* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download unique features of the plant life cycle and their consequences
Transposable element wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene expression programming wikipedia , lookup
Genetically modified crops wikipedia , lookup
Oncogenomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genetically modified organism containment and escape wikipedia , lookup
Genetic engineering wikipedia , lookup
Point mutation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Minimal genome wikipedia , lookup
X-inactivation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Genome (book) wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genome evolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome editing wikipedia , lookup
History of genetic engineering wikipedia , lookup
Microevolution wikipedia , lookup
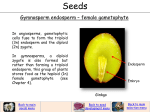
REVIEWS UNIQUE FEATURES OF THE PLANT LIFE CYCLE AND THEIR CONSEQUENCES Virginia Walbot* and Matthew M. S. Evans‡ Continuous development, the absence of a germline, flexible and reversible cellular differentiation, and the existence of haploid and diploid generations — both of which express genes — are characteristics that distinguish plants from animals. Because these differences alter the impact of mutations, animals and plants experience varied selection pressures. Despite different life-cycles, both flowering plants and multicellular animals have evolved complex sensing mechanisms that act after fertilization as ‘quality checks’ on reproduction, and that detect chromosome dosage and the parent of origin for specific genes. Although flowering plant embryos escape such surveillance in vitro, embryo success in the seed often depends on a healthy endosperm — a nutritive tissue that is produced by a second fertilization event in which maternal and paternal gene contributions can be monitored immediately after fertilization and throughout development. DOUBLE FERTILIZATION The process by which two cells in a megagametophyte fuse with two sperm (typically from the same pollen grain) to produce both a diploid embryo and an accessory organ — the endosperm. Double fertilization is characteristic of angiosperms, but also occurs in other taxa in which the result is usually the production of two embryos. *Department of Biological Sciences, Stanford University, Stanford, California 94305-5020, USA. ‡ Department of Plant Biology, Carnegie Institution of Washington, 260 Panama Street, Stanford, California 94305, USA. e-mail: [email protected]; mmevans@andrew2. stanford.edu doi:10.1038/nrg1064 The complete genome sequences of complex animals and plants indicate that a similar number of genes is required for a fruitfly, human and rice plant1. Furthermore, many gene classes are conserved although it is >1 billion years since the last shared progenitor of plants and animals1. There are, however, fundamental differences in the life cycles of plants and animals (FIG. 1) that alter both the timing and the stringency of natural selection in these kingdoms. The presence of genetically active haploid (gametophyte) and diploid (sporophyte) phases of the life cycle, the absence of a germline, and a DOUBLE FERTILIZATION event during sexual reproduction to produce both an embryo and a nutritive tissue (ENDOSPERM) are unique features of flowering plants. These plant-specific life-history components alter the impact of mutations and present opportunities for distinctive developmental regulation. Here, we illustrate these features of plant development using examples primarily from Arabidopsis and maize that we compare with animals, and comment on the resulting genetic and evolutionary consequences of the plant life cycle. In plants, the gametophytic stage presents an opportunity for natural selection on the haploid genome — as NATURE REVIEWS | GENETICS in other haploid organisms, recessive deleterious alleles that would be masked in diploid organisms are removed from the population2. Over evolutionary history, there has been a steady reduction in the diversity of structures during the haploid phase; for example, gametophytes in flowering plants are non-photosynthetic and contain few cell types. Nonetheless, on the basis of RNA hybridization kinetics, 20,000–25,000 genes are expressed in the pollen of maize and Tradescantia3,4; this represents roughly onethird to one-half of the probable total gene number5. We propose that supplementary mechanisms have also evolved to bolster the ‘quality check’ of the genome. These include imprinting — the silencing of either paternally or maternally derived alleles — as well as checks on the ratio of maternal to paternal endosperm ploidy and chromosome-arm dosage. In maize and Arabidopsis, these quality checks ensure that sexual reproduction is tightly regulated, in contrast to the innate flexibility of plant development. These quality-checking mechanisms could be seen to parallel the sensitivity of mammalian development to perturbations in chromosome dosage, and the requirement for both a sperm and an egg. Also, imprinting extends the haploid sufficiency VOLUME 4 | MAY 2003 | 3 6 9 © 2003 Nature Publishing Group REVIEWS a b Tassel Zebrafish Ear Zebrafish Mature sporophyte (2n) Meiosis Egg (n) Megaspore (n) Microspore (n) Sperm (n) Mitosis Mitosis Megagametophyte (n) Egg cell Polar nuclei Microgametophyte (n) (germinated pollen grain) Vegatative nucleus Sperm nuclei Fertilization Endosperm (3n) Primordial germ cells Embryo (2n) Figure 1 | Comparison of plant and animal life cycles. a | Animal life cycle, in which gametes are produced directly through meiosis, and gene expression is restricted to diploid cells. In the embryo, the germline is set aside early (primordial germ cells) and these are the only cells that are competent to undergo meiosis in the adult. b | Typical flowering plant life cycle (maize), in which meiosis, followed by mitotic divisions, produces two types of haploid organism that are genetically active — the female megagametophyte and the male microgametophyte (pollen grain). Mitotic division in the microgametophyte results in a pair of sister cells that differentiate into sperm. In the megagametophyte, a haploid egg differentiates and is fertilized by one sperm cell to produce an embryo, and two haploid nuclei fuse to form a diploid central cell that is fertilized by another sperm cell to become triploid endosperm. The endosperm is a terminal nutritive tissue that contributes to normal embryo development in flowering plants (note that double fertilization has been documented in some non-flowering plants86, and endosperm ploidy in flowering plants can be diploid or other ploidy values that are distinct from triploid87). Red arrows show plantspecific features. Photograph provided by E. Raz; figure reproduced with permission from REF. 88 (2002) Company of Biologists Ltd. ENDOSPERM A tissue, found in flowering plants, which is generated by the fusion of the central cell of the megagametophyte and a sperm. In most angiosperms, the endosperm is triploid, with two genome equivalents from the maternal line and one from the paternal line; however, there are many exceptions to this general rule. AUTOTROPHIC Able to independently acquire a nutrient. INFLORESCENCE The reproductive portion of a plant that bears a cluster of flowers in a specific pattern. 370 test transiently in the early stages of the diploid phase of the life cycle. Similarly, in mammalian embryos, imprinting creates effective haploid selection, but only for a small number of loci early in embryo development6. Selection on haploid cells Plants have two phases to their life cycle: the diploid sporophytic stage that ends in meiosis to produce haploid cells, and the haploid gametophytic phase in which mitotic proliferation produces a haploid plant that includes the differentiation of a subset of cells as gametes. Both the egg-producing haploid plant (the megagametophyte) and the sperm-producing haploid plant (the microgametophyte) are genetically active —therefore, the phenotype reflects the haploid genotype. It is often argued that haploid individuals are at a selective disadvantage compared to diploid individuals as deleterious recessive mutations are exposed7,8, but an advantage is that haploidy is an opportunity to purge the population of deleterious mutations9,10. As a | MAY 2003 | VOLUME 4 consequence, plants probably have a lower genetic load than animals, the gamete properties of which are determined almost entirely by gene expression in progenitor diploid cells. Recessive lethal and deleterious alleles can accumulate in diploid animal populations, because they are masked in the heterozygous diploid individuals10. Despite its possible advantages, natural selection has favoured the reduction of the haploid phase in plants in three ways: the proportion of the total life cycle, physical size and the range of biological processes. Although there is great diversity in algal life cycles11, in the green alga Chlamydomonas reinhardtii, which is a model plant for genetic analysis now that the nuclear genome is sequenced12, most of the life cycle is haploid — gametes fuse to produce a diploid cell, which can immediately undergo meiosis to reconstitute haploid cells that proliferate by mitosis. The haploid phase expresses most of the genome, whereas the diploid phase is centred on meiosis13. In the mosses and liverworts, which are basal green plants, the haploid phase is also dominant. Individual gametophytes are free-living, AUTOTROPHIC and sustain the diploid sporophytes, which remain embedded in the gametophytes11. In flowering plants, this state is completely reversed as the haploid phase usually consists of 2–7 cells that depend on the diploid parent for nutrient acquisition11. Therefore, given the reduction in size and metabolic contribution in the most recently evolved plant taxa, has the scope or intensity of the haploid sufficiency test diminished? Despite the limited size and lifespan of pollen, RNA transcripts that correspond to many genes are present, and, in some cases, there is proof that transcription occurred post-meiotically from the haploid genome3,4. Many genes are required to produce functional pollen, as is clear from random mutagenesis experiments. In Arabidopsis, ~1.3% of mutated lines are gametophytic lethal, which indicates that at least 200 genes are vital for gametophyte development14. This is probably an underestimate, as seed is not recovered if either gametophyte fails before competent gamete formation in the self-fertilizing flowers of Arabidopsis 15,16. In maize, the microgametophytes and megagametophytes develop in separate INFLORESCENCES — the tassel and ear, respectively17. After Mu transposon mutagenesis of maize, complete male sterility (100% defective pollen, which indicates a sporophytic failure) and semi-steriles (50% pollen death presumably as a result of the segregation of defective genes required in the microgametophyte) are among the most common phenotypic classes18. There is also overlap in the genes that contribute to gametophytic and saprophytic success. For a few metabolic characteristics, including alcohol dehydrogenase activity and starch composition3,18, there is clear evidence of selection that occurs on the gametophyte for sporophytic traits. This could be widespread for many developmental regulatory genes. Patterson and others exploited >850 reciprocal translocation stocks, representing 1,700 deficiencies in maize, and established that they all caused pollen abortion and that ~90% resulted in megagametophyte www.nature.com/reviews/genetics © 2003 Nature Publishing Group REVIEWS Box 1 | Identification of loci essential for gametophyte development The identification of loci that are required for the gametophyte phase and gametogenesis requires special tools, because natural and induced mutants that are gametophytic lethal are lost in standard mutagenesis approaches, even if they are viable in heterozygotes. As gametophytes inherit either the functional or lethal allele, a semisterility phenotype is observed, in which 50% of pollen or eggs are viable. If knockouts result in gametopytic lethality, no diploid organism is generated and, therefore, it is impossible to score the effects of mutations in diploid tissues. Similarly, genes that cause lethality early in embryo development must be rescued to uncover their functions at a later stage in life. To do this, transgenes that are expressed selectively at the earlier stage (haploid or embryo) allow development and, therefore, allow the evaluation of phenotypes at later stages. A recent example of this strategy showed that the FERTILIZIATION INDEPENDENT ENDOSPERM (FIE) locus of Arabidopsis is responsible not only for suppressing precocious endosperm development but also for suppressing inappropriately early flowering81. SOMA The cells of the body that cannot undergo meiosis. In plants, this comprises the entire plant body until the late specification of reproductive cells in flowers. By contrast, animals have a somatic body and a germline that differentiates early in development; at reproductive maturity, the germ cells proliferate, undergo meiosis and the meiotic products differentiate into gametes. lethality19. Presumably, some mutations that cause nutritional defects that are lethal to pollen, which is sealed from metabolic exchange with the surrounding diploid tissues for days, can be compensated for in megagametophytes, which continue to absorb nutrients from the vegetative plant. In BOX 1, we describe how genes that are involved in gametophyte development can be studied. Gametophytic lethality might be considered surprising, given the extent of gene duplication in flowering plant genomes. Autotetraploidy (chromosome doubling in a species) and allotetraploidy (chromosome doubling after a cross between two species) are common events in all taxa. Approximately 70% of the Arabidopsis genome consists of an ancient (>100 million years (Myr) ago) duplication20,21. Furthermore, ~17% of Arabidopsis loci have a linked recent duplication22. In maize, the modern genome is the result of a complete duplication in the past 11–16 Myr23. At the time of genome-wide duplications, gametophytic functions must be redundantly represented by a minimum of two loci, yet today maize is effectively a diploid species for many loci because mutations at one locus in a gene family produce a visible phenotype18. Such observations are, at present, discussed in the context of the subfunctionalization model24, in which the main mode of divergence of duplicate genes involves the loss of portions of the gene-expression programme at individual loci (or more precisely, their constituent alleles). As a result, both loci are required during the life cycle, because at particular stages only one locus is expressed. For animals, subfunctionalization is invoked to explain selection that favours the retention of duplicate genes that act in different cell types; in plants, subfunctionalization could also have acted to distinguish loci that are essential for the haploid phase from those that are now expressed in the diploid phase. Testing this idea requires locus-specific methods to distinguish between the expression patterns of closely related genes. Plants produce meiotic cells late in development A crucial distinction between plants and animals concerns the germline. In animals, germ cells are designated early in embryo development and remain as a separate stem-cell population throughout the life of the animal25; the germ-cell derivatives are the only cells that NATURE REVIEWS | GENETICS can undergo meiosis to produce eggs and sperm. Germcell specification occurs by several mechanisms, including the differential distribution of material stored in the egg (in worms and flies) and fate decisions at the time of gastrulation (in mammals)26,27. By contrast, the stemcell population that directs the growth of the plant shoot only produces somatic structures — leaves and shoots — until environmental and developmental signals trigger the switch to reproduction28,29; that is, the apical stem cells switch from producing strictly somatic organs to producing flowers, in which a few cells are programmed to undergo meiosis to produce megagametophytes, but many more cells are programmed to differentiate into microgametophytes11. There is high regularity in the anatomical placement of pre-meiotic cells in most flowers, but this does not reflect a stringent cell-lineage relationship — plant development is generally governed by positional information (for example, cell–cell interactions) that results in a high regularity of structure28. In plants, we are beginning to define the genes that are crucial for the cell-fate decision that, at a particular mitosis, restricts one cell to the SOMA and the sister cell to the pre-meiotic fate. Among the handful of defined genes that alter cell-fate determination in the anthers of Arabidopsis are two leucine-rich repeat receptor kinases30,31 that, if mutated, result in an excess of meiotic cells and a deficiency of somatic cells. A nuclear protein of unknown role is required for meiotic-cell specification32. Several other loci that alter anther cell fate have been defined in Arabidopsis but are not yet cloned33, and, in maize, the multiple archesporial cells 1 (mac1) mutation disrupts the somatic-to-germinal switch in both male and female flowers34, which results in an excess of meiotic cells. Also, genes that are expressed in the megagametophyte are important during post-meiotic and post-fertilization development35 — a theme that we return to in the discussion of imprinting. The failure to set aside a germline provides a potential problem for plants, because gametes could carry many mutations that have accumulated during somatic growth. This would create an incipient disaster for animals, given the absence of haploid selection. Somatic diversification occurs in plants, as is readily observed in mangroves in which the developing seeds germinate while they are attached to the maternal plant; this allows observation of the segregation of albino mutants on some branches but not others36. Similarly, plants that carry a transposable element inserted in an anthocyanin pigment gene can have red sectors that represent somatic reversion; if red sectors occur in flowers, a revertant red allele can be transmitted to the next generation37. Stringent selection during the haploid gametophytic phase and selection during somatic growth — as cell lineages carrying deleterious mutations are impaired in growth and development and so are less likely to contribute to gamete formation — might abrogate the problem of accumulated mutations before meiosis38. As reviewed in BOX 2, it has been argued that these features make plants more tolerant of somatically active transposable elements39. VOLUME 4 | MAY 2003 | 3 7 1 © 2003 Nature Publishing Group REVIEWS APOMIXIS The production of seed without embryo fertilization, which can involve direct embryogenesis from somatic cells or the development of meiotic products into embryos. GYNOGENETIC An individual that develops from a cell in the megagametophyte (typically the egg) and, therefore, contains only maternal chromosomes. ANDROGENETIC An individual that develops from a sperm and, therefore, contains only paternal chromosomes. Plasticity of plant embryogenesis Formation of a mammalian embryo requires either the fertilization of an egg by a sperm or, nowadays, the implantation of a somatic nucleus in the cytoplasm of an egg40. In hundreds of insect41 and a few vertebrate42 species, sexual reproduction is circumvented by the direct development of embryos from eggs. In plants, reproduction is innately more flexible. There are many examples of asexual plant reproduction, ranging from the production of miniature plantlets on leaf margins, as seen in Kalanchoë (FIG. 2), to APOMIXIS in which seeds can develop in flowers without egg fertilization. Apomixis occurs in diverse taxa43, and seeds can be produced if either somatic cells or meiotic products undergo embryogenesis43,44. Further developmental plasticity is observed in vitro, as diploid somatic cells45 and haploid pollen grains46 can form embryos in tissue culture. Although haploid plants are sterile, chromosome doubling can produce fertile homozygous plants. Therefore, a haploid genome of paternal origin is fully competent to organize an embryo in tissue culture, and an endosperm is dispensable for development outside the context of a seed. The spontaneous production of haploid embryos in situ is line dependent in maize47,48, but the frequency is generally 10–3 for GYNOGENETIC origin and 10–5 for 49 ANDROGENETIC origin . Therefore, neither the maternal nor the paternal genome is absolutely required for maize embryo development in a flower. Further insights into the in semine (in seed) requirements come from the analysis of an important regulator of gametophyte fate — indeterminate gametophyte (ig). This maize gene seems to suppress the capacity of haploid cells to form an embryo and to help maintain the proper ploidy of megagametophyte cells. In loss-offunction ig mutants, a percentage of the progeny are haploid, and androgenetic haploids are as frequent as the gynogenetic derivatives49. Curiously, in all cases, the ig haploid plants have a normal triploid endosperm, which indicates that endosperm fertilization has occurred. Therefore, in maize, a haploid embryo probably needs a normal triploid endosperm to survive. Interestingly, even in plants that produce seed by apomixis, most taxa produce an endosperm by fertilization of the central cell by a sperm cell50. However, endosperm is probably not required in all flowering plants, with an extreme case being orchids, in which a second fertilization usually occurs but the triploid endosperm nucleus quickly degenerates51. Non-equivalence of parental genomes In most angiosperms, both an embryo and its endosperm are part of a developing seed — the unit of reproductive dispersal (FIG. 3). The functional relationship of the endosperm to the embryo parallels that of the placenta to the fetus in mammals, because the endosperm nourishes the embryo. Several lines of evidence show that seed production requires contributions from both parents, which indicates that the maternal and paternal genomes are not equivalent. Parent-of-origin effects can be detected by varying the relative germplasm contributions from the two parents and by examining mutations Box 2 | The plant life cycle establishes conditions for tolerating transposons The combination of haploid selection and the production of gametes from diverse somatic lineages might allow plants to harbour active transposons with few consequences. Obviously, human selection for transposon activity at colour genes resulted in the diversity of striped and dotted maize kernels, and of flecked horticultural flower varieties. But transposon excision from non-vital colour genes should be accompanied by insertions elsewhere. Insertions in essential genes should result either in selection against transposon activity or the loss of viability in the line, because many new insertions disrupt gene function. Newly generated insertion alleles contribute to the somatic mutations in the plant, but can only accumulate in the population if they survive haploid selection. The combination of purifying selection of the haploid phase and the diversity of somatic lineages that can generate flowers, buffers the plant from transposon excess. New dominant-lethal mutations in presumptive germ cells do not eliminate reproduction elsewhere, and lethality in the gametes removes deleterious alleles from the population. The more permissive conditions for ‘genome experimentation’ in plants might help explain the enormous range of genome sizes, even among closely related taxa. Genome composition analysis shows that, in many cases, genome size increases reflect the recent amplification of transposable elements82,83. In some cases, transposon behaviour is directly tied to important features of the plant life cycle. Mutator activity in maize results from the mobility of MuDR/Mu elements, which collectively can increase forward mutation frequency 50–100-fold84. These highly active multicopy transposons selectively insert in maize genes, but do not kill the host. The MuDR/Mu transposon family is mobile only late in the life cycle during the last few somatic cell divisions; this minimizes the impact on somatic growth, because only a few cells will carry any newly arising insertion mutant. Furthermore, each transposon excision and insertion generates two chromosome breaks, but this genotoxic stress does not impact the stem-cell populations or organs at early stages of their development. By contrast, the Ac/Ds and Spm element families studied by Barbara McClintock are active at all somatic stages, but as they are present in only one or a few copies, their activities cause only a modest disruption compared with what might happen if 50–100 Mu elements were mobile throughout the life cycle84. Because of the late onset of all MuDR/Mu activities, heritable Mu insertions occur in the pre-germinal cells before meiosis as well as in gametophytes, with 20% of the events occurring after the last mitotic division that separates the two sperm85. Consequently, Mu transposons amplify their copy number and create many new insertional mutants coincident with the stringent check of genome quality that occurs in the haploid phase. 372 | MAY 2003 | VOLUME 4 www.nature.com/reviews/genetics © 2003 Nature Publishing Group REVIEWS Figure 2 | Seedlings produced along the Kalanchoë leaf margin. This is an example of the flexibility in plant reproduction, as plantlets form along the leaf margin. Reproduced with permission from Curtis Clark © (2003) Curtis Clark, Biological Sciences, California State Polytechnic University, Pomona, USA. that cause aberrant development if inherited through one parent but not the other. There are several possible mechanisms for parent-specific mutant phenotypes; these include imprinting, a requirement for a particular dose of active gene copies in the endosperm and the cytoplasmic inheritance of gene products from one of the haploid gametophytes. a Maize seed b Arabidopsis seed Endosperm Embryo Embryo 1 mm 60 µm Figure 3 | Relative contribution of embryo and endosperm to mature seeds. a | A median longitudinal section of a mature maize seed, stained with Evans blue to show the programmed cell death of the starchy endosperm. The bulk of the seed consists of the endosperm rather than the embryo. b | By contrast, in the median longitudinal section of a mature Arabidopsis seed, the embryo almost completely fills the seed coat and little endosperm persists. Reproduced with permission from J. A. Long and M. K. Barton, Carnegie Institution, Stanford, California, USA. NATURE REVIEWS | GENETICS Although all flowering plants must produce a successful embryo, species differ substantially in the relative importance of the endosperm (FIG. 3); they range from having no endosperm, as in orchids52, to species with a transitory organ that supports embryo development, and to cases in which the endosperm stores the nutritional reserves for seedling establishment. In Arabidopsis, the endosperm is crushed by embryo development and its nutrients are transferred to the embryo. In cereals, such as maize, the endosperm can be 90% of the seed weight, extending the reliance of the embryo on the endosperm to seedling germination. Given the variability of the reliance of the embryo on the endosperm, we might expect variability in the extent of parent-of-origin effects on seed development. What is the function of parent-of-origin effects? We have already introduced the concept that the extension of functional haploidy after fertilization could serve as a further check on genome quality. Consistent with this, evidence now points to functional haploidy for maternal alleles acting early in endosperm and embryo development, and this could effectively extend haploid selection53. Also, parent-of-origin effects act as a quality check on sexual reproduction, because seed is only produced if the fertilization of appropriately programmed male and female gametes occurs. An additional explanation for parent-of-origin effects originates from the parental conflict theory, in which maternal and paternal alleles are differentially expressed after fertilization to modulate resource allocation to the progeny52. The main prediction is that there should be specific loci in which either the maternal or the paternal allele is expressed after fertilization. The differences in parental contribution to seed success are substantial: the paternal parent provides little beyond chromosomes, whereas the maternal parent supplies the nutrients for embryo and endosperm development, and maternal tissues form the seed coat and the surrounding fruit tissues. The parental conflict theory proposes that paternal alleles evolve to maximize nutrient allocation to the individual seed (the selfish paternal parent), whereas the maternal parent must support a suite of seeds that develop in a fruit; so, maternal alleles are probably under selection to reduce resource allocation and avoid a negative impact on the overall reproductive potential of the mother. The genetic basis for the conflict between the parents is that the maternal plant is equally related to each embryo and endosperm, and, therefore, has a similar investment in each seed. By contrast, if there are different paternal parents, individual seeds are not full siblings and so are in competition for maternal resources52. The substantial evidence that specific chromosome arms of paternal origin are essential for maize endosperm development — which implies either that the dosage of some genes is crucial, or that paternal alleles are expressed whereas the corresponding maternal alleles are less effective — fits with the parental conflict theory. Paternal deficiency for 8 of the 19 maize chromosome arms that have been tested using B-A chromosome translocations (FIG. 4), resulted in a severe reduction in VOLUME 4 | MAY 2003 | 3 7 3 © 2003 Nature Publishing Group REVIEWS Mitosis I Mitosis II A B A B B A B A B A Sperm cells Generative cell A B B A A B Vegetative nucleus Microspore nucleus A B A B B A B A Figure 4 | Microgametophyte development and B-A chromosome behaviour. The B chromosomes in maize are peculiar; they carry no essential genes but can undergo translocation with the standard maize A chromosomes. In this example, the meiotic product inherited one copy of the A-B and B-A chromosomes that had participated in an exact reciprocal exchange at the centromere. The first mitotic division (mitosis I), produces a vegetative cell and a smaller generative cell, which have the same genetic constitution. The generative cell then divides (mitosis II) to produce two sperm, each of which participate in fertilization. B-A chromosome nondisjunction during this mitotic division routinely produces one sperm with an A-arm deficiency, whereas the second sperm has a duplication for this arm. This enables the ploidy of A-chromosome arms to be manipulated. The vegetative cell — which supplies most or all of the metabolic functions — has a balanced chromosome set, and, therefore, gametophyte viability is generally unaffected by these translocations. B-A chromosome translocations in maize have been used to investigate the effect of paternal deficiency of different chromosome arms54,55, and to mark different sperm nuclei to assess their fate65. DICOT A flowering plant with two embryonic initial leaves, known as cotyledons. MONOCOT A flowering plant with a single cotyledon in the embryo. 374 endosperm size54,55. Increasing the number of maternal copies did not overcome these defects and could exacerbate them56. Collectively, the data from Arabidopsis and maize support the parental conflict model, in which paternal alleles control the growth of the endosperm because their expression is required or is dominant to the maternal alleles. The function of stringent ploidy ratio requirements for endosperm formation could be linked to the parental conflict theory, and could also provide a quality check to ensure that viable seed is formed. FIGURE 1 shows that flowering plant gametophytes have two cells that can fuse with a partner of the opposite sex, resulting in a diploid embryo and a typically triploid endosperm. One hypothesis, which results from the parental conflict theory, is that if the ploidy of the endosperm is altered, the competition between the maternal and paternal genomes is disrupted, which results in aberrant development. These predictions have been tested using ig1 maize mutants, which produce megagametophyte central cells with normal diploidy, or in many cases higher ploidy, which shows that the endosperm that is produced after fertilization has a range of ploidy values. Endosperm with a ratio of maternal to paternal genomes of 2:1 (2m:1p) are normal, whereas 3m:1p endosperms are viable but abnormal. Higher maternal ploidy (tetraploid to octoploid) is lethal to the endosperm and results in maize kernel abortion57. However, it is the ratio of maternal:paternal genomes that is crucial for endosperm development, not ploidy per se, as both 4m:2p and 2m:1p ratios promote normal endosperm development57. Similar results are seen in interploidy crosses in other DICOT and MONOCOT plants, including wheat, barley and | MAY 2003 | VOLUME 4 species of Solanum and Primula 50,58. In Arabidopsis, there is a greater tolerance for deviations in the maternal to paternal genome ploidy ratio59, but normal development only occurs in crosses producing endosperm with a 2m:1p ratio. Furthermore, the phenotype of the endosperm in these interploidy crosses fits the predictions of the parental conflict theory — in which the relative contribution of the maternal and paternal parent must be balanced to allow normal development — for many of the species tested50,58. In these interploidy crosses, both the endosperm and embryo ploidy ratios are altered and, therefore, these experiments do not resolve whether the endosperm or the embryo is the crucial tissue for sensing parental contributions, although the phenotype is seen in the endosperm. Only in maize has it been shown that the endosperm ploidy ratios, separate from those of the embryo, are crucial for normal development57. Parent-of-origin effects of individual loci Finer resolution of the regulation of parent-of-origin effects is possible if the focus switches to individual genes. Imprinting breaks one of Mendel’s laws — namely that alleles are unchanged by their mode of transmission. For imprinted alleles, the parent of origin determines expression in the next generation. The first documented case of gene imprinting 60 was the maize R regulatory gene, which encodes a basic helix–loop–helix transcription factor and controls the pigmentation of the endosperm epidermis61. Maternal transmission of R in a cross with an r male, results in an RR maternal/r paternal endosperm with a solid red epidermis. In the reciprocal cross involving an r female, however, the rr/R endosperm is white with red sectors. A requirement for two doses of R www.nature.com/reviews/genetics © 2003 Nature Publishing Group REVIEWS POLYCOMB A class of proteins — originally described in Drosophila melanogaster — that repress the expression of the genes with which they are associated. There are several classes of polycomb proteins and in higher plants they are organized into gene families. for full pigmentation can be ruled out using B-A translocations (FIG. 4) to boost R copy number in the sperm — in the rr/RR genotype the kernels are still mottled62,63. Therefore, R is silenced in microgametophytes and the activity is restored stochastically during epidermal development, which results in mottled pigmentation. Ploidy constraints, the requirement for paternal transmission of some loci and the paternal downregulation of R expression — all of which are observed in the maize endosperm — have not been observed in the maize embryo. For example, some R alleles that are subject to paternal imprinting are uniformly visibly expressed in embryo structures62,63. These data have been interpreted to mean that any paternal mark is readily reversed after fertilization in the embryo but not in the endosperm. Alternatively, there could be differential imprinting of the two sperm in a pollen grain followed by preferential fertilization; in a few species, sperm dimorphism and preferential fertilization have been documented64. Arguing against this, using maize sperm that were rendered dimorphic after non-disjunction of B chromosomes (FIG. 4), Faure et al.65 showed that both sperm (one without B chromosomes and one with two B chromosomes) can fertilize the egg in vitro. However, in vivo there is preferential fertilization of the egg by the B-containing sperm, although this does not result from an inability of one sperm to fuse with the egg. Investigation of paternal gene expression in the maize embryo indicates that paternally transmitted green fluorescent protein (GFP) transgenes are expressed shortly after in vitro fertilization66. The highly condensed chromatin of maize sperm begins to decondense immediately after fertilization, GFP RNA is detectable at 4 hours and fluorescence is seen 6 hours after fertilization — many hours before the first zygotic cell division. Therefore, paternal alleles can be readily expressed in the maize zygote, at least in vitro. Future experiments in which a sperm is fused with a central cell might show that imprinting has occurred. The question of whether the paternal genome undergoes similar early activation in semine as well as in vitro has been difficult to answer. Studies of the expression of many different genes in developing endosperms and embryos of Arabidopsis have addressed this question with conflicting results. Using allele-specific reverse-transcription polymerase chain reaction (RTPCR) to differentiate maternally and paternally transmitted alleles from different strains, Vielle-Cazada et al. found that the zygotes and early embryos of Arabidopsis contained transcripts for the maternal allele of two genes, but that the paternal allele transcripts were not detectable for several days53. Also, using β-glucuronidase expression under the control of 18 different promoters that are active in zygotes and early embryos, they confirmed that maternal expression was readily scored but paternal expression was absent. These results were surprising, because most loci that are required for embryo development show classic Mendelian transmission. Self-pollination of a strain that is heterozygous at an embryo-lethal locus produces 25% dead embryos, not the 50% lethality that is expected from a maternal NATURE REVIEWS | GENETICS effect situation. In a few cases, such as emb30 (GNOM), mutants have a weak, early maternal defect that embryos recover from later in development53. Weijers et al. were quick to provide evidence that some paternally transmitted transgene markers could be expressed in a two-cell embryo (although most of their examples were from later stages) and argued that the genetic data show that paternal alleles must be expressed early in development67. These studies, and others that are summarized in TABLE 1, show non-equivalence of alleles: both are expressed, but with quantitative differences. Differing results probably reflect different experimental methods (assaying transcription only versus transcription and translation of the test gene) and the use of various developmental stages. Furthermore, transcript detection after fertilization could reflect either de novo transcription or transmission of stored mRNA. Whether or not the entire paternal genome is silent immediately after fertilization, it seems that the expression of maternal and paternal alleles is not equivalent for many loci during early development. The delay in full activation of the paternal genome might parallel the delay in zygotic gene activity in many animals — the plant version of the mid-blastula transition25. The maternal genome might escape this because the female gametes, unlike the sperm cells, do not have condensed chromatin and are more metabolically active. Regulation of imprinting in Arabidopsis The analysis of several maternal-effect lethal and lossof-DNA-methylation mutants in Arabidopsis has greatly advanced understanding of the mechanisms that underlie parental allele inequality and the complexity of its regulation. FERTILIZATION INDEPENDENT ENDOSPERM (FIE) mutants of Arabidopsis show a striking phenotype: the diploid central cell can proliferate in the absence of fertilization to form an aberrant endosperm68. So, in a mutant background, the usual requirement for paternal and maternal contributions to endosperm development are bypassed. FIE encodes a WD group POLYCOMB protein that acts to repress endosperm development68. If a FIE/fie heterozygous plant is pollinated by wild-type pollen, half of the embryos abort at the heart stage; in a reciprocal cross, all of the embryos are viable. Therefore, the maternal fie allele alone controls viability, and the paternal FIE allele is either imprinted completely or its expression is too late to restore normal endosperm development. A second locus, MEDEA (MEA), which encodes a SET (Su(var)3–9; enhancer of zeste; trithorax) domain polycomb protein, is similarly required in the megagametophyte to suppress precocious endosperm development69,70. After fertilization, only a maternally derived MEA allele suffices to prevent embryo abortion71. Paternal MEA alleles are not expressed in the endosperm, but are expressed in embryo structures70,72, echoing the story of the R alleles of maize. A third gene that might be subject to imprinting is FERTILIZATION INDEPENDENT SEED2 (FIS2), which encodes a zinc-finger protein and is also involved in the suppression of central cell proliferation before fertilization73. VOLUME 4 | MAY 2003 | 3 7 5 © 2003 Nature Publishing Group REVIEWS Table 1 | Parent-of-origin effects on gene expression in Arabidopsis Gene MEA Method Tissue Whole Embryo Endosperm RT-PCR of whole seed M>P – – H, T, WS, M 70 RT-PCR of dissected seed – M M M>P or M T WS M 70 – M=P M=P M=P RT-PCR of whole siliques M – – G 72 In situ hybridization – – M – 72 M Rare P M M,P ‡ in cyst PG EG 73 GUS protein fusion Embryo stage* References VPE RT-PCR of dissected seed – – M=P § WS 70 FIE RT-PCR of dissected seed – M=P M=P M=P M>P None None T WS M 75 GFP reporter – – – M None M,P‡ M None M,P‡ F–S G H 75 GUS protein fusion – – M M,P ‡ M M,P ‡ EG H 73 FIS2 GUS protein fusion – M M PG, later § 73 SIN1 GUS reporter – – – M None M,P ‡ M None M,P ‡ EG LG–T WS 79 PRL RT-PCR of whole siliques M M,P ‡ – – – – G T 53 GUS reporter – – – – M M,P || PG MG 53 In situ hybridization to GUS reporter – M=P M>P O, H 89 GUS reporter – M M PG–G § 53 18 genes EMB30/GN RT-PCR of whole siliques M – – PG 53 LTP1 2-component GUS reporter – – – – M>P M M M,P ‡ – – – – H, T O G H 67 90 CYC 2-component GUS reporter – – – M M# M,P ‡ – – – Z O LG 90 RPS5A GUS reporter – M,P – PG, later 67 *Embryo stages (in order of development): Z, zygote; F, four-cell; O, octant; S, sixteen-cell; EG, early globular; MG, mid globular; LG, late globular; H, heart; T, torpedo; WS, walking stick; M, mature. The stages before globular (G) are collectively referred to as pre-globular (PG). ‡ Maternal (M) and paternal (P) alleles were both expressed but no quantitative comparison was made. §Data not shown in the original reference. || M everywhere, P in cyst only. #M expressed, P variable. CYC, CYCLOIDEA; EMB30/GN, EMBRYO DEFECTIVE 30/GNOM; FIE, FERTILIZATION INDEPENDENT ENDOSPERM; FIS2, FERTILIZATION INDEPENDENT SEED 2; GFP, green flourescent protein; GUS, β-glucuronidase; LTP1, LIPID TRANSFER PROTEIN 1; MEA, MEDEA; PRL, PROLIFERA; RPS5A, RIBOSOMAL PROTEIN S5A; RT-PCR, reverse-transcription polymerase chain reaction; SIN1, SHORT INTEGUMENT 1; VPE, VACUOLAR PROCESSING ENZYME. An important step in understanding MEA was the recent discovery that a DNA glycosylase that is encoded by DEMETER (DME) is expressed in the central cell preceding fertilization. DME expression is required for the activation of the megagametophyte copy of MEA, which is consequently most strongly expressed in the central cell74. After fertilization, DME transcripts are no longer detected, although maternal MEA continues to be expressed. No DME transcripts are detected in microgametophytes, which indicates that one component of the maternal requirement for MEA is the selective activation of an essential gene in the megagametophyte. There are no detectable paternal MEA transcripts in the endosperm, except in transgenic plants that ectopically express DME after fertilization, in which the paternal 376 | MAY 2003 | VOLUME 4 allele is presumably activated by the DME protein. At present, MEA is the only known target of DME action. In contrast to MEA, the expression of paternally transmitted FIE alleles is detectable not only in the embryo but also in the endosperm after an initial maternal expression period75. So, although the FIE and MEA proteins interact75,76, the duration (or severity) of paternal lack of activity is distinctive. Furthermore, the imprinting of these loci is differentially sensitive to loss of function of the DECREASE IN DNA METHYLATION (DDM1) gene. Maternal defects of mea mutants can be rescued by ddm1 in two distinct ways. Seeds inheriting mea can complete development if they are also homozygous for ddm1 (REF. 72). Also, pollen from a ddm1/ddm1 MEA/MEA plant suppresses maternal mea www.nature.com/reviews/genetics © 2003 Nature Publishing Group REVIEWS PHENOCOPY A mimic of a phenotype that is caused by a known mutation. defects75. One hypothesis is that lowered methylation levels allow the expression of the paternal MEA allele and that this is sufficient to result in normal development; if so, paternal MEA should be independent of activation by DME. Alternatively, reducing methylation levels in sperm could allow the precocious expression of paternal genes that are important in development, circumventing some MEA-mediated process. Interestingly, a ddm1/ddm1 FIE pollen donor cannot relieve the defect in fie maternal mutants75. A second experiment exploited plants with low levels of DNA methyl transferase 1 activity, achieved using an antisense strategy. Paternal FIE delivery from the antisense METHYL TRANSFERASE 1 (MET1) plants, which lack almost 90% of normal DNA methylation levels, can rescue a fie1 maternal defect77. Also, pollination by MET1 antisense plants rescues both maternal fis2 and mea defects, but, unlike FIE, there is no requirement for a wild-type allele from the pollen parent73. Some caution is required in interpreting these results, as different ecotypes and allele combinations have been studied. Nonetheless, using genetic backgrounds with altered levels of DNA methylation shows possible mechanistic differences between FIE and MEA or FIS2. The available results indicate that the paternal hypomethylation of downstream targets can circumvent the requirement for MEA and FIS2, but not FIE. Unravelling the suite of target genes and learning how their expression is affected by maternal and paternal activity — or the lack of it — remains a substantial challenge. Significantly, the antisense MET1 plants show developmental defects similar to those observed in crosses between diploid and hexaploid Arabidopsis 78. The pollination of diploid females by MET1 antisense males PHENOCOPIES crosses with excess maternal dosage. Conversely, the pollination of MET1 antisense females by diploid males phenocopies crosses with excess paternal dosage. Presumably, the demethylation of alleles in the endosperm de-represses genes that are normally only expressed from the opposite parent. A final known player is SHORT INTEGUMENT 1 (SIN1), which encodes a Dicer homologue79,80 and is therefore predicted to process RNA into short molecules that can modulate gene expression post-transcriptionally by RNA destruction, or transcriptionally by directing chromatin remodelling. This gene acts in the sporophyte before meiosis to modulate megagametophyte development. We can now return to the initial quandary of a model in which paternal gene expression is generally repressed but there is Mendelian transmission of most developmental genes. The genetic results indicate that a paternal allele suffices for a normal embryo, but does not prove, or even require, that paternal alleles are routinely expressed immediately after fertilization. Recall that plant embryos can form from somatic cells and from haploid microgametophytic cells in culture — cells that must entirely reprogramme their gene expression. Clearly, forming an embryo in these situations does not require precisely imprinted maternal and paternal alleles, neither does it require either fertilization event. NATURE REVIEWS | GENETICS It seems possible that paternal alleles could be less readily expressed initially and still suffice to organize an embryo when they are eventually expressed hours or days after fertilization. It is probable that not all of these genes have parent-of-origin effects. Possible mechanisms for parent-of-origin effects are: gametophytic expression, such as DME; prolonged silencing of the paternal allele, such as R, MEA and FIS2; an early requirement for the gene before general paternal genome expression; or a combination of these mechanisms. Perhaps the maternal effects of the absence of MEA, FIE and FIS2 proteins are readily recognized because they encode components of the system that is responsible for differential maternal and paternal allele activity. Also, silencing of the paternal alleles of MEA and FIS2 persists much longer than general paternal silencing in Arabidopsis. Most loci that are expressed in the embryo might, in fact, be imprinted in the sperm for reduced initial activity, but also programmed to escape from this downregulation early in embryo development. The parental conflict theory and an extension of the effective haploid phase are two distinct phenomena that might have some overlapping elements. During the growth phase of the maize endosperm, it is clear that paternal alleles at many loci are required to attain a normal size, as reflected in the stringent requirement for the ploidy ratio of maternal:paternal chromosomes. It seems that maternal alleles have a dominant role immediately after fertilization, whereas paternal alleles are required later, with the consequence that both parents contribute uniquely to foster normal endosperm and embryo development. Conclusions The complexities of the flowering-plant life cycle have developmental, genetic and evolutionary consequences that generate fundamental differences between plants and animals. The existence of a genetically active haploid phase shows that plants accumulate fewer deleterious recessive alleles and might be more tolerant of DNA transposons than are animals. Conversely, the absence of a germline and the late switch of somatic lineages in the plant shoot to floral development allow genetic mutations to accumulate and be represented in the gametes. Therefore, plants generate greater genetic diversity, but, through haploid selection, impose more stringent selection than animals. Despite this function of the haploid phase, the diversity of structures and pathways that are expressed in gametophytes has steadily declined over evolutionary time. It is plausible that, with the advent of double fertilization in flowering plants, a supplementary mechanism has evolved whereby parent-of-origin effects, including imprinting and ploidy, can act as an extra quality check. The delay in the full activation of the paternal genome extends the effective haploid phase to early embryo development. Also, flowering plants, like the mammals, conduct a rigorous genome quality check after fertilization, but do so primarily by assessing the ploidy level in the endosperm — an accessory seed-restricted organ that is produced by a second fertilization event. In many VOLUME 4 | MAY 2003 | 3 7 7 © 2003 Nature Publishing Group REVIEWS plant species, a normal endosperm is a prerequisite for robust embryo development and seed success; therefore, ploidy requirements and the imprinting of endospermexpressed alleles can act to ensure that plant sexual reproduction only occurs if two sets of appropriately programmed gametes are united. Given the present 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. 17. 18. 19. 20. 21. 22. 23. 24. 25. 26. 378 Meyerowitz, E. M. Plants compared to animals: the broadest comparative study of development. Science 295, 1482–1485 (2002). Gu, Z. L. et al. Role of duplicate genes in genetic robustness against null mutations. Nature 421, 63–66 (2003). Mascarenhas, J. P. Gene activity during pollen development. Annu. Rev. Plant Physiol. Plant Mol. Biol. 41, 317–338 (1990). Mascarenhas, J. P. Pollen gene-expression — molecular evidence. Int. Rev. Cytol. 140, 3–18 (1992). Brendel V., Kurtz, S. & Walbot V. Comparative genomics of Arabidopsis and maize: prospects and limitations. Genome Biol. Rev. 3, 1005.1–1005.6 (2002). John, R. M. & Surani, M. A. Genomic imprinting, mammalian evolution, and the mystery of egg-laying mammals. Cell 101, 585–588 (2000). Otto, S. P. & Marks, J. C. Mating systems and the evolutionary transition between haploidy and diploidy. Biol. J. Linn Soc. 57, 197–218 (1996). Hughes, J. S. & Otto S. P. Ecology and the evolution of biphasic life cycles Am. Nat. 154, 306–320 (1999). Mable, B. K. & Otto, S. P. The evolution of life cycles with haploid and diploid phases. Bioessays 20, 453–462 (1998). Agrawal, A. F. & Chasnov, J. R. Recessive mutations and the maintenance of sex in structured populations. Genetics 158, 913–917 (2001). Raven, P. H., Evert, R. F. & Eichhorn, S. E. Biology of Plants 6th edn (Freeman & Co., New York, 1999). Joint Genome Institute. Chlamydomonas reinhardtii v.1.0, http://genome.jgi-psf.org/chlre1/chlre1.home.html (2002). Harris, E. H. Chlamydomonas as a model organism. Annu. Rev. Plant Physiol. Plant Mol. Biol. 52, 363–406 (2001). Bonhomme, S. et al. G T-DNA mediated disruption of essential genes in Arabidopsis is unexpectedly rare and cannot be inferred from segregation distortion alone. Mol. Gen. Genet. 260, 444–452 (1998). Desfeux, C., Clough, S. J. & Bent, A. F. Female reproductive tissues are the primary target of Agrobacterium-mediated transformation by the Arabidopsis floral-dip method. Plant Physiol. 123, 895–904 (2000). Bechtold, N. et al. The maternal chromosome set is the target of the T-DNA in the in planta transformation of Arabidopsis thaliana. Genetics 155, 1875–1887 (2000). Coe, E. H., Neuffer, M. G. & Hoisington, D. A. in Corn and Corn Improvement (eds Sprague, G. F. & Dudley, J. W.) 81–258 (American Soc. Agronomy, Inc., Madison, Wisconsin, 1988). Maize Gene Discovery Project. ZmDB: Maize Phenotype Database. http://www.zmdb.iastate.edu/zmdb/ phenotypeDB/index.htm (2003). Patterson, E. B. in Maize Breeding and Genetics (ed. Walden, D. B.) 693–710 (Wiley and Sons, New York, 1978). Vision, T. J., Brown, D. G. & Tanksley, S. D. The origins of genomic duplications in Arabidopsis. Science 290, 2114–2117 (2000). This article defines the extent and timing of duplication events on the basis of genomic sequence, and illustrates important principles about the genomic duplications that have been found in all flowering plants examined so far. Blanc, G., Barakat, A., Guyot, R. Cooke, R. & Delseny, I. Extensive duplication and reshuffling in the Arabidopsis genome. Plant Cell 12, 1093–1101 (2000). The Arabidopsis Genome Initiative. Nature 438, 796–815 (2000). Gaut, B. S., Le Thierry d’Ennequin, M., Peek, A. S. & Sawkins, M. C. Maize as a model for the evolution of plant nuclear genomes. Proc. Natl Acad. Sci. USA 97, 7008–7015 (2000). Lynch, M. & Conery, J. S. The evolutionary fate and consequences of duplicate genes. Science 280, 1151–1155 (2000). Gilbert, S. F. Developmental Biology 5th edn (Sinauer Associates, Inc. Sunderland, Massachusetts, 2000). Starz-Gaiano, M. & Lehmann, R. Moving towards the next generation. Mech. Dev. 105, 5–18 (2001). dominance of flowering plants in terrestrial habitats, the double fertilization event resulting in a sexually derived nutritive tissue seems to have been an evolutionary innovation of great consequence and we believe that this is, in part, because of the genome quality checks that occur in this organ. 27. Hogan, B. Decisions, decisions. Nature 418, 282–283 (2002). 28. Weigel, D. & Jürgens, G. Stem cells that make stems. Nature 415, 751–754 (2002). 29. Simpson, G. G. & Dean, C. Flowering — Arabidopsis, the rosetta stone of flowering time? Science 296, 285–289 (2002). 30. Zhao, D. Z., Wang, G. F., Speal, B. & Ma, H. The EXCESS MICROSPOROCYTES1 gene encodes a putative leucinerich repeat receptor protein kinase that controls somatic and reproductive cell fates in the Arabidopsis anther. Genes Dev. 16, 2021–2031 (2002). This study addresses the important question of how pre-meiotic cells are specified in the anther, and indicates that there is a trade off between somatic and pre-germinal cell proliferation that is regulated through the kinase defined in the genetic study. 31. Canales, C., Bhatt, A. M., Scott, R. & Dickinson, H. EXS a putative LRR receptor kinase regulates male germline cell number and tapetal identity and promotes seed development in Arabidopsis. Curr. Biol. 12, 1718–1727 (2002). 32. Yang, W. C., Ye, D., Xu, J. & Sundaresan, V. The SPOROCYTELESS gene of Arabidopsis is required for initiation of sporogenesis and encodes a novel nuclear protein. Genes Dev. 13, 2108–2117 (1999). 33. Sorensen, A., Guerineau, F., Canales-Holzeis, C., Dickinson, H. G. & Scott, R. J. A novel extinction screen in Arabidopsis thaliana identifies mutant plants defective in early microsporangial development. Plant J. 29, 581–594 (2002). 34. Sheridan, W. F., Golubeva, E. A., Abrhamova, L. I. & Golubovskaya, I. N. The mac1 mutation alters the developmental fate of the hypodermal cells and their cellular progeny in the maize anther. Genetics 153, 933–941 (1999). A genetic and anatomical study of the consequences of the excess proliferation of germinal cells at the expense of strictly somatic cells in maize, for comparison with Zhao et al. (reference 30) who examined a parallel phenomenon in Arabidopsis. 35. Drews, G. N. & Yadegari, R. Development and function of the angiosperm female gametophyte. Annu. Rev. Genet. 36, 99–124 (2002). 36. Klekowski, E. J., Lowenfeld, R. & Klekowski, E. H. Mangrove genetics. 4. Postzygotic mutations fixed as periclinal chimeras. Int. J. Plant Sci. 157, 398–435 (1996). 37. Chaparro, J. X., Werner, D. J., Whetten, R. W. & O’Malley, D. M. Characterization of an unstable anthocyanin phenotype and estimation of somatic mutation-rates in peach. J. Hered. 86, 186–193 (1995). 38. Pineda-Krch, M. & Fagerstrom, T. On the potential for evolutionary change in meristematic cell lineages through intraorganismal selection. J. Evol. Biol. 12, 681–688 (1999). 39. Walbot, V. Sources and consequences of phenotypic and genotypic plasticity in flowering plants. Trends Plant Sci. 1, 27–32 (1996). 40. Rideout, W. M., Eggan, K. & Jaenisch, R. Nuclear cloning and epigenetic reprogramming of the genome. Science 293, 1093–1098 (2001). 41. Normark, B. B. The evolution of alternative genetic systems in insects. Annu. Rev. Entomol. 48, 397–423 (2003). 42. Crews, D. & Fitzgerald, K. T. Sexual behavior in parthenogenetic lizards (Cnemidophorus). Proc. Natl Acad. Sci. USA 77, 499–502 (1980). 43. Chaudhury, A. M. et al. Control of early seed development. Annu. Rev. Cell Dev. Biol. 17, 677–699 (2001). 44. Grimanelli, D., Leblanc, O., Perotti, E. & Grossniklaus, U. Developmental genetics of gametophytic apomixes. Trends Genet. 17, 597–604 (2001). 45. von Arnold, S., Sabala, I., Bozhkov, P., Dyachok, J. & Filonova, L. Developmental pathways of somatic embryogenesis. Plant Cell Tissue Organ Cult. 69, 233–249 (2002). 46. Wan, Y., Petolino, J. F. & Widholm, J. M. Efficient production of doubled haploid plants through colchicine treatment of anther-derived maize callus. Theor. Appl. Genet. 77, 889–892 (1989). | MAY 2003 | VOLUME 4 47. Chase, S. S. Monoploids and diploids of maize: a comparison of genotypic equivalents. Am. J. Bot. 51, 928–933 (1964). 48. Chase, S. S. Monoploids and monoploid derivatives of maize (Zea mays L.). Bot. Rev. 35, 117–167 (1969). 49. Kermicle, J. L. Androgenesis conditioned by a mutation in maize. Science 166, 1422–1424 (1969). 50. Vinkenoog, R. & Scott, R. J. Autonomous endosperm development in flowering plants: how to overcome the imprinting problem? Sex. Plant Reprod. 14, 189–194 (2001). 51. Arditti, J. Fundamentals of Orchid Biology (Wiley, New York, 1992). 52. Haig, D. & Westoby, M. Parent specific gene expression and the triploid endosperm. Am Nat. 134, 147–155 (1989). 53. Vielle-Cazada, J.-P., Baskar R. & Grossniklaus, U. Delayed activation of the paternal genome during seed development. Nature 404, 91–94 (2000). Based on differential RT-PCR to detect maternal versus paternal transcripts of two endogenous genes, and reporter protein expression from promoter:GUS fusion genes, the authors propose that the entire paternal genome is silent for up to 2 days after fertilization. 54. Lin, B.-Y. Association of endosperm reduction with parental imprinting in maize. Genetics 100, 475–486 (1982). This report shows that the crucial parental genome ratios that are required for normal seed development occur in the endosperm, by separately manipulating ratios in the embryo and endosperm using B-A translocations. 55. Birchler, J. A. & Hart, J. R. Interactions of endosperm size factors in maize. Genetics 117, 307–317 (1987). 56. Birchler, J. A. Dosage analysis of maize endosperm development. Annu. Rev. Genet. 27, 181–204 (1993). 57. Lin, B.-Y. Ploidy barriers to endosperm development in maize. Genetics 107, 103–115 (1984). 58. Haig, D. & Westoby, M. Genomic imprinting in endosperm: its effect on seed development in crosses between species, and between different ploidies of the same species, and its implications for the evolution of apomixes. Phil. Trans R. Soc. Lond. B 333, 1–13 (1991). 59. Scott, R. J., Spielman, M., Bailey, J & Dickinson, H. G. Parent-of-origin effects on seed development in Arabidopsis thaliana. Development 125, 3329–3341 (1998). 60. Kermicle, J. L. Dependence of the R-mottled aleurone phenotype in maize on mode of sexual transmission. Genetics 66, 69–85 (1970). The first report of an imprinted gene, and a model of the genetic analysis of parent-of-origin effects. 61. Ludwig, S. R., Habera, L. F., Dellaporta, S. L. & Wessler, S. R. Lc, a member of the R gene family responsible for tissuespecific anthocyanin production, encodes a protein similar to transcriptional activators and contains the mychomology region. Proc. Natl Acad. Sci. USA 86, 7092–7096 (1989). 62. Alleman, M. & Doctor, J. Genomic imprinting in plants: observations and evolutionary implications. Plant Mol. Biol. 43, 147–161 (2000). 63. Kermicle, J. L. & Alleman, M. Gametic imprinting in maize in relation to the angiosperm life cycle. Development (Suppl.) 9–14 (1990). 64. Southworth, D., Strout, G. & Russell, S. D. Freeze-fracture of sperm of Plumbago zeylanica L. in pollen and in vitro. Sex. Plant Reprod. 10, 217–226 (1997). 65. Faure J-E. et al. Double fertilization in maize: the two male gametes from a pollen grain have the ability to fuse with egg cells. Plant J. 33, 1051–1062 (2003). 66. Scholten, S., Lörz, H. & Kranz, E. Paternal mRNA and protein synthesis coincides with male chromatin decondensation in maize zygotes. Plant J. 32, 221–231 (2002). This work reports the early activation of spermtransmitted green fluorescent protein in zygotes after in vitro fertilization. 67. Weijers, D., Geldner, N., Offringa, R. & Jürgens, G. Seed development: early paternal gene activity in Arabidopsis. Nature 414, 709–710 (2001). www.nature.com/reviews/genetics © 2003 Nature Publishing Group REVIEWS 68. Ohad, N. et al. Mutations in FIE, a WD polycomb group gene, allow endosperm development without fertilization. Plant Cell 11, 437–415 (1999). 69. Kiyosue T. et al. Control of fertilization-independent endosperm development by the MEDEA polycomb gene Arabidopsis. Proc. Natl Acad. Sci. USA 96, 4186–4191 (1999). 70. Kinoshita, T., Yadegari, R., Harada, J. J., Goldberg, R. B. & Fischer, R. L. Imprinting of the MEDEA polycomb gene in the Arabidopsis endosperm. Plant Cell 11, 1945–1952 (1999). More information about the expression domain and role of an important gene in the maternal control of endosperm and embryo development. 71. Grossniklaus, U., Vielle-Calzada, J. P., Hoeppner, M. A. & Gagliano, W. B. Maternal control of embryogenesis by medea, a polycomb group gene in Arabidopsis. Science 280, 446–450 (1998). First report of an imprinted gene in Arabidopsis, which, along with reference 70, begins to define the scope of parent-of-origin effects. 72. Vielle-Calzada, J.-P. et al. Maintenance of genomic imprinting at the Arabidopsis medea locus requires zygotic DDM1 activity Genes Dev. 13, 2971–2982 (1999). This report lays the framework for establishing the link between DNA methylation and the setting up and maintenance of the repressive imprinting of paternal alleles. 73. Luo, M., Bilodeau, P., Dennis, E. S., Peacock, W. J. & Chaudhury, A. Expression and parent-of-origin effects for FIS2, MEA, and FIE in the endosperm and embryo of developing Arabidopsis seeds. Proc. Natl Acad. Sci. USA 97, 10637–10642 (2000). 74. Choi, Y. H. et al. DEMETER, a DNA glycosylase domain protein, is required for endosperm gene imprinting and seed viability in Arabidopsis. Cell 110, 33–42 (2002). Elucidation of a gene with a product that acts on MEDEA, a gene imprinted to be expressed from maternal but not paternal alleles, in the endosperm. Expression of DEMETER in megagametophytes, but 75. 76. 77. 78. 79. 80. 81. 82. 83. not microgametophytes, establishes conditions for the parent-of-origin effects of MEDEA. Yadegari, R. et al. Mutations in the FIE and MEA genes that encode interacting polycomb proteins cause parent-oforigin effects on seed development by distinct mechanisms. Plant Cell 12, 2367–2381 (2000). Shows that the protein products of two imprinted genes interact but can be distinguished in their impact; the observations indicate considerable complexity in the regulation of parent-of-origin effects. Spillane, C. et al. Interaction of the Arabidopsis Polycomb group proteins FIE and MEA mediates their common phenotypes. Curr. Biol. 10, 1535–1538 (2000). Vinkenoog, R. et al. Hypomethylation promotes autonomous endosperm development and rescues postfertilization lethality in fie mutants. Plant Cell 12, 2271–2282 (2000). Adams, S., Vinkenoog, R., Spielman, M., Dickinson, H. G. & Scott, R. J. Parent-of-origin effects on seed development in Arabidopsis thaliana require DNA methylation. Development 127, 2493–2502 (2000). Golden, T. A., et al. Short Integuments1/suspensor1/Carpel Factory, a Dicer homolog, is a maternal effect gene required for embryo development in Arabidopsis. Plant Physiol. 130, 808–822 (2002). Ray, S., Golden, T. & Ray, A. Maternal effects of the short integument mutation on embryo development in Arabidopsis. Dev. Biol. 180, 365–369 (1997). Kinoshita, T., Harada, J. J., Goldberg, R. B. & Fischer, R. L. Polycomb repression of flowering during early plant development. Proc. Natl Acad. Sci. USA 98, 14156–14161 (2001). Bennetzen, J. L. & Kellogg, E. A. Do plants have a one-way ticket to genomic obesity? Plant Cell 9, 1509–1514 (1997). Sanmiguel, P. & Bennetzen, J. L. Evidence that a recent increase in maize genome size was caused by the massive amplification of intergene retrotransposons. Annals Bot. 82, 37–44 (1998). NATURE REVIEWS | GENETICS 84. Walbot, V. & Rudenko, G. N. in Mobile DNA II (eds. Craig, N. L., Craigie, R., Gellert, M. & Lambowitz, A.) 533–564 (American Soc. Microbiology, Washington DC, 2002). 85. Robertson, D. S. Mutator activity in maize: timing of its activation in ontogeny. Science 218, 1515–1517 (1981). 86. Williams, J. H. & Friedman, W. E. Identification of diploid endosperm in an early angiosperm lineage. Nature 415, 522–526 (2002). 87. Friedman, W. E. The evolution of double fertilization and endosperm: an ‘historical’ perspective. Sex. Plant Reprod. 11, 6–16 (1998). 88. Weidinger, G. et al. Regulation of zebrafish primordial germ cell migration by attraction towards an intermediate target. Development 129, 25–36 (2002). 89. Springer, P. S., Holding, D. R., Groover, A., Yordan, C. & Martienssen, R. A. The essential Mcm7 protein PROLIFERA is localized to the nucleus of dividing cells during the G1 phase and is required maternally for early Arabidopsis development. Development 127, 1815–1822 (2000). 90. Baroux, C., Blanvillain, R. & Gallois, P. Paternally inherited transgenes are downregulated but retain low activity during early embryogenesis in Arabidopsis. FEBS Lett. 509, 11–16 (2001). Online links DATABASES The following terms in this article are linked online to: MaizeGDB: http://www.maizegdb.org ig | R TAIR: http://www.arabidopsis.org DDM1 | DME | emb30 | FIE | FIS2 | MEA | SIN1 FURTHER INFORMATION Virginia Walbot’s laboratory: http://www.stanford.edu/%7Ewalbot Zea mays Database (ZmDB): http://zmdb.iastate.edu Access to this interactive links box is free online. VOLUME 4 | MAY 2003 | 3 7 9 © 2003 Nature Publishing Group