* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Protein stabilization: a common consequence of mutations
Histone acetylation and deacetylation wikipedia , lookup
Protein (nutrient) wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Cytokinesis wikipedia , lookup
Magnesium transporter wikipedia , lookup
Protein moonlighting wikipedia , lookup
Signal transduction wikipedia , lookup
Biochemical switches in the cell cycle wikipedia , lookup
Protein domain wikipedia , lookup
List of types of proteins wikipedia , lookup
Phosphorylation wikipedia , lookup
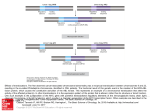
ã Oncogene (1999) 18, 7552 ± 7558 1999 Stockton Press All rights reserved 0950 ± 9232/99 $15.00 http://www.stockton-press.co.uk/onc Protein stabilization: a common consequence of mutations in independently derived v-Myc alleles Paul R Gavine1, James C Neil2 and Dorothy H Crouch*,1 1 Biomedical Research Centre, University of Dundee, Ninewells Hospital and Medical School, Dundee DD1 9SY, UK; 2Department of Veterinary Pathology, University of Glasgow Veterinary School, Bearsden, Glasgow G61 1QH, UK Myc is overexpressed in many cancers as a result of gene rearrangement or ampli®cation, but coding sequence changes which cluster in the N-terminal transactivation domain also appear to play a role in tumour progression. The prototypic v-Myc gene of MC29 virus diers from avian c-Myc by a series of mutations, including a change at a regulatory phosphorylation site within the mutational hotspot (thr-61) which is known to potentiate transformation in vitro. We now show that the mutation at thr-61 stabilizes the v-Myc protein (turnover dierence) and that this single mutation is both necessary and sucient for the phenotype. A major involvement of the proteasome in Myc degradation was con®rmed, but surprisingly, a dilysine motif adjacent to thr-61 proved not to be the ubiquitin target. Two other v-Myc genes which carry a mutation at thr-61 (avian MH2) or a large deletion encompassing this domain (feline T17) were found to be stabilized to a similar extent as MC29, showing that stabilization is a common feature of independently derived Myc oncogenes. These results suggest a common selective process in the genesis of these three viral oncoproteins and a mechanistic link with Jun, Fos and Myb oncoproteins which are also stabilized relative to their cellular counterparts. Keywords: Myc; viral mutations; stabilization Introduction Myc, originally identi®ed as the transforming component of avian leukosis virus, MC29, functions as a sequence speci®c nuclear transcription factor, working in association with its obligate partner, Max. Both dimerization with Max through the C-terminal helix ± loop ± helix/leucine zipper (HLH-LZ) domain and sequence-speci®c DNA binding mediated by the adjacent basic region (B) are prerequisites for Myc to function, and the resulting Myc : Max complex transactivates target genes through an N-terminal transactivation domain in Myc (Henriksson and Luscher, 1996). myc expression is strictly controlled in the cell ensuring regulated growth through the correct expression of its downstream target genes. Escape from this control through retroviral transduction, chromosome ampli®cation or retroviral insertion, however, results in elevated levels of Myc, aberrant gene expression and *Correspondence: DH Crouch Received 26 April 1999; revised 21 July 1999; accepted 21 July 1999 deregulated growth. Mutational changes in Myc, either somatic or virally-derived, can also contribute signi®cantly to the development of the neoplastic phenotype, and intriguingly, these are clustered in a hotspot region in the amino terminal transactivation domain (Henriksson and Luscher, 1996; Hoang et al., 1995; Smith-Sorensen et al., 1996). As a consequence of oncogenic activation, the normal signalling mechanisms which regulate cellular protooncogenes are subverted. Unravelling the molecular consequences of these activating events will therefore contribute both to our understanding of how cellular proteins are controlled by normal cell signalling and how viral oncoproteins escape this. Examples of activating mutations are deletion of key regulatory regions (d domain in Jun (Gillespie, 1991)) or mutation of speci®c phosphorylation sites (Myc, Fos (Curran et al., 1994; Henriksson et al., 1993; Okazaki and Sagata, 1995; Pulverer et al., 1994)). A common feature of c-Myc and other nuclear transcription factors (Jun, Fos) is their short half life, which ensures the maintenance of low steady state levels required for controlled growth. Degradation of these, and other cell cycle regulators and tumour suppressors, is mediated by the 26S proteasome (Ciechanover, 1998; Gross-Mesilaty et al., 1998; Musti et al., 1997), following conjugation of ubiquitin to lysine residues of the target protein (Ciechanover, 1998). Protein stabilization as a result of disruption in proteasome-directed degradation of key regulatory proteins may represent a potential mechanism of oncogenic activation since inappropriate levels of these proteins in the cell would result. Indeed, some key nuclear transcription factors are stabilized as a consequence of oncogenic activation. Both viral Fos and the leukaemia-speci®c form of c-Myb are more stable than their cellular counterparts due to carboxy terminal deletions (Bies and Wol, 1997; Curran et al., 1994). v-Jun, however, is stabilized through a deletion termed the d domain in the N-terminus, which abrogates ubiquitin dependent degradation (Treier et al., 1994). We set out to establish whether the prototypic vMyc from the avian MC29 virus is stable. We now report that MC29 v-Myc is considerably more stable than c-Myc as a consequence of a single point mutation in a major phosphorylation site, and propose that loss of phosphorylation at this site stabilizes the Myc protein. Furthermore, we show that two independently derived v-Myc isolates with analogous mutations at thr-61 (avian virus MH2) or a deletion spanning this domain (feline virus T17) are Stabilization of viral Myc proteins PR Gavine et al also stable. Finally, we propose that protein stabilization is a common mechanism of oncogenic activation of nuclear transcription factors such as Myc, Fos, Jun and Myb (Bies and Wol, 1997; Curran et al., 1994; Isaksson et al., 1996; Okazaki and Sagata, 1995). Results MC29 v-Myc oncoprotein is stable To address whether the MC29 v-Myc oncoprotein is more stable than c-Myc, we generated avian retroviral vectors which encode either v-myc sequences derived from the avian leukosis virus, MC29 (Walter et al., 1986) or avian c-Myc (La Rocca et al., 1994). The location of the MC29 v-Myc point mutations with respect to the structure of c-Myc is outlined in Figure 1a. MC29 v-Myc diers from c-Myc by ®ve point mutations, which are scattered throughout the protein (Walter et al., 1986). Signi®cantly, thr-61, found within the highly conserved N-terminal Myc Box 1 (MBI: amino acids 45 ± 63), is the most commonly mutated amino acid in tumour-derived Myc genes (Henriksson and Luscher, 1996; Hoang et al., 1995; Smith-Sorenson et al., 1996). The constructs were transfected into chick embryo ®broblasts (CEF) and subjected to G418 selection. To measure protein turnover, the resulting cultures were treated with the protein synthesis inhibitor, emetine, for the appropriate lengths of time, and Western blots used to analyse Myc degradation in the total cellular pool of Myc. This was felt to preclude dierential labelling of c-Myc and v-Myc in pulse-chase experiments due to dierences in half life and methionine content of the mutants. Equal amounts of protein lysates were analysed by Western blot, and Myc proteins visualized using an anti-myc antibody (Hann Figure 1 MC29 v-Myc is more stable than c-Myc. (a) Schematic representation of MC29 v-Myc-derived point mutations relative to c-Myc. The domain structure of Myc shows an N-terminal transactivation domain and C-terminal basic (B), helix ± loop ± helix (HLH) and leucine zipper (LZ) domains. Thr-61, the most mutated residue in tumour-derived Myc genes (Henricksson and Luscher, 1996; Hoang et al., 1995; Smith-Sorensen et al., 1996) is highlighted. Each construct was cloned into the avian retroviral expression vector, SFCV-sa7 (La Rocca et al., 1994) and transfected into chick embryo ®broblasts (CEF). (b) The turnover of avian MC29 v-Myc and c-Myc proteins in CEF was detected by Western blot of total cell lysates using an antimyc antibody following treatment with the protein synthesis inhibitor, emetine, for the indicated periods of time et al., 1983). From Figure 1b, it can clearly be seen that MC29 v-Myc is considerably more stable than cMyc, suggesting that the virally-derived point mutations stabilize the v-Myc protein. Densitometric scanning reveals that the half life of MC29 v-Myc protein is extended to 61 min, compared to 20 min for c-Myc. The Ras/Raf/MAPK pathway has been shown to result in stabilization of Myc (Sears et al., 1999). Therefore, to exclude any emetine-mediated eects on MAP kinase signalling, we showed that no alterations in MAPK activity occurred during emetine treatment using an anti-phospho-p44/p42 MAPK antibody (New England Biolabs) which detects the active p44/p42 MAPK. In addition, in combination with the use of the speci®c MAPK inhibitor, U0126 (Promega), we conclude that the stabilization observed in v-Myc was not due to emetine-induced activation of MAPK which speci®cally targeted v-Myc (data not shown). Threonine 61 regulates Myc degradation To focus on the roles of individual point mutations in MC29 v-Myc, we generated each point mutation in isolation (Figure 2a). The resulting constructs were transfected into CEF and analysed for the stability of the Myc proteins following emetine treatment. The thr-61 to met-61 mutation (c-MycT61M) is sucient to stabilize the protein to levels seen in v-Myc (Figure 2b). To discover whether the other mutations contribute to altered Myc turnover, we generated a construct, v-MycM61T, which contained, in addition to the four other virally-derived point mutations, a backmutation of met-61 to thr-61 (Figure 3a). These results clearly show that the thr-61 to met-61 mutation is sucient to stabilize the c-Myc protein, whilst the met-61 to thr-61 mutation destabilizes the v-Myc protein, even in the presence of the other viral mutations (Figure 3b). As has previously been reported (Ciechanover, 1998; Gross-Mesilaty et al., 1998; Salghetti et al., 1999), we con®rm a major involvement of the proteasome in Myc degradation in the avian system using the speci®c proteasome inhibitor, lactacystin (Fenteany et al., 1995; Gross-Mesilaty et al., 1998). Treatment with lactacystin resulted in a clear stabilization of both v-Myc and cMyc proteins (Figure 4). Use of the less speci®c proteasome inhibitor, ALLnL (Krappmann et al., 1996) also resulted in stabilization of the Myc oncoproteins (data not shown). Mutation of thr-61 does not completely prevent proteasomal degradation of v-Myc, which could be interpreted in a number of ways. Firstly, it is likely that the thr-61 to met-61 mutation aects the rate of proteasomal degradation of v-Myc resulting in reduced degradation of v-Myc, but not its complete stabilization. Alternatively, we cannot rule out a nonproteasomal mechanism of degradation, although this has not previously been reported for Myc. Proteasome function is unaected by Myc transformation Oncogenes can interfere with the activity of the ubiquitin-proteasome pathway (Nakajima et al., 1998), and it has been suggested that Myc may 7553 Stabilization of viral Myc proteins PR Gavine et al 7554 aect proteasome function (Henriksson and Luscher, 1996). Therefore, transformation by v-Myc, but not cMyc, may have a general eect on proteasome function. To address this possibility, we investigated the turnover of a well characterized proteasomedegraded protein, c-Jun (Musti et al., 1997; Treier et al., 1994). Our results show that c-Jun is unstable in both c-Myc- and v-Myc-transformed CEF (Figure 5), suggesting that transformation by v-Myc does not aect proteasome function. A dilysine motif within Myc Box I is not the ubiquitin target Proteasomal degradation of IkB results when TNF-adirected phosphorylation signals ubiquitination of an Figure 2 The single MC29 mutation, Thr-61 to Met-61, stabilizes c-Myc. (a) Point mutations of MC29 v-Myc and comparison with avian c-Myc. Each MC29 v-Myc point mutation was generated in isolation, cloned into the avian retroviral vector, SFCV-sa7 for turnover studies in CEF. (b) The eect of each MC29 point mutation on the turnover of the Myc protein was performed by Western blot analysis as outlined in Figure 1b Stabilization of viral Myc proteins PR Gavine et al 7555 Figure 5 Proteasome function is unaected by Myc transformation. The turnover of c-Jun, another proteasome-degraded nuclear oncoprotein, in v-Myc and c-Myc-transformed CEF was shown by Western blot using an anti-jun antibody B. Figure 3 Back mutation of Met-61 to Thr-61 in v-Myc is sucient to destabilize MC29 v-Myc. (a) Myc constructs were generated which contained the mutations, thr-61 to met-61 (T61M) and met-61 to thr-61 (M61T) in a c-Myc and v-Myc background respectively. (b) The eect of thr-61 to met-61 (T61M) and met-61 to thr-61 (M61T) mutation on the turnover of c-Myc and v-Myc respectively was shown by Western blot of total cell lysates as outlined in Figure 1b. Densitometric scanning shows that T61M mutation results in a 1.7 fold stabilization (half lives of c-Myc and c-Myc (T61M) are 14 and 24 min respectively). The M61T mutation results in a 2.2 fold destabilization (half lives of vMyc and v-Myc (M61T) are 47 and 20.7 min respectively) Figure 6 A dilysine motif within MBI is not the ubiquitin target in Myc. (a) Myc Box I sequence surrounding the phosphorylation sites, ser-65 and thr-61, which are subject to hierarchical phosphorylation by MAPK and GSK-3a respectively (Lutterbach and Hann, 1994). A dilysine motif, KK54,55, adjacent to these phosphorylation sites was mutated to diarginine residues, RR54,55 (Rodriguez et al., 1996) to determine the eect on Myc turnover. (b) The eect of the KK54,55RR54,55 mutation on the turnover of c-Myc was shown by Western blot of total cell lysates as outlined in Figure 1b adjacent dilysine motif (Rodriguez et al., 1996). The arrangement of a dilysine motif with respect to known phosphorylation sites in IkB is similar to that in the Nterminus of Myc (Rodriguez et al., 1996) (Figure 6a). In view of this similarity, we asked whether the two lysine residues adjacent to the known MBI phosphorylation sites, thr-61 and ser-65 (Figure 6a) (Lutterbach and Hann, 1994) are ubiquitin targets in Myc. We generated a mutant, Myc (KK54,55RR54,55,) in which the dilysine motif, KK54,55, was mutated to a diarginine motif, RR54,55 (Rodriguez et al., 1996). The half life of Myc (KK54,55RR54,55,) in CEF was determined and Figure 6b shows that these lysines are not ubiquitin targets, since this mutation does not stabilize the protein. Indeed, the protein appears more unstable than c-Myc, which was con®rmed by analysis of earlier time points (data not shown). Two independently isolated v-Myc alleles, avian MH2 and feline T17, are also stable Figure 4 A major component of Myc in chick embryo ®broblasts is degraded by the proteasome. The speci®c proteasome inhibitor, lactacystin, was added to v-Myc and cMyc-transfected CEF prior to emetine treatment to con®rm proteasome-directed degradation of Myc. Stabilized Myc proteins were detected by Western blot as described in Figure 1b To establish whether stabilization is a common feature of independently derived v-Myc alleles, we generated avian retroviral vectors encoding the v-myc sequences derived from the avian leukosis virus, MH2 (Walter et al., 1986) and feline leukaemia virus T17 (Fulton et al., Stabilization of viral Myc proteins PR Gavine et al 7556 Figure 7 Two independently isolated v-Myc isolates, avian MH2 and feline T17, are more stable than c-Myc. (a) Schematic representation of avian MH2 and feline T17 v-Myc relative to cMyc. The T17 v-Myc mutations are localized as indicated (Fulton et al., 1996), however, due to the extensive nature of the mutations in MH2 v-Myc (Walter et al., 1986), these are not accurately represented. Thr-61, the only mutation in common between MC29 v-Myc (Figure 1a) and MH2 v-Myc, is also deleted in T17 v-Myc, is highlighted. Each viral construct was cloned into the avian retroviral expression vector, SFCV-sa7 (La Rocca et al., 1994) and transfected into chick embryo ®broblasts (CEF). (b) The turnover of MH2 and T17 v-Myc proteins in CEF detected by Western blot of total cell lysates as described in Figure 1b. Chimaeras between T17 v-Myc and feline c-Myc con®rm that the T17 deletion is solely responsible for the stabilization (data not shown) 1996) (Figure 7a). In addition to extensive point mutations, MH2 has a small deletion. T17 v-Myc, however, contains an extensive 74 amino acid deletion in the transactivation domain, together with two point mutations in the C-terminus (Fulton et al., 1996). Signi®cantly, the T17 deletion spans the mutational hotspot in Myc and the highly conserved Myc Box I (MBI: amino acids 45 ± 63). Analysis of MH2 and T17 v-Myc degradation show both viral products are more stable than their cellular counterparts (Figure 7b), leading us to conclude that stabilization is a common occurrence in viral Myc isolates. Using chimaeras between T17 v-Myc and cMyc, we con®rmed that the T17 deletion was solely responsible for stabilizing the protein (data not shown). In addition, the common mutation at thr-61 (MC29 and MH2 v-Myc) or deletion spanning this domain (T17 v-Myc) suggests a common mechanistic basis for the stabilization. Discussion Three independent viral isolates of Myc (two avian and one feline) encode proteins which are considerably Figure 8 Mutations in MBI regulate Myc degradation. (a) Hierarchical phosphorylation of the Myc Box I (MBI ± amino acids 45 ± 63) amino acids, ser-65 and thr-61 by MAPK and GSK3a (Lutterbach and Hann, 1994) respectively regulate Myc degradation. MBI is depicted by the hatched area. The proteasome inhibitor, lactacystin, signi®cantly stabilizes Myc in the avian system, which is consistent with current evidence suggesting Myc is degraded through the proteasome (Ciechanover, 1998; Gross-Mesilaty et al., 1998). (b) MC29 and MH2 vMyc mutations within MBI, and speci®cally the GSK-3a phosphorylation site, thr-61, reduce Myc degradation. Thr-61 is clearly pivotal in controlling Myc degradation, but on the basis of current available data, a role for phosphorylation on ser-65 cannot be discounted. Unlike IkB degradation, ubiquitination of Myc does not occur through a dilysine motif upstream of key regulatory phosphorylation sites. (c) Deletion of MBI residues 49 ± 124 in T17 v-Myc also results in reduced degradation. This may be analogous to the d domain of c-Jun, which serves as a recognition site for the ubiquitination machinery. The d domain deletion in ASV17 v-Jun therefore abrogates ubiquitin-dependent degradation of c-Jun (Isaksson et al., 1996) more stable than c-Myc, suggesting that reduced degradation may be a common mechanism for enhancing the oncogenicity of the Myc protein. Indeed, stabilization may be a common mechanism for all nuclear transcription factor oncoproteins, since activated forms of Jun, Fos and Myb are also more stable than their cellular counterparts (Bies and Wol, 1997; Curran et al., 1994; Okazaki and Sagata, 1995; Treier et al., 1994). Posttranscriptional mechanisms which result in reduced degradation may therefore contribute to the development of the neoplastic phenotype. Loss of thr-61, a major phosphorylation site, is a key feature of three independently derived viral Myc isolates suggesting a common selective process, and implicating MBI as a major determinant in degradation of Myc (Salghetti et al., 1999). Mutation at thr-61 also potentiates Myc transformation in vitro, suggesting a correlation between stabilization and transforming function (Henriksson et al., 1993; Pulverer et al., 1994). In chick cells, the appearance of colonies in soft agar produced by Myc(T61M) more closely resembles vMyc than c-Myc (data not shown), further supporting the hypothesis that stabilization potentiates Myc transformation. However, the role of the other MC29 point mutations in regulating Myc transforming function cannot be discounted, particularly since a CKII phosphorylation site is also mutated in MC29 vMyc (ser-325 to leu-325) (Luscher et al., 1989; Walter Stabilization of viral Myc proteins PR Gavine et al et al., 1986). In the case of T17 v-Myc, the MBI deletion is responsible for stabilizing T17 v-Myc in both ®broblasts and T cells (Fulton et al., 1996), but the protein is inactive for in vitro transformation and apoptosis in ®broblasts, while retaining the potential to induce T cell lymphomas in vivo. These results suggest that Myc has pleiotropic eects, which contribute dierentially to transformation of dierent cell types. Ser-65 and thr-61 are subject to hierarchical phosphorylation by MAPK and GSK-3a respectively (Lutterbach and Hann, 1994). However, since an increase in phosphorylation at ser-65 is detected when thr-61 is mutated (Lutterbach and Hann, 1994), the regulatory mechanism for Myc stabilization could either be through loss of thr-61 phosphorylation or an increase in ser-65 phosphorylation. Myc degradation would, therefore, be regulated by the balance between these two regulatory kinases in the cell. In the absence of GSK-3a, ser-65 phosphorylation by MAPK may result in Myc stabilization (Sears et al., 1999), and subsequent thr-61 phosphorylation by GSK-3a will destabilize Myc (Figure 8a). However, as a consequence of altered thr-61 phosphorylation, viral Myc mutants such as MC29 and MH2 v-Myc would be stabilized due to reduced degradation (Figure 8b). The T17 v-Myc deletion, however, removes MBI and the N-terminal regulatory phosphorylation sites (Figure 8c). Intriguingly, this deletion is reminiscent of the d domain deletion spanning the transactivation domain in ASV17 v-Jun (Isaksson et al., 1996; Treier et al., 1994), which serves as a recognition site for the ubiquitin machinery and abrogates ubiquitin-dependent degradation of v-Jun. Therefore, by disrupting the Myc degradation signal sequence, these viral Myc mutations may interfere with the ability of the ubiquitin machinery to recognize and degrade Myc (Figure 8). Phosphorylation dierentially regulates proteasomal degradation of a number of important transcription factors. For example, phosphorylation destabilizes MyoD (Song et al., 1998), whilst Fos and Jun are stabilized (Chen et al., 1996; Musti et al., 1997; Okazaki and Sagata, 1995). Degradation of Myc, however, may be regulated by two phosphorylation events, one which stabilizes the protein (possibly ser-65 (Sears et al., 1999) or an alternative (Colman and Ostrowski, 1996) and one which destabilizes Myc (thr61) (Figure 8). The balance between signalling pathways, phosphorylation and degradation of transcription factors is clearly important for maintaining appropriate levels in the cell, and viral oncoproteins subvert this control through the acquisition of point mutations (May et al., 1998; Musti et al., 1997). Despite having a similar organization of dilysine residues and phosphorylation sites which are required for the signal-induced degradation of IkB, Myc degradation does not appear to be through a similar mechanism. In fact, when mutated, the lysine residues adjacent to MBI can appear to destabilize c-Myc even further, suggesting that the primary MBI sequence can regulate the rate of Myc degradation (Salghetti et al., 1999). MBII, a domain which is also critical for Myc transformation and plays a role in Myc degradation (Flinn et al., 1998; Stone et al., 1987) contains two lysine residues which may be the potential ubiquitin targets. Alternatively, ubiquitination may involve a novel target residue (Breitschopf et al., 1998), and we are currently determining the key ubiquitin targets in Myc. In summary, we have described three independently derived v-Myc alleles which are considerably more stable than their cellular counterparts, suggesting stabilization may be a common consequence of mutations in Myc. We have discussed possible mechanisms of how Myc degradation may be regulated through mutation of either key phosphorylation sites or ubiquitin regulating sequences in view of current evidence suggesting that Myc is degraded through the proteasome (Ciechanover, 1998; GrossMesilaty et al., 1998). Experiments are therefore currently underway to determine the precise biological and biochemical consequence of these virally derived mutations of Myc. Materials and methods Cell culture and transfections Cell culture and transfection of appropriate SFCV ± Myc constructs (10 mg) together with RCAN (A) helper (4 mg) into secondary chick embryo ®broblasts (CEF) was performed essentially as described (La Rocca et al., 1994). Following G418 selection, cultures were expanded and used to analyse Myc turnover. Construction of retroviruses The construction of SFCV ± c-Myc, SFCV ± v-Myc and SFCV ± T17 v-Myc have previously been described (Fulton et al., 1996; La Rocca et al., 1994). With the exception of SFCV ± c-Myc(T61M), SFCV ± v-Myc(M61T) and SFCV ± cMyc(KK54,54RR54,55), all Myc mutants were generated by site directed mutagenesis essentially as described (Crouch et al., 1990; La Rocca et al., 1994). c-Myc(T61M) was generated by replacing an 80 bp SacI ± PvuII fragment into pAT ± c-Myc (Crouch et al., 1990) and subsequent cloning into SFCV-sa7 as previously described (Crouch et al., 1990; La Rocca et al., 1994). To generate SFCV ± v-Myc(M61T), the 3' ClaI fragment of c-Myc was replaced with a 607 bp ClaI fragment from v-Myc. c-Myc(KK54,54RR54,55) was generated by ligation of a PstI ± SacI linker containing the appropriate mutation into spt ± c-Myc (Crouch et al., 1993), and the subsequent HindIII fragment ligated into SFCV-sa7 (La Rocca et al., 1994). For the construction of SFCV ± MH2 vMyc, a 285 bp amino terminal fragment and a carboxy terminal 621 bp fragment were generated by PCR from a pMH2 proviral cDNA template (Coll et al., 1983). The primers pairs used were 5'-gacaagcttatcgataccatgccgctcagcgccagc-3' and 5'-cgtcaccatctccagctggtcggcgg-3' for the amino terminal fragment, and 5'-gacaagcttctatgcacgagagttcttt-3' and 5'-gcggctctgctgggggtcgacacgccgccc-3' for the carboxy terminal fragment. After HindIII/PstI digestion and HindIII/ClaI digestion of the amino terminal and carboxy terminal fragments respectively, the puri®ed fragments were ligated into the HindIII site of SFCV-sa+, together with a central 584 bp PstI/ClaI spanning exon 2/exon 3 boundary, which was isolated from pMH2 (Coll et al., 1983). All sequences generated by PCR or mutagenesis were con®rmed by doublestranded sequencing using T7-Sequenase v2.0 essentially as described by the manufacturers (Amersham). Oligonucleotide site directed mutagenesis Site directed mutagenesis was performed using the Sculptor mutagenesis system (Amersham) as described previously (Crouch et al., 1990). The mutagenic oligonucleotides used 7557 Stabilization of viral Myc proteins PR Gavine et al 7558 were as follows: S325L: 5'-ccccgcacgttagact-3'; S350R: 5'ctgaagctgcgtttctttgcc-3', I383L: 5'-gttctgtctctccaatcggacgag-3'; K407N: 5'-ttgaaacacaaccttgaggagc-3'. Western blotting Cell lysates were prepared by lysing cultures in SDS-sample buer containing 1% SDS, but no bromophenol blue or mercaptoethanol. Protein concentrations were measured using Micro BCA reagent (Pierce), and readjusted for bromophenol blue and mercaptoethanol before loading onto 7.5% SDS ± PAGE gels. Transfer to nitrocellulose and Western blotting was performed essentially as described (La Rocca et al., 1994), except that incubation with the primary antibody was performed in 2% BSA in TBST. Proteins were detected using speci®c rabbit antibodies (anti-v-myc 12C antibody (Hann et al., 1983) and anti-jun 730 (Kilbey et al., 1996) and visualized using enhanced chemiluminescence. Protein turnover analysis Myc turnover was analysed in G418-selected cultures following treatment with the protein synthesis inhibitor, emetine (Sigma) at a ®nal concentration of 0.1 mM (Crouch et al., 1990) for the indicated times. Proteasomal involvement was determined using the inhibitor lactacystin (Aniti Research Products) at a concentration of 10 mM (GrossMesilaty et al., 1998). The inhibitor was added 45 min prior to the addition of emetine, and was present throughout the experiment. Acknowledgements We wish to thank Jen Blake for technical assistance, RN Eisenman and DAF Gillespie for providing the anti-v-myc 12C and anti-jun antibodies respectively, and CNA Palmer and DW Meek for critical reading of the manuscript. References Bies J and Wol L. (1997). Oncogene, 14, 203 ± 212. Breitschopf K, Bengal E, Ziv T, Admon A and Ciechanover A. (1998). EMBO J., 17, 5964 ± 5973. Chen R-H, Juo PC-H, Curran T and Blenis J. (1996). Oncogene, 12, 1493 ± 1502. Ciechanover A. (1998). EMBO J., 17, 7151 ± 7160. Coll J, Righi M, de Taisne C, Dissou C, Gegonne A and Stehelin D. (1983). EMBO J., 12, 2189 ± 2194. Colman MS and Ostrowski MC. (1996). Nucleic Acids Res., 24, 1971 ± 1978. Crouch DH, Fisher F, Clark W, Jayaraman PS, Goding CR and Gillespie DAF. (1993). Oncogene, 8, 1849 ± 1855. Crouch DH, Lang C and Gillespie DAF. (1990). Oncogene, 5, 683 ± 689. Curran T, Miller AD, Zokas L and Verma IM. (1984). Cell, 36, 259 ± 268. Fenteany G, Standaert RF, Lane WS, Choi S, Corey EJ and Schreiber SL. (1995). Science, 268, 726 ± 731. Flinn EM, Busch CMC and Wright APH. (1998). Mol. Cell. Biol., 18, 5961 ± 5969. Fulton R, Gallagher R, Crouch D and Neil JC. (1996). J. Virol., 170, 1154 ± 1162. Gillespie DAF. (1991). Seminars Virol., 2, 329 ± 339. Gross-Mesilaty S, Reinstein E, Bercovich B, Tobias KE, Schwartz AL, Kahana C and Ciechanover A. (1998). Proc. Natl. Acad. Sci. USA, 95, 8058 ± 8063. Hann SR, Abrams HD, Rohrschneider LR and Eisenman RN. (1983). Cell, 34, 789 ± 798. Henriksson M, Bakardjiev A, Klein G and Luscher B. (1993). Oncogene, 8, 3199 ± 3209. Henriksson M and Luscher B. (1996). Adv. Cancer Res., 68, 109 ± 182. Hoang AT, Lutterbach B, Lewis BC, Yano T, Chou T-Y, Barrett JF, Raeld M, Hann SR and Dang CV. (1995). Mol. Cell. Biol., 15, 4031 ± 4042. Isaksson A, Musti AM and Bohmann D. (1996). Biochim. Biophys. Acta, 1288, F21 ± F29. Kilbey A, Black EJ, Unlu M and Gillespie DAF. (1996). Oncogene, 12, 2409 ± 2418. Krappmann D, Wulczyn FG and Scheidereit C. (1996). EMBO J., 15, 6716 ± 6726. La Rocca SA, Crouch DH and Gillespie DAF. (1994). Oncogene, 9, 3499 ± 3508. Luscher B, Kuenzel EA, Krebs EG and Eisenman RN. (1989). EMBO J., 8, 1111 ± 1119. Lutterbach B and Hann SR. (1994). Mol. Cell. Biol., 14, 5510 ± 5522. May GHW, Funk M, Black EJ, Clark W, Hussain S, Woodgett JR and Gillespie DAF. (1998). Curr. Biol., 8, 117 ± 120. Musti AM, Treier M and Bohmann D. (1997). Science, 275, 400 ± 402. Nakajima T, Morita K, Tsunoda H, Imajoh-Ohmi S, Tanaka H, Yasuda H and Oda K. (1998). J. Biol. Chem., 273, 20036 ± 20045. Okazaki K and Sagata N. (1995). EMBO J., 14, 5048 ± 5059. Pulverer BJ, Fisher C, Vousden K, Littlewood T, Evan G and Woodgett JR. (1994). Oncogene, 9, 59 ± 70. Rodriguez MS, Wright J, Thompson J, Thomas D, Baleux F, Virelizier J-L, Hay RT and Arenzana-Seisdedos F. (1996). Oncogene, 12, 2425 ± 2435. Salghetti SE, Kim SY and Tansey WP. (1999). EMBO J., 18, 717 ± 726. Sears R, Leone G, DeGregori J and Nevins JR. (1999). Molecular Cell, 3, 169 ± 179. Smith-Sorensen B, Hijmans EM, Beijersbergen RL and Bernards R. (1996). J. Biol. Chem., 271, 5513 ± 5518. Song A, Wang Q, Goebl MG and Harrington MA. (1998). Mol. Cell. Biol., 18, 4994 ± 4999. Stone J, de Lange T, Ramsay G, Jakobovits E, Bishop JM, Varmus H and Lee W. (1987). Mol. Cell. Biol., 7, 1697 ± 1709. Treier M, Staszewski LM and Bohmann D. (1994). Cell, 78, 787 ± 798. Walter N, Jansen W, Trachmann C and Bister K. (1986). Virology, 154, 219 ± 223.