* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Nucleosomal structure of sea urchin and starfish sperm chromatin
Transcriptional regulation wikipedia , lookup
Genomic library wikipedia , lookup
DNA profiling wikipedia , lookup
SNP genotyping wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Non-coding DNA wikipedia , lookup
Point mutation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Molecular cloning wikipedia , lookup
Biochemistry wikipedia , lookup
Artificial insemination wikipedia , lookup
DNA supercoil wikipedia , lookup
Histone acetylation and deacetylation wikipedia , lookup
Biosynthesis wikipedia , lookup
Gel electrophoresis wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Agarose gel electrophoresis wikipedia , lookup
Community fingerprinting wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
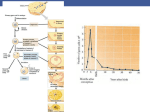
volume 9 Number 3 1981 Nucleic A c i d s Research Nucleosomal structure of sea urchin and starfish sperm chromatin. Histone H2B is possibly involved in determining the length of linker DNA I.A.Zalenskaya, V.A.Pospelov, A.O.Zalensky* and V.I.Vorob'ev Institute qf Cytology, Academy of Sciences of the USSR, Leningrad 190121, and institute of Marine Biology, Far East Scientific Center, Academy of Sciences of the USSR, Vladivostok 690022, USSR Received 16 December 1980 ABSTRACT Comparison has been made between sea urchin and starfish, sperm chromatin. The only protein by which chxomatins from these sources differ significantly is histone H2B. Sea urchin sperm H2B is known to contain an elongated N-texminal region enriched in Arg. Analysis of the micrococcal nuclease digests of sea urchin and starfish nuclei in one- and two-dimensional electrophoresis has shown that sperm chromatin of both animals consists of repeated units similar in general features to those of rat thymus or liver. However, i)NA repeat length in chromatin of sea urchin sperm (237 bp) is higher than that of starfish sperm (224 bp), while the core DNA length does not differ and is the same as in the chromatin of rat liver or thymus. A suggestion has been made that the N-teuninal region of histone H2B is associated with the linker DNA and is responsible for the increased length of sea urchin linker DNA. INTRODUCTION In a periodically repeated chromatin subunit, a nucleosome, two main parts can be distinguished: the core particle and the region between two neighbour core particles which is termed the linker (for rev. see 1, 2 ) . The core particle consists of DNA n/\i\Q base pairs (bp) long combined with an octamer of "small" histones (H4+H3+H2A+H2B)x2. The linker DNA is probably associated with histone H1 (3-6). While the length of the core DNA is the same in all the cells studied, the length of the linker DNA varies from 15 to 100 bp depending on the cell type. It has been suggested that the conservation in length of the core DNA is due to the conservation in primary structure of the core histones, while the variability of the linker DNA is connected with changes in the structure of histone H1 (6, 7 ) . IIRL Press Limited. 1 Falconberg Court, London W 1 V 5FG. U.K. 47 3 Nucleic Acids Research At the same time, some recent investigations have shown that the linker DNA might also be associated with histones H2A and H2B (8-10). These results as well as data on some variability of histones H2A and H2B (11) lead to an assumption that histoaes H2A and H2B could, together with H1, define the length of the linker DNA (8, 1, 2). In this paper we report findings obtained losing a comparative approach on the role of hlstone H2B in the organization of a nucleosome. Comparison has been made between sea urchin and starfish sperm chromatins which contain substantially different hiatones H2B (12-15). At the same time their histones H4, H3, H2A and, what is especially important for this study, histone H1 are very similar (see Results and Discussion). We believe that the impact of histone H2B on the structure of a nucleosome may be inferred from comparison of parameters of the chromatin subunits from these two sources. The sperm cells of these animals are apparently very convenient for such analysis: (I) they represent a highly homogeneous cell population; (ii) the content of non-histone proteins in sperm cells is very low, and therefore their influence on nucleosomal structure can be excluded. The results presented here are consistent with the point of view that histone H2B is associated not only with the core particle, but also with the linker DNA and affects the length of the latter. MATERIALS AND METHODS Biological material. Sea urchin StronKylocentrotus intermedlus and starfish Aphelasteriaa japonica (Sea of Japan) were used in this study. Preparation of sperm cells and sperm nuclei. The procedures has been described else--where (14, 15)« Briefly, the sperm was collected by centrifugation, washed in STC buffer (0.25 M sucrose, 10 mil Tris-HOl, pH 7.5, 3 mM Ca01 2 , 0.1 mM phenylmetylsulfonylfluorid - PMSF), and disrupted in the same buffer. The nuclei were washed once in. STC buffer, containing 0.5% Triton X-100 and twice in the STC buffer without Triton. 474 Nucleic Acids Research Bat thymus nuclei were isolated by the same method as sperm nucleio Eat liver nuclei were additionally puriiffied by centrifugation through 2.2 M sucrose, 10 mM Tris-HCl, pH 7»5» 3 mM 0aCl 2 , 0.1 mM PMSP. Histone isolation. Total histone and histone fractions H1 and H2B were isolated according to Oliver et al. (16). The histones H1 and RUB from sea urchin and starfish sperm have been obtained in highly purified form by this method (14, 15). The purity of the fractions being analysed in the present paper is no less than 98% (as has been judged from electrophoretic data). Nuclease digestion. Nuclei were suspended at a concentration of 1 ing DNA per ml of buffer containing 0.3 M sucrose, 10 mM Tris-HCl, pH 7.5, 1 mM CaCl^, 0.1 mM FMSF. Micrococcal nuclease (Worthington) was added up to a concentration of 30 units per mg of DNA. The digestion was carried out at 37° 0 during 3» 15 and 30 mia. The reaction was stopped by chilling the samples on ice. The samples were centrifuged at 6,000 rpm for 2 min. The nuclear pellet was lysed in 5 mM Tris-HCl, 2 mM BDTA, pH 7«5» After 15-20 min the lysed nuclei were centrifuged at 6,000 rpm for 10 min. She supernatant, which contained soluble chromatin fragments was used for electrophoretic analysis and isolation of DNA fragments. Eat liver nuclei were digested with DNase I as described in (17). DNA isolation. DNA was isolated from the soluble chromatin fragments as described in (10). Gel electrophoresis. The chromatin fragments were separated on 5% polyacrylamide gel in 10 mM Tris-borate buffer, pH 8.3 containing 1 mM EDTA (TBE buffer) as described in (18). To determine the DNA repeat length, the DNA fragments isolated from nuclei treated with micrococcal nuclease during 3 min were separated on 1,8% agarose gel in 20 mM Tris-acetate buffer, pH 7.7 containing 2 mM EDTA (buffer TAB). Th.e values for the DNA repeat length were obtained from the slope of the regression line from a plot of fragment size versus band number as described in (7)« The DNA fragments, isolated from microccccal nuclease digests of rat thymus or liver nuclei were used as standards. A value of 195 bp was established for both rat thymus and rat liver DNA repeat using BFI DNA fragments from 475 Nucleic Acids Research phage rfX174 produced by r e s t r i c t a s e Bsp. The DNA fragments were a kind g i f t from Dr. R.E.Streeck and Dr. H.G.Zachau. To compare DNA length of the core particles the DNA fragments were electropuoresed on 6.5% acrylamide gel in TAE buffer and on 10% acrylamide gel in double TBE buffer, containing 6.5 M urea. Before electrophoresis on aerylamide-urea gel the DNA samples were dissolved in 8 M urea and denatured by boiling for 5 min. The electrophoresis of histones was carried out on 15% polyacrylamide gel according to Laemmli (19). A second dimension of electrophoresis was used to analyse the protein and DNA composition of the chromatin fragments. The s t r i p s of 5% polyacrylamide gel after low ionic strength e l e c trophoresis of chromatin fragments were incubated for 30 min in 0.0625 M Tris-HCl, pH 6.8 containing O.2J6 SDS and l a i d horizont a l l y on the top of the preformed slab of Laemmli g e l . After completion of the run, the g e l was washed in 50% ethanol to remove SDS. The gel was f i r s t stained with ethidium bromide (0.1 mg/ml) to visualize DNA, and then with Coomassi b r i l l i a n t blue to stain the proteins. Amino acid composition was determined on KLM (Hitachi) amino acid analyzer following 24 h hydrolysis of the proteins in 6 N HG1 at 110° C. No corrections have been made for hydrolytic losses. KESUI/rS Histones from sperm of the sea urchin Strongylocentrotua inteimedius and the starfish Aphelasterias .japonica separated by SDS-gel electrophoresis are shown in fig. 1 (rat liver and rat or calf thymus are used in this study as standard objects). Histones H3 and H4 have identical electrophoretic properties in the sperm cells and in calf thymus. Electrophoretic mobility of histone H2A from sperm of the sea urchin and the starfish is practically the same, and it is higher than that of calf thymus. Also similar are the sperm histones H1, both migrating slower than calf thymus H 1 . On the other hand, histones H£B from sperm of starfish and 476 Nucleic Acids Research Pig. 1. SDS gel electrophoresis of histones: (a) sea urchin sperm, (t>) starfish sperm, (c) calf thymus. sea urchin, unlike the histones H4, H3, H2A and H1, differ markedly. Mol* w. of H2B subfractions from sperm of the sea urchin S. intermedlus as determined by SflS-gel electrophoresis are 17,550 and 16,200 dalton, while mol. w. of starfish sperm H2B is close to 13,000 (14, 15). The electrophoretic data on similarity and difference of the histone fractions was confirmed by the comparison of their amino acid compositions (Table 1 , 2 ) . A characteristic feature of histone H1 from sperm of both organisms is a high arginine content - 16.4 and 14.5 mole% as well as a high ratio of basic to acidic amino acids - 9.3 and 8.8 in sea urchin and starfish, respectively. In calf thymus H1 arginine content is 2 mole% and basic to acidic amino acids ratio is 5*9 (20). Amino acid composition of histone H2A is similar in sea urchin sperm, starfish gonade and calf thymus (Table 2) (21-23). Significant difference in amino acid composition is found between histones H2B from sea urchin and starfish sperm. The most striking variation is in arginine content. The values are: Nucleic Acids Research Table 1. Amino acid composition of some histories from the sea urchin Strongylocentrotus intermedius and the starfish Aphelasterias .japonica (values are moles # ) . Lys His Arg Asp Tnr Ser Glu Pro Gly Ala Cys Val Met lie Leu Tyr Phe Lys/Arg basic/acidic H1 H1 (15) sea urchin starfish 24.4 27.8 0.8 1.2 16.4 14.5 1.5 2.7 2.4 1.7 7*7 2.2 8.1 2.3 21.4 — 4.8 _ 1.3 2.2 traces traces 1.7 9.3 9.5 3.0 4.6 4.6 31.4 - 1.7 traces 0.8 0.7 traces traces 1.7 8.8 H2B 1 +H2b 2 sea urchin 12.6 1.4 18.4 3.9 5.9 11.2 6.6 4.2 9.4 8.1 - 5-8 traces 3.6 4.7 2.5 1.4 0.7 3.1 H2B (15) starfish 11.7 1.4 7.7 5.8 7.4 9^1 2.0 10.1 11.5 — 7.7 5.3 5.2 3.3 traces 2.5 1.5 1.4 Table 2. Lys to Arg and basic to acidic amino acids ratios of some histone fractions Lys/Arg basic/acidic H1 Sperm of the sea urchin Stronsylocentrotus intermedius 1.7 9.3 Sperm of the starfish 8.8 Aphelasterias japonica 1.7 Calf thymus (20) 15.2 5.9 H2A Sperm of the sea urchin 1.4 1.2 Psammechinus miliaris (21) Gonades of the starfish 1.4 Asterias rubens (22) 1.0 1.2 Calf thymus (23) 1.5 H2B Sperm of the sea urchin StronKylocentrotus intermedius 3.1 0.7 Sperm of the strafish 1.4 Aphelasterias ,-japonica 1.5 Calf thymus (24) 2.5 1.9 478 Nucleic Acids Research 18.4- mole% in sea urchin sperm in comparison with 7.7 mole$J in starfish and 6.2 mole% in calf thymus HZB (24). The nuclei from sea urchin and starfish sperm, rat thymus and rat liver were digested with micrococcal nuclease. The chromatin fragments from the digests were separated by low ionic strength polyacrylamide gel electrophoresis (Fig. 2 ) . To analyse the DMA. and the proteins which are contained in the chromatin particles, second dimension electrophoresis was carried out (Pig. 3 ) . The general electrophoretic pattern of chromatin fragments from sperm of sea urchin and starfish and from rat thymus is similar. Zones of mononucleosomes lacking H1 (MN-H1), mononucleosomes containing H1 (MN+H1), dinucleosomes (DN) and so on are displayed. At the same time, some specific features may be noted. In sperm of both organisms (Fig. 2a, c) the zone of MN-H1 migrates faster than that of rat thymus (Fig. 2b). Furthermore, in sea urchin sperm the complete nucleosome particle DN .—! MN+H1 MN-H1 —- Fig. 2. Chromatin fragments obtained from 15 min micrococcal nuclease digests: (a) sea urchin sperm, (b) rat thymus, (c) starfish sperm. 479 Nucleic Acids Research First MN >o <n c MN dimension DN .MN MN H1 ~H1 + DN +MN.MN I:'- c o w — E Fig. 3« Two-dimensional electrophoresis. First dimension - separation of chromatin fragments on 5% acrylamide gel. Second dimension - separation of proteins and DNA on 15% acrylamide gel. (a) sea urchin sperm, (b) starfish sperm, (c) rat thymus. (MN+H1) separates into two subfractions. The data from two-dimensional electrophoresis show that both subfractions contain the same set of histones but their DNAs differ in length (Fig. 3a). It is not yet clear whether tae fact that the complete nucleosome displays a double zone demonstrates the heterogeneity of native chromatin or this reflects the different extent of nuclease digestion of the same particle. Since both subfractions have the same protein composition and contain DNA of different length, the second possibility seems to be more plausible. One of the main parameters of a nucleosome which supposedly could be defined by the proteins it contains is the DNA re480 Nucleic Acids Research peat length. The size of DNA fragments from sea urchin and starfish sperm micrococcal nuclease digests was determined on 1.856 agarose gel (Fig. 4, Table 3) as described in Material and Methods. The method of nucleosome repeat determination used eliminates the effect of UNA degradation from the ends (1, 7 ) . DNA fragments from rat liver digest were used as markers (see Materials and Methods). A value of 237 (+5) bp was found for sea urchin S.intermedius sperm DNA repeat in agreement with data reported for other sea urchin species (25, 26). A DMA repeat length of 224 (*6) bp was determined for starfish sperm. Thus, DBA contained in the complete nucleosome from sperm of the sea urchin S.intermedius is about 13 bp longer than that from sperm of the starfish A..iaponica. In order to find out whether the idea on the core DNA. inva- B 1353 1078 872 606 310 2 34 1 94 Fig.4. Agaro8e gel electrophoresis of DNA fragments from micrococcal nuclease digests. A: (a) rat thymus, 5 min digestion, (b) rat liver, 3 min digestion, (c) restriction fragments, B: a) and (b) sea urchin sperm, (a) 15 min, (b) 3 min digestion, fc) rat liver. 3 min digestion, (d) and (e) starfish sperm, [d) 3 min, (e) 15 min digestion. 481 Nucleic Acids Research Table 3. Sizes of DNA fragments isolated from short (3 min) micrococcal nuclease digest of rat liver and sea urchin and starfish sperm nuclei as determined from mobilities in agarose gel (see Materials and Methods). Band number 1 2 3 4 5 6 7 8 Slope Rat liver marker Base pairs Sea urchin sperm 370 570 775 960 1150 1350 450 700 930 1165 1410 1630 1870 195+3 237+5 Starfish sperm 425 627 850 1090 1340 1530 224+6 riability can be extended to toe cells studied here, the DHA fragments isolated from 30 min micrococcal nuclease digests were compared on polyacrylamide gel in two systems. Data from electrophoresis of DNA fragments on nondenaturating 6.5% acrylamide gel show that the position of the sharp edge of the fastest band which corresponds to the core particle DNA (7) is indistinguishable from each other in rat liver and sea urchin and starfish sperm (Fig.5). The same results were obtained when DNA from 30 min micrococcal digests denatured by boiling for 5 min in 8 M urea was electrophoresed on denaturating 10% acrylamide gel containing urea. (Pig.6). The fastest bands of singlestranded DNA from all three sources have a mobility that corresponds to 146 bases (27). Thus, there is no doubt that the nucleosomes from sea urchin and starfish sperm differ in the linker DNA length. DISCUSSION It is very likely that variation in length of linker DNA is connected with changes in structure of the proteins which interact with the DNA. Histone H1 might be one of such proteins since it is bound to at least a part of the linker DNA (36) and is known to be the most variable of all his tones. At the same time some data exist suggesting that also the 482 Nucleic Acids Research Fig* 5« Nondenaturating polyacrylamide gel el ectrophoresis of DNA fragments from micrococcal nuclease digests (36 min digestion;, (a) sea urchin sperm, (b) starfish sperm, (c) rat liver. Fig. 6. Comparison of micrococcal and DNase 1 digests on 10% polyacrylamide-urea gel. Denaturated DNA fragments from (a-c) micrococcal nuclease digests of nuclei; (a) sea urchin sperm, (b) rat liver, (c) starfish sperm; (d) DNase I digest of rat liver nuclei. According to corrected data (2 7) the bands are multiples of 10.4 bases. The band numbers were estimated as in (17). Nucleic Acids Research histories of intermediate variability, H2A and H2B, could by their N-terminal parts interact with the linker DNA and define together with H1, the linker length (8-10). If so, pronounced changes in these proteins would have an effect on the linker. Sea urchin sperm H2B is substantially different from its counterparts from other sources (12-15). H2B subtractions of sea urchin sperm have mol. w. markedly higher than that of starfish sperm H2B. The amino acid composition of sea urchin sperm H2B is characterized by a very high arginine content. The primary sequence analysis of six H2B subtractions from sperm of three sea urchin species has shown that the increase in mol. w. is due to the elongation of the N-terminal regions (12). These extensions contain basic repetitive pentapeptides Pro-Thr-Lys-Arg-Ser or Pro-Arg-Lys-Gly-Ser and bear some resemblance to protamines (28). To find out how the alteration in N-terminal part of the histone H2B from sea urchin sperm could be reflected in chromatin structure we compared using micrococcal nuclease some parameters of sea urchin and starfish sperm chromatin. Starfish sperm cells have been chosen for such analysis since all the histones they contain, for exeption of histone H2B, seemed to be very similar as judged from their amino acid compositions and electrophoretic behavior in two systems. To verify the above we considered also the data available on the primary structures of these proteins. Histones H3 and H4 are evolutionary highly stable. Only limited changes have been round in variable N-terminal part of sea urchin sperm YLtLk (23). The very important point is histone H1 since it is one of the proteins that according to current hypothesis might be responsible for variation of linker DNA length. Electrophoretic data show that histone H1 from sea urchin and starfish sperm have very similar (if not identical) mol. w. and charge (14, 15). Amino acid compositions of the both are also alike, the peculiarities that distinguish them from somatic H1 being of the same character. Moreover, from the vast body of the data on primary structures of histone H1 family (including sea urchin sperm H1), Von Holt et al. (12) concluded that H1 variants isolated from comparable cells show high degree of homology. 484 Nucleic Acids Research Contrary to the histone H1, the histone H2B from starfish sperm is approximately 20 amino acid residues shorter than that from sea urchin sperm and does not contain the specific N-terminal domain that has been found in sea urchin sperm H2B (13)* In primary structure and amino acid composition starfish sperm H2B resembles that from somatic cells. Thus, the only protein in which sea urchin and starfish sperm nucleosomes significantly differ is histone H2B. At the same time the linker DNA from sea urchin sperm nucleosome is about 13 bp longer than that from starfish sperm. On the basis of the data presented in this paper it can be suggested that the increase in linker DNA length of sea urchin sperm nucleosome might be due to ohanges in the N-terminal region of histone H2B: its elongation and enrichment by basic amino acid residues with a high affinity to DNA. The structural homologies between the reiterated pentapeptides present in N-terminal part of H2B and the protamines suggest that this region can be involved in condensing the genetic material (29). In this sense the basic N-terminal region of sea urchin sperm H2B may play a role similar to that of histone H1. The influence of the basic N-terminal region of H2B on the linker length can be obviously exerted if this region interacts with the linker DNA. A suggestion that the basic regions of H2B and H2A have binding sites outside the core particle has also been made on the basia of PME data (9)» In addition, the analysis of Serratia endonuclease digest of trypsin-treated chromatin also demonstrates a possibility of interaction between N-terminal regions of core histones and the linker DNA (10). It should be noted that histones H2A and H2B with markedly changed electrophoretic properties were found in plants (30). The repeat DNA length in plant chromatin is however about 200 bp which is characteristic of the cells with the "usual" set of core histones. (31). This apparent contradiction to our conclusion may be explained if we take into consideration data on amino acid composition and partial amino acid sequence of plant histones (32). Like sea urchin sperm histone H2B, pea histones H2B and H2A both contain evolutionary conserved middle and carboxyterminal region, whereas their N-terminal regions are vari485 Nucleic Acids Research able. However changes in N-terminal regions in pea and aea urchin histones are of different character. The basicity and arginine content of pea H2A and H2B are virtually the same or even lower than these values in their calf thjmus counterparts (32^. Therefore, these data are not inconsistent with our suggestion that the specific changes in H2B molecule, namely those which are connected with the affinity of the N-terminal region to DNA, influence the linker DNA length. Our findings support the assumption that conservation in length of core DNA might be due to the invariability of histones H3 and H4 and conserved hydrophobic central and C-terminal regions of nistone H2B (and probably H2A) responsible for histone interactions (33) while the linker DNA changes would be defined by evolutionary variable histone H1 and N-terminal regions of histone H2B at least. ACKNOWLEDGEMENT We thank Dr. E.Ch. Ibragimov for amino acid analysis. REFERENCES 1 2 3 4 5 6 7 8 9 10 11 12 13 486 Kornberg, R.D. (1977) Ann. Rev. Biochem. 46, 931-954 Chambon, P. (1978) Cold Spring Harbor Symp. on Quant.Biol. v.42, 1211-1236 Shaw, B.R., Hermann, T.M., Kovacic, R.T., Beaudreau, G.C. and Van Holde, K.E. (1976) Proc. Nat. Acad. S c i . 73. 505-509 Simpson, R.T. and Whit lock, J . P . (1976) Cell 9, 34-7-353 Varshavsky, A . J . , Bakayev, V.V. and Georgiev, G.P. (1976) Nucl. Acids Res. 3, 4-7-492 Noll, M. and Kornberg, R.D. (1977) J. Mol. Biol. 109, 393404 Morris, N.R. (1976) Cell 9, 627-632 Oudet, P . , Germond, J . E . , Bellard, M., Spadafora, C. and Chambon, P. (1977) Philos. Trans. R. Soc. Lond. Cary, P.D., Moss, T. and Bradbury, E.M. (1978) Eur. J . Biocham. 89, 475-482 Pospelov, V.A., Svetlikova, S.B. and Vorob'ev, V . I . (1979) FEBS L e t t . 99, 123-128 Panyim, S . , Bilek, D. and Chalkley, R. (1971) J . Biol. Chem. 246, 4206-4215 Von Holt, C , Strickland, W.N., Brandt, V7.F. and S t r i c k land, M.S. (1979) FEBS L e t t . 100, 201-217 S t r i c k l a n d , M.S., Strickland W.N. and Von Holt, C. (19«0) Eur. J . Biochem. 106, 541-548 Nucleic Acids Research 14 15 16 17 18 ly 20 21 22 23 24 25 26 27 28 29 30 31 32 33 Zalenskaya, I.A. and Zalensky, A.O. (1980) Comp. Biochem. Physiol. 65B, 369-373 Zalenskaya, I.A., Zalenskaya E.O. and Zalensky, A.O. Ibid. 65B, 375-378 Oliver, D., Sommer, K.R., Panyim, S., Spiker, St. and Chalkley, B. (1972) Biochem. J. 129, 34-9-353 Noll, M. (1974) Nucl. Acids Ees. 1, 1573-1578 Pospelov, V.A., Svetlikova, B.B. and Vorob'ev, V.I. (1977) Nucl. Acids Res. 4, 3267-3279 Laemmli, U.K. (1970) Nature 227, 680-685 Kinkade, J.M. and Cole, E.D. (1966) J. Biol. Chem. 241, 5798-5805 Wouters-Tyrou, D., S a u t i e r e , P. and Biserte, G. (1974) Biochim. Biophys. Acta 342, 360-366 Vanhoutte-Durand, G., Mizon, J . , Sautiere, P. and B i s e r t e , G. (1977) Gomp. Biochem. Physiol. 57b, 121-126 Sautiere, P . , Tyrou, D., Laine, B., Mizon, J . , Ruffin, P. and Biserte, G. (1974) Eur. J . Biochem, 4 1 , 563-576 Iwai, K., Ishikawa, K. and Hayashi, H. (1970) Nature 226, 1056-1058 Spadafora, C , Bellard, M., Oompton, J.L. and Chambon, P. (1976) PEBS L e t t . 69, 281-285 Keichline, L.D. and Wassarman, P.M. (1979) Biochemistry 18, 214-219 L u t t e r , L.C. Nucl. Acids. Res. (1979) 6, 41-56 Strickland, M., Strickland, W., Brandt, W.F., Von Holt, C , Wittmann-Liebold, B. and Lehmann, A. (1978) Bur. J . Biochem. 89, 443-452 Brandt, W.P. and Von Holt, G. (1978) Biochim. Biophys. Act a , 537. 177-181 Nadeau, P . , P a l l o t a , D. and Lafontaine, J.-G. (1974) Arch. Biochem. Biophys. 161, 171-177 Chea, K.S.E., Osborn, D.J. (1977) Biochem. J . 163, 141-144 Hayashi, H., Iwai, K., Johnson, J.D. and Bonner, J . (1977) J . Biochem. 82, 503-510 Spiker, S . , Isenberg, I . (1977) Biochemistry 16, 1819-1827 487 Nucleic Acids Research