATP molecules and misfolded protein binds to chaperonin through hydrophobic interactions 2. GroES binds to GroEL resulting in changes of GroEL cis ring structure, changes in misfolded protein- cavity interactions 3. Hydroglyses 7 ATP molecules 4. Binding 7 ATP to trans ring and concomitant release of folded protein, ADP molecules and GroES from cis ring, binding of misfolded protein to trans ring 5. Cis ring becomes trans ring and cycle can repeat 1e. Protein folding predictions Molecular Dynamics (MD) In molecular dynamics simulation, we simulate motions of atoms as a function of time according to Newton’s equation of motion. The equations for a system consisting on N atoms can be written as: d ri t 2 mi dt 2 Fi t , (i 1, 2, , N ). (1) Here, ri and mi represent the position and mass of atom i and Fi(t) is the force on atom i at time t. Fi(t) is given by Fi iV r1 , r2 , , rN , (2) where V(r1, r2, …, rN) is the potential energy of the system that depends on the positions of the N atoms in the system. ∇i is i i j k x y z (3) Integration Using a Finite Difference Method The positions at times (t + Δt ) and (t − Δt ) can be written using the Taylor expansion around time t, 1 1 2 3 4 ri t t ri t ri t t ri t t ri t t O t , 2 6 1 1 2 3 4 ri t t ri t ri t t ri t t ri t t O t . 2 6 The sum of two equations is ri t t ri t t 2ri t ri t t O t . 2 4 (4a) (4b) (5) Using eq. (1), the following equation is obtained: ri t t 2ri t ri t t 1 2 4 Fi t t O t . mi (6) We should calculate eq. (6) iteratively to obtain trajectories of atoms in the system (Verlet algorithm). Energy Functions used in Molecular Simulation Φ r Θ Bond stretching term Angle bending term Vtotal Dihedral term K r r K K 1 cosn 2 b 2 0 bonds angles dihedrals Cij Dij 12 10 van der Waals r Hbonds rij ij i , j pairs H-bonding term O r H 0 Aij Bij qi q j 12 6 r electrosta tic r r ij ij i , j pairs ij Van der Waals term r The most time demanding part. Electrostatic term + r ー System for MD Simulations Without water molecules With water molecules # of atoms: 304 # of atoms: 304 + 7,377 = 7,681 MD Requires Huge Computational Cost Time step of MD (Δt) is limited up to about 1 fsec (10-15 sec). ← The size of Δt should be approximately one-tenth the time of the fastest motion in the system. For simulation of a protein, because bond stretching motions of light atoms (ex. O-H, C-H), whose periods are about 10-14 sec, are the fastest motions in the system for biomolecular simulations, Δt is usually set to about 1 fsec. Huge number of water molecules have to be used in biomolecular MD simulations. ← The number of atom-pairs evaluated for non-bonded interactions (van der Waals, electrostatic interactions) increases in order of N 2 (N is the number of atoms). It is difficult to simulate for long time. Usually a few tens of nanoseconds simulation is performed. Time Scales of Protein Motions and MD Permeation of an ion in Porin channel Elastic vibrations of proteins α-Helix folding β-Hairpin folding Bond stretching Protein folding 10-15 10-12 10-9 10-6 10-3 100 (fs) (ps) (ns) (μs) (ms) (s) MD Time It is still difficult to simulate a whole process of a protein folding using the conventional MD method. To perform MD simulations parallelization is the key Special-purpose computer Calculation of non-bonded interactions is performed using the special chip that is developed only for this purpose. For example; MDM (Molecular Dynamics Machine) or MD-Grape: RIKEN MD Engine: Taisho Pharmaceutical Co., and Fuji Xerox Co. Parallelization A single job is divided into several smaller ones and they are calculated on multi CPUs simultaneously. Today, almost MD programs for biomolecular simulations (ex. AMBER, CHARMm, GROMOS, NAMD, MARBLE, etc) can run on parallel computers. Brownian Dynamics (BD) The dynamic contributions of the solvent are incorporated as a dissipative random force (Einstein’s derivation on 1905). Therefore, water molecules are not treated explicitly Since BD algorithm is derived under the conditions that solvent damping is large and the inertial memory is lost in a very short time, longer time-steps can be used BD method is suitable for long time simulation. The folding of Villin headpiece subdomain Solved using Molecular Dynamics simulations with massively parallelized computation: distributed dynamics with Folding@home 2. Protein Quaternary Structures Levels of protein structure Primary Secondary Tertiary Quaternary Quaternary structure Quaternary structure refers to the organization and arrangement of subunits in a protein with multiple subunits Same physical forces involved than in intramolecular interactions in monomeric proteins (also disulfides, metal coordination...) Quaternary structure Can have more than two subunits Subunits are individual polypeptides Pyruvate dehydrogenase complex: 60 subunits! The flagella assembly of Salmonella sp.