* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Supplemental Data
Genome evolution wikipedia , lookup
Maximum parsimony (phylogenetics) wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Gene nomenclature wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Protein moonlighting wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
History of genetic engineering wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Microevolution wikipedia , lookup
Computational phylogenetics wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene expression programming wikipedia , lookup
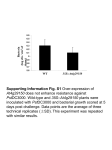
Supplementary Figures. Figure S1 Phylogenetic tree of subgroup I the GT47 family. Phylogenetic tree constructed using the MEGA program including the Arabidopsis GT47 family proteins together with the 5 members of the human EXT family (EXT1, EXT2, EXTL1, EXTL2 and EXTL3). Figure S2 Phylogenetic tree of the GT43 family members from Arabidopsis and other species. Phylogenetic relationship between the Arabidopsis IRX9, IRX9-L, IRX14, IRX14-L proteins, homologs from rice, poplar and cotton and the animal GlcAT-I, GlcAT-P, GlcAT-S proteins. The tree was made using the MEGA programme. Figure S3. RT-PCR analysis of the irx9 and irx9-L1 alleles RT-PCR was performed on RNA prepared from irx9-L1, irx9 and irx9-L1 irx9 plants using primers specific for IRX9-L and IRX9. Primers for the 18S rRNA were used as a control. Figure S4 Gene expression profiles obtained from the Genevestigator resource. The expression pattern of Meta-Analyzer data were obtained from AtGenExpress at the Genevestigator site (https://www.genevestigator.ethz.ch/). Signal intensities of different organs and stages from weak to strong were labelled from white to deep blue. Figure S5 Overexpression of minor function GX synthesis genes is able to restore the phenotype of the mutant in the homologous major function gene. Radial sections of stems from wild type (A), irx9 (B), irx14 (D), fra8 (F), irx9 transformed with 35S:IRX9-L (C), irx14 transformed with 35S:IRX14-L (E), fra8 transformed with 35S:F8H (G). Scale bar, 20 m Figure S6 Complementation of the irx10 mutant by overexpression of IRX10-L. (A) RT-PCR detection of IRX10-L (upper panel) and IRX10 (middle panel) transcripts in wild-type, irx10 and 35S:IRX10-L expressing lines in an irx10 mutant background. The expression of the 18S rRNA gene was used as a loading control (lower panel). 1 (B-D) Restoration of primary xylem vessels in the transgenic Arabidopsis irx10 plants overexpressing the IRX10-L gene. Wild type (B), irx10 (C) and 35S:IRX10-L (irx10 mutant background) (D). Scale bar, 20 m Figure S7 Phenotypes of double mutant combinations between major and minor function genes. (A) Wild-type, (B) irx10 irx14-L, (C) irx14 irx9-L and (D) irx14 irx10-L, (E) irx9, (F) irx10-L, (G) irx9 irx14-L and (H) irx9 irx10 5 week old soil grown plants. (I) From left to right are wt, irx10-L f8h, irx10 f8h. (J) A magnified view of the floral tissues of irx10 f8h which are highlighted in a light grey box in (I). Figure S8 Reduction in cell wall xylose content is correlated with the severity of the mutant phenotype. Cell wall xylose composition of stems of wild-type, irx9-L, irx14-L, f8h, irx9, irx14, fra8, irx7, irx9 irx9-L, irx14 irx14-L, fra8 f8h and irx10 irx10-L. Standard error bars are shown (n=3). 2