* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Inducible uptake and metabolism of glucose by the phosphorylative
Survey
Document related concepts
Basal metabolic rate wikipedia , lookup
Cryobiology wikipedia , lookup
Isotopic labeling wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Paracrine signalling wikipedia , lookup
Biosynthesis wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Microbial metabolism wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biochemical cascade wikipedia , lookup
Citric acid cycle wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Glyceroneogenesis wikipedia , lookup
Phosphorylation wikipedia , lookup
Transcript
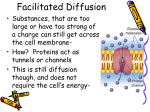
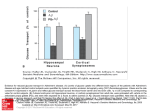
Inducible uptake and metabolism of glucose by the phosphorylative pathway in Pseudomonas putida CSV86 Aditya Basu & Prashant S. Phale Biotechnology group, School of Biosciences and Bioengineering, Indian Institute of Technology-Bombay, Powai, Mumbai, India Correspondence: Prashant S. Phale, Biotechnology Group, School of Biosciences and Bioengineering, Indian Institute of Technology-Bombay, Powai, Mumbai 400 076, India. Tel.: 191 22 25767836; fax: 191 22 25723480; e-mail: [email protected] Received 28 January 2006; revised 19 April 2006; accepted 20 April 2006. First published online May 2006. doi:10.1111/j.1574-6968.2006.00285.x Editor: Wilfrid Mitchell Keywords Pseudomonas ; glucose metabolism; transport; enzyme induction; aromatic compound catabolism. Abstract Pseudomonas putida CSV86 utilizes glucose, naphthalene, methylnaphthalene, benzyl alcohol and benzoate as the sole source of carbon and energy. Compared with glucose, cells grew faster on aromatic compounds as well as on organic acids. The organism failed to grow on gluconate, 2-ketogluconate, fructose and mannitol. Whole-cell oxygen uptake, enzyme activity and metabolic studies suggest that in strain CSV86 glucose utilization is exclusively by the intracellular phosphorylative pathway, while in Stenotrophomonas maltophilia CSV89 and P. putida KT2442 glucose is metabolized by both direct oxidative and indirect phosphorylative pathways. Cells grown on glucose showed five- to sixfold higher activity of glucose-6-phosphate dehydrogenase compared with cells grown on aromatic compounds or organic acids as the carbon source. Study of [14C]glucose uptake by whole cells indicates that the glucose is taken up by active transport. Metabolic and transport studies clearly demonstrate that glucose metabolism is suppressed when strain CSV86 is grown on aromatic compounds or organic acids. Introduction A variety of aromatic compounds are present in fresh and marine waters and in agricultural lands. The most effective and economical way to remove these compounds from the environment is by microbial degradation (Alexander, 1981). The bottleneck in the efficient degradation of these compounds is the presence of simple carbon sources like organic acids and sugars, which are preferentially utilized by microorganisms, and unless the simpler carbon sources are completely depleted, the toxic aromatic compounds are not degraded (Collier et al., 1996). In pseudomonads, glucose utilization follows two routes: (i) the direct oxidative pathway, which converts glucose to gluconate, 2-ketogluconate and then subsequently to 6-phosphogluconate by extracellular, high affinity, glucose dehydrogenase and gluconate dehydrogenase (Quay et al., 1972; Midgley & Dawes, 1973; Roberts et al., 1973; Lessie et al., 1979), and (ii) the intracellular, low affinity, nucleotide-dependent phosphorylative pathway (Lessie & Neidhardt, 1967; Tiwari & Campbell, 1969; Eisenberg et al., 1974; Guymon & Eagon, 1974) wherein glucose is converted to 6-phosphogluoconate by glucokinase and glucose 6-phosphate dehydrogenase (Zwf). The key intermediate for both pathways is 6-phosphogluconate, which enters the TCA cycle via glyceraldehyde-3FEMS Microbiol Lett 259 (2006) 311–316 phosphate and pyruvate through the Entner-Doudoroff pathway (Fig. 1a) (Entner & Doudoroff, 1952; Tiwari & Campbell, 1969). Depending on the physiological conditions, one or other of the pathways predominates (Lessie & Phibbs, 1984). It has been reported that degradation of aromatic compounds is repressed by glucose as well as by organic acids (Worsey & Williams, 1975; Duetz et al., 1994; Holtel et al., 1994; Schleissner et al., 1994; Muller et al., 1996, 1997; McFall et al., 1997). Such preferential utilization of simple carbon sources represses degradation of recalcitrant compounds in nature (referred to as carbon catabolite repression). Attempts have been made to overcome this repression by generating mutants defective in glucose utilization which will mineralize complex carbon sources like naphthalene efficiently even in the presence of glucose (Samanta et al., 2001). Pseudomonas putida CSV86 has been shown to metabolize aromatic compounds preferentially over glucose and cometabolize aromatic compounds and organic acids (Basu et al., 2006). Here we report that strain CSV86 utilizes glucose only by the intracellular, phosphorylative pathway. The strain utilizes aromatic compounds and organic acids faster compared with glucose, and the glucose-metabolizing 2006 Federation of European Microbiological Societies Published by Blackwell Publishing Ltd. All rights reserved c 312 (a) A. Basu and P.S. Phale Gluconate Glucose Gcd Gad Gct Glucose ATP 2-Ketogluconate Kgt 2-Ketogluconate Gluconate Gck ATP ATP Gnk Kgk 2-Keto-6-P-gluconate Glucose-6-P Kgr Zwf NAD(P)H NAD(P) 6-P-Gluconate Pgd 2-Keto-3-Deoxy-6-P-Gluconate Kdga Glyceraldehyde-3-P (b) Glucose Gluconate x Gcd Gct Glucose ATP Pyruvate x TCA Cycle 2-Ketogluconate x Gck Glucose-6-P Zwf NAD(P) 6-P-Gluconate Pgd 2-Keto-3-Deoxy-6-P-Gluconate Kdga Glyceraldehyde-3-P Pyruvate enzyme, Zwf, and glucose transport are inducible and suppressed by growth on aromatics or organic acids. Materials and methods Growth conditions Bacterial cultures used in this study were P. putida CSV86 (Mahajan et al., 1994), Stenotrophomonas maltophilia CSV89 (Phale et al., 1995) and P. putida KT2442. Cultures were grown in 150 mL mineral salt medium [MSM; Basu et al., 2003] at 30 1C on a rotary shaker (200 r.p.m.). The medium was supplemented aseptically with appropriate amounts of aromatic compounds (0.1%), glucose (0.25%) or organic acids (0.25%) as carbon source. Growth was monitored spectrophotometrically at 540 nm. Metabolism of glucose Glucose grown mid-log phase cells (200 mg) were harvested, washed twice with sterile distilled water, suspended in 50 mL of 10 mM glucose prepared in sterile distilled water 2006 Federation of European Microbiological Societies Published by Blackwell Publishing Ltd. All rights reserved c TCA Cycle Fig. 1. Metabolic pathways involved in the utilization of glucose by (a) pseudomonads and (b) Pseudomonas putida CSV86. The enzymes involved are: Gcd, Gad: glucose and gluconate oxidase; Gct, Kgt: glucose- and 2-ketogluconate-transporter; Gck, Gnk, Kgk: glucose-, gluconate- and 2-ketogluconate-kinase; Zwf: glucose 6-phosphate dehydrogenase; Kgr: 2-keto 6-phospho gluconate reductase; Pgd: 6-phosphogluconate dehydratase; Kdga: 2Keto-3-deoxy-6-phospho gluconate aldolase Cross indicates the inability of strain to utilize gluconate and 2-ketogluconate as carbon source. and incubated at 30 1C for 3.5 h. The pH of the suspension was constantly monitored and maintained at pH 7.0 with KOH (6 M). After incubation, the cells were removed by centrifugation. The supernatant was filtered through a 0.2 mm filter and lyophilized to a dry powder. Products formed from glucose were resolved by TLC (one dimension) and detected as described (Pujol & Kado, 2000). Whole-cell oxygen uptake Mid-log phase cells grown on the appropriate carbon source were used. Respiration rates were measured at 30 1C using oxygraph (Hansatech, UK) fitted with a Clark-type O2 electrode as described earlier (Basu et al., 2003). Respiration rates were corrected for endogenous O2 consumption and expressed as nmol O2 consumed min1 mg1 of cells (wet weight). Preparation of cell-free extracts and enzyme assays Cell-free extracts were prepared as described earlier (Basu et al., 2003). Protein was estimated using folin-phenol FEMS Microbiol Lett 259 (2006) 311–316 313 Glucose metabolism in P. putida CSV86 reagent (Lowry et al., 1951). Glucose dehydrogenase (Matsushita & Ameyama, 1982), gluconate dehydrogenase (Matsushita et al., 1982) and glucose-6-phosphate dehydrogenase (Lessmann et al., 1975) activities were monitored as described. Enzyme activities are expressed either as nanomoles of substrate consumed or product formed or NADH formed or consumed per min. Specific activities are expressed as nmoles per min per mg of protein. Log OD 540 nm 10 1 0.1 [14C]Glucose uptake, binding assay Uptake of [U-14C]glucose was studied by modifying the method described previously (Sly et al., 1993). Cells were grown till late-log phase, harvested, washed twice and resuspended in MSM to an absorbance of 0.20 at 540 nm. Cell suspension was incubated at 30 1C for 10 min in a shaking water bath. To the prewarmed cell suspension (10 mL), 5 nmol of [14C]glucose (BRIT, India, sp. ac. 140 mCi mmol1) was added, and samples (100 mL) were withdrawn and rapidly filtered through 0.45 mm cellulose ester filters (Pall). The filters were immediately washed twice with sterile MSM (1 mL), air dried and vigorously mixed in scintillation cocktail (0.4% PPO and 0.025% POPOP in toluene). Radioactivity was measured using a liquid scintillation counter (Rackbeta LKB1209) and expressed as pmoles [14C]glucose accumulated. All the experiments were performed at least three times in triplicate and the observed standard deviation was less than 5%. Results and discussion Pseudomonas putida CSV86 metabolizes naphthalene via the catechol meta-cleavage pathway, and benzyl alcohol via the catechol ortho-cleavage pathway (Mahajan et al., 1994; Basu et al., 2003). It also utilizes benzoic acid, and p-and o-hydroxy benzoic acid and p-and o-hydroxybenzyl alcohols (Basu et al., 2003). Besides aromatic compounds, strain CSV86 also utilizes glucose and glycerol, but it failed to grow on gluconate, 2-ketogluconate, fructose and mannitol. The growth profile on various carbon sources is shown in Fig 2. The observed specific growth rate (m, h1) and doubling time (h) on various carbon sources were found to be: naphthalene (0.51, 1.36), salicylate (0.34, 2.04), benzyl alcohol (0.54, 1.28), benzoate (0.59, 1.17) glucose (0.22, 3.15) and succinate (0.76, 0.91). Growth profile and kinetic analysis clearly demonstrated that CSV86 utilized aromatic compounds and organic acids faster than glucose. The observed m and doubling time values on glucose for strains S. maltophilia CSV89 and P. putida KT2442 were 0.52 and 1.33; and 0.55 and 1.26, respectively, indicating that strain CSV86 utilizes glucose at a lower rate. To elucidate the glucose metabolic pathway, O2 uptake and enzyme activity studies were carried out. Strain CSV86 FEMS Microbiol Lett 259 (2006) 311–316 0.01 0 5 10 15 Time (h) 20 25 Fig. 2. Growth profile of Pseudomonas putida CSV86 on naphthalene ( ), salicylate (,), benzyl alcohol (&), benzoate (}), glucose (n) and succinate ( ). grown on glucose showed O2 uptake with glucose but failed to respire on gluconate and 2-ketogluconate. Naphthaleneand succinate-grown cells showed poor O2 uptake when incubated with glucose. Strains CSV89 and KT2442 showed O2 uptake in the presence of glucose, gluconate and 2ketogluconate (Table 1). These results suggest that, unlike the other strains, CSV86 does not have the ability to metabolize gluconate and 2-ketogluconate. These observations were supported by measurements of enzyme activities and analysis of the products formed during metabolism of glucose. Specific activities for various enzymes involved in glucose metabolism are depicted in Table 2. Cell-free extracts prepared from all three strains showed activity of Zwf. Compared with glucose-, succinate- and naphthalenegrown cells of CSV86 showed five- to six-fold lower activity of Zwf, suggesting that the enzyme is inducible. Strain CSV86 showed activity of glucose oxidase but failed to show activity of gluconate oxidase, while strains CSV89 and KT2442 showed activity for both enzymes. TLC analysis of metabolites produced during metabolism of glucose by CSV86 detected very little gluconate and no 2-ketogluconate; however, strains CSV89 and KT2442 showed spots corresponding to gluconate (Rf 0.18, light blue) and 2ketogluconate (Rf 0.18, dark brown), data not shown. Therefore, TLC analysis, whole-cell respiration and enzyme activity studies suggest that CSV86 does not metabolize glucose via gluconate and 2-ketogluconate, thus indicating the absence of the direct oxidative pathway. However, the presence of Zwf activity suggests that CSV86 utilizes glucose by the phosphorylative pathway. Based on these results, the proposed pathway for glucose metabolism in P. putida CSV86 is shown in Fig. 1b. In strains CSV89 and KT2442, both the phosphorylative and direct oxidative pathways appear to be operational for glucose metabolism (Fig. 1a). To study glucose transport, [14C]glucose uptake by cells grown under different conditions was monitored. Glucose2006 Federation of European Microbiological Societies Published by Blackwell Publishing Ltd. All rights reserved c 314 A. Basu and P.S. Phale Table 1. Oxygen uptake rates with various substrates for Pseudomonas putida CSV86, Stenotrophomonas maltophilia CSV89 and P. putida KT2442 O2 uptake rates (nmol min1 mg1) Pp CSV86w Sm CSV89z Pp KT2442‰ Substrate Glcz Naph Suc Glc Glc Glucose Gluconate 2-Ketogluconate 2.31 ND ND 0.4 NDk ND 0.3 ND ND 4.93 0.78 0.37 4.12 4.61 1.0 Oxygen uptake has been corrected for endogenous cell respiration. w P. putida CSV86. S. maltophilia CSV89. ‰ P. putida KT2442. z Cells were grown on: Naph, Naphthalene; Glc, Glucose; Suc, Succinate. k ND, not detected by this method. z Table 2. Specific activities of glucose metabolizing enzymes in Pseudomonas putida CSV86 grown on naphthalene (0.1%) and glucose (0.25%), and Stenotrophomonas maltophilia CSV89 and P. putida KT2442 grown on glucose (0.25%) Specific activity (nmol min1 mg1 protein) Pp CSV86 Enzyme Naph Glc Suc Sm CSV89 Pp KT2442 Glucose oxidase Gluconate oxidase Glucose-6-phosphate dehydrogenase (Zwf) 13 ND 14 15.7 ND 63.4 14 ND 11 17.1 285.9 95.1 16 25 170 grown CSV86 cells showed high rates of glucose uptake (Fig. 3). Addition of sodium azide (25 mM) after 1 min, inhibited glucose uptake, and cells preincubated for 10 min with sodium azide (25 mM) or formaldehyde (25 mM) did not show any uptake. These results demonstrate that glucose uptake in CSV86 is by active transport. Cells grown on succinic acid or naphthalene alone showed very low glucose uptake compared with glucose-grown cells. Similar results were observed when cells were grown on benzyl alcohol or pyruvate (data not shown). These results suggest that glucose transport in CSV86 is suppressed when cells are grown on organic acids or aromatic compounds. Metabolic studies clearly demonstrated that glucose is metabolized in P. putida CSV86 by the phosphorylative pathway and not by the direct oxidative pathway. It has been shown that pseudomonads sequester glucose as gluconate and under glucose limitation conditions gluconate is utilized as the carbon source (Schleissner et al., 1997). However, this is not the case with strain CSV86 as it failed to accumulate gluconate in the medium, and did not utilize it as a carbon source. Although a very small amount of gluconate was formed during glucose metabolism, the inability to utilize it as the carbon source could be due to either lack of gluconate transport or gluconate metabolizing enzymes, or both. CSV86 utilizes glucose at slower rate 2006 Federation of European Microbiological Societies Published by Blackwell Publishing Ltd. All rights reserved c [14C]Glucose uptake (pmol) ND, not detected. 30 20 10 0 0 5 10 15 20 Time (min) 25 30 35 Fig. 3. [14C]Glucose uptake by Pseudomonas putida CSV86 cells grown on naphthalene ( ), succinate (,) and glucose (&). Inhibition of [14C]glucose uptake was observed when cells were exposed to sodium azide (25 mM) after one min (arrow,B) or pretreated for 10 min with sodium azide (25 mM, ) or formaldehyde (25 mM, %). compared with aromatic compounds and organic acids. The slow utilization of glucose may be due to regulation of glucose metabolizing enzymes and/or the transport process. In strain CSV86 the activity of Zwf was found to be inducible; five- to six-fold higher activity was observed when cells were grown on glucose compared with naphthalene and FEMS Microbiol Lett 259 (2006) 311–316 315 Glucose metabolism in P. putida CSV86 succinate. The Zwf activity was 35% and 65% lower in CSV86 compared with strains CSV89 and KT2442, respectively. As reported for other pseudomonads (Midgley & Dawes, 1973), glucose transport in CSV86 was sensitive to sodium azide and formaldehyde demonstrating the active transport of glucose, while significantly reduced glucose uptake by cells grown on aromatic compounds and organic acid indicates that this transport system is also inducible. Based on these results, we conclude that in P. putida CSV86 the metabolism of glucose is regulated at least at enzyme and transport level. The absence of the direct oxidative pathway and the low activity of Zwf in CSV86 may be responsible for the slow growth rate on glucose. Few strains have been previously reported to have only the phosphorylative pathway or to be defective in glucose metabolism (Lessie & Phibbs, 1984). It has been reported that glucose and organic acid repress the utilization of various aromatic compounds by pseudomonads (Holtel et al., 1994; Schleissner et al., 1994; Muller et al., 1996; Rentz et al., 2004) and attempts have been made to improve the utilization of aromatic compounds in the presence of glucose by conjugal transfer of naphthalene and salicylate degradation genes into P. putida strains generated by mutagenesis which are deficient in glucose metabolism (Samanta et al., 2001). Such strains would utilize aromatic compounds with a higher specific growth rate compared with glucose and hence would be an asset for bioremediation. However, the viability of such strains in the environment is not known. Pseudomonas putida CSV86 is a natural isolate and grows on aromatic compounds with a higher specific growth rate than on glucose. It also showed a preference for aromatic compounds over glucose and cometabolizes aromatic compounds and organic acids (Basu et al., 2006). These unique properties, coupled with further characterization and subtle modification, may make this strain a potential candidate for bioremediation and environmental clean up. Acknowledgements A. B. thanks the University Grants Commission, India for the award of a senior research fellowship. References Alexander M (1981) Biodegradation of chemicals of environmental concern. Science 211: 132–138. Basu A, Dixit SS & Phale PS (2003) Metabolism of benzyl alcohol via catechol ortho-pathway in methylnaphthalene-degrading Pseudomonas putida CSV86. Appl Microbiol Biotechnol 62: 579–585. FEMS Microbiol Lett 259 (2006) 311–316 Basu A, Apte SK & Phale PS (2006) Preferential utilization of aromatic compounds over glucose by Pseudomonas putida CSV86. Appl Environ Microbiol 72: 2226–2230. Collier DN, Hager PW & Phibbs PV Jr (1996) Catabolite repression control in the Pseudomonads. Res Microbiol 147: 551–561. Duetz WA, Marques S, de JC, Ramos JL & van Andel JG (1994) Inducibility of the TOL catabolic pathway in Pseudomonas putida (pWW0) growing on succinate in continuous culture: evidence of carbon catabolite repression control. J Bacteriol 176: 2354–2361. Eisenberg RC, Butters SJ, Quay SC & Friedman SB (1974) Glucose uptake and phosphorylation in Pseudomonas fluorescens. J Bacteriol 120: 147–153. Entner N & Doudoroff M (1952) Glucose and gluconic acid oxidation of Pseudomonas saccharophila. J Biol Chem 196: 853–862. Guymon LF & Eagon RG (1974) Transport of glucose, gluconate, and methyl alpha-D-glucoside by Pseudomonas aeruginosa. J Bacteriol 117: 1261–1269. Holtel A, Marques S, Mohler I, Jakubzik U & Timmis KN (1994) Carbon source-dependent inhibition of xyl operon expression of the Pseudomonas putida TOL plasmid. J Bacteriol 176: 1773–1776. Lessie TG & Neidhardt FC (1967) Formation and operation of the histidine-degrading pathway in Pseudomonas aeruginosa. J Bacteriol 93: 1800–1810. Lessie TG & Phibbs PV Jr (1984) Alternative pathways of carbohydrate utilization in pseudomonads. Annu Rev Microbiol 38: 359–388. Lessie TG, Berka T & Zamanigian S (1979) Pseudomonas cepacia mutants blocked in the direct oxidative pathway of glucose degradation. J Bacteriol 139: 323–325. Lessmann D, Schimz KL & Kurz G (1975) D-glucose-6-phosphate dehydrogenase (Entner–Doudoroff enzyme) from Pseudomonas fluorescens. Purification, properties and regulation. Eur J Biochem 59: 545–559. Lowry OH, Rosebrough NJ, Farr AL & Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193: 265–275. Mahajan MC, Phale PS & Vaidyanathan CS (1994) Evidence for the involvement of multiple pathways in the biodegradation of 1- and 2-methylnaphthalene by Pseudomonas putida CSV86. Arch Microbiol 161: 425–433. Matsushita K & Ameyama M (1982) D-Glucose dehydrogenase from Pseudomonas fluorescens, membrane-bound. Methods Enzymol 89: 149–154. Matsushita K, Shinagawa E & Ameyama M (1982) D-Gluconate dehydrogenase from bacteria, 2-keto-D-gluconate-yielding, membrane-bound. Methods Enzymol 89: 187–193. McFall SM, Abraham B, Narsolis CG & Chakrabarty AM (1997) A tricarboxylic acid cycle intermediate regulating transcription of a chloroaromatic biodegradative pathway: fumaratemediated repression of the clcABD operon. J Bacteriol 179: 6729–6735. 2006 Federation of European Microbiological Societies Published by Blackwell Publishing Ltd. All rights reserved c 316 Midgley M & Dawes EA (1973) The regulation of transport of glucose and methyl alpha-glucoside in Pseudomonas aeruginosa. Biochem J 132: 141–154. Muller C, Petruschka L, Cuypers H, Burchhardt G & Herrmann H (1996) Carbon catabolite repression of phenol degradation in Pseudomonas putida is mediated by the inhibition of the activator protein PhlR. J Bacteriol 178: 2030–2036. Phale PS, Mahajan MC & Vaidyanathan CS (1995) A pathway for biodegradation of 1-naphthoic acid by Pseudomonas maltophilia CSV89. Arch Microbiol 163: 42–47. Pujol CJ & Kado CI (2000) Genetic and biochemical characterization of the pathway in Pantoea citrea leading to pink disease of pineapple. J Bacteriol 182: 2230–2237. Quay SC, Friedman SB & Eisenberg RC (1972) Gluconate regulation of glucose catabolism in Pseudomonas fluorescens. J Bacteriol 112: 291–298. Rentz JA, Alvarez PJ & Schnoor JL (2004) Repression of Pseudomonas putida phenanthrene-degrading activity by plant root extracts and exudates. Environ Microbiol 6: 574–583. Roberts BK, Midgley M & Dawes EA (1973) The metabolism of 2-oxogluconate by Pseudomonas aeruginosa. J Gen Microbiol 78: 319–329. Samanta SK, Bhushan B & Jain RK (2001) Efficiency of naphthalene and salicylate degradation by a recombinant 2006 Federation of European Microbiological Societies Published by Blackwell Publishing Ltd. All rights reserved c A. Basu and P.S. Phale Pseudomonas putida mutant strain defective in glucose metabolism. Appl Microbiol Biotechnol 55: 627–631. Schleissner C, Olivera ER, Fernandez-Valverde M & Luengo JM (1994) Aerobic catabolism of phenylacetic acid in Pseudomonas putida U: biochemical characterization of a specific phenylacetic acid transport system and formal demonstration that phenylacetyl-coenzyme A is a catabolic intermediate. J Bacteriol 176: 7667–7676. Schleissner C, Reglero A & Luengo JM (1997) Catabolism of D-glucose by Pseudomonas putida U occurs via extracellular transformation into D-gluconic acid and induction of a specific gluconate transport system. Microbiology 143: 1595–1603. Sly LM, Worobec EA, Perkins RE & Phibbs PV Jr (1993) Reconstitution of glucose uptake and chemotaxis in Pseudomonas aeruginosa glucose transport defective mutants. Can J Microbiol 39: 1079–1083. Tiwari NP & Campbell JJ (1969) Enzymatic control of the metabolic activity of Pseudomonas aeruginosa grown in glucose or succinate media. Biochim Biophys Acta 192: 395–401. Worsey MJ & Williams PA (1975) Metabolism of toluene and xylenes by Pseudomonas (putida (arvilla) mt-2): evidence for a new function of the TOL plasmid. J Bacteriol 124: 7–13. FEMS Microbiol Lett 259 (2006) 311–316