* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Esperimento di genetica 17.1
DNA vaccination wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Molecular cloning wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
Genomic library wikipedia , lookup
Genetic testing wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Point mutation wikipedia , lookup
Y chromosome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
DNA supercoil wikipedia , lookup
Designer baby wikipedia , lookup
Genetic engineering wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Genome (book) wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Deoxyribozyme wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
X-inactivation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Neocentromere wikipedia , lookup
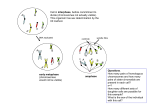
Robert J. Brooker - Genetica Esperimento di genetica 17.1 The Staining of Harlequin Chromosomes Can Reveal Recombination Between Sister Chromatids Our understanding of crossing over and homologous recombination has come from a variety of experimental approaches, including genetic, biochemical, and cytological analyses. Chromosomal staining methods have allowed researchers to visualize the genetic exchange between eukaryotic chromosomes. In the 1970s, the Russian cytogeneticist A. F. Zakharov and colleagues developed methods that improved our ability to identify chromosomes. They made the interesting observation that chromosomes labeled with the nucleotide analog 5-bromodeoxyuridine (BrdU) bind certain types of stain to a different degree compared to normal chromosomes. In 1974, Paul Perry and Sheldon Wolff extended this approach to differentially stain sister chromatids and microscopically identify SCEs. Before we consider the experiment of Perry and Wolff, let’s examine how their staining procedure allowed them to accurately distinguish the two sister chromatids. In their approach, eukaryotic cells were grown in a laboratory and exposed to BrdU for two rounds of DNA replication. After the second round of DNA replication, one of the sister chromatids in each pair contained one unla- beled strand and one BrdU-labeled strand. The other sister chromatid had two BrdU-labeled strands (Figure EG17.1.1). When treated with two dyes, Hoechst 33258 and Giemsa, the sister chromatid containing two strands with BrdU stains very weakly and appears light, whereas the sister chromatid with only one strand containing BrdU stains much more strongly and appears very dark. In this way, the two sister chromatids can be distinguished microscopically. Chromosomes stained in this way are referred to as harlequin chromosomes, because they are reminiscent of a harlequin character’s costume with its variegated pattern of light and dark patches. In these chromosomes, SCEs can be clearly identified as exchanges between light and dark chromatids. The steps in Perry and Wolff ’s protocol are shown in Figure EG17.1.2. They began with Chinese hamster ovary cells, a commonly used mammalian cell line, and exposed the cells to BrdU for two rounds of DNA replication. Near the end of the second round, colcemid was added to prevent the completion of mitosis. The cells were treated with KCl to spread out the chromosomes, which were subsequently fixed and then stained with Hoechst 33258 and Giemsa. THE HYPOTHESIS Crossing over may occur between sister chromatids. FI GU RE EG1 7 .1.1 H arle quin c hrom osom e s. © 2010 The McGraw-Hill Companies, S.r.l. - Publishing Group Italia Robert J. Brooker - Genetica Starting material: A laboratory cell line of Chinese hamster ovary (CHO) cells. FI GU RE EG1 7 .1.2 T he st a ining of ha rle quin c hrom osom e s re ve a ls sist e r c hrom a t id e xc ha nge . © 2010 The McGraw-Hill Companies, S.r.l. - Publishing Group Italia Robert J. Brooker - Genetica THE DATA INTERPRETING THE DATA Reprinted by permission from Macmillan Publishers Ltd. Nature, New Giemsa method for the differential staining of sister chromatids. Perry P. & Wolff S. 251:5471, 156–158, 1974. A micrograph of their results is shown in the data of Figure EG17.1.2. As seen here, the chromosomes show the classic harlequin appearance due to the differential staining of the sister chromatids. Furthermore, examples of SCE are clearly visible. The arrows depict regions where crossing over has taken place. In this study, Perry and Wolff found that SCEs occurred at a frequency of approximately 0,67 per chromosome. This method has provided an accurate (and dramatic) way to visualize genetic exchange between sister chromatids. Many subsequent studies have used the harlequin staining method to study the effects of agents that may influence the frequency of genetic exchanges. Researchers have found that DNA damage caused by radiation and chemical mutagens tends to increase the level of genetic exchange. When cells are exposed to these types of mutagens, the technique of harlequin staining has revealed a substantial increase in the frequency of SCEs. In addition, certain genetic disorders that result in higher levels of chromosome breakage also show elevated SCEs. For example, Bloom syndrome is a rare autosomal recessive disorder characterized by short stature, skin abnormalities, and a predisposition for developing certain forms of cancer. The defect is associated with a gene that is involved with DNA replication. In the cells of Bloom syndrome patients, chromosome breaks are more frequent during DNA replication. Likewise, SCE is typically 10- to 15-fold more frequent in Bloom syndrome patients compared to unaffected individuals. © 2010 The McGraw-Hill Companies, S.r.l. - Publishing Group Italia