* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Chapter 10 Gene Mutation: Origins and Repair Processes
Comparative genomic hybridization wikipedia , lookup
Genome evolution wikipedia , lookup
Holliday junction wikipedia , lookup
Genetic code wikipedia , lookup
Agarose gel electrophoresis wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Maurice Wilkins wikipedia , lookup
Community fingerprinting wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Molecular cloning wikipedia , lookup
Biosynthesis wikipedia , lookup
Non-coding DNA wikipedia , lookup
DNA supercoil wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Molecular evolution wikipedia , lookup
Deoxyribozyme wikipedia , lookup
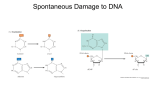
Chapter 10 Gene Mutation: Origins and Repair Processes Mutation is the process wherby genes change from one allelic form to another. Genetic analysis would not be possible without mutants. Gene mutations at the molecular level Type of mutation Examples Forward mutations at DNA level Transition Purin replaced by different purin, or pyrimidine replaced by a different pyrimidine: AT ⇒ GC; CG ⇒ TA Transversion Purine replaced by a pyrimidine, or pyrimidine replaced by a purine: AT ⇒ CG; CG ⇒ GC Forward mutations at protein level Silent mutation Triplets code for same amino acid: AGG ⇒ CGG (Arg) Synonymous missence mutation Codon specifies a chemically similar amino acid Nonsynonymous missence mutation Codon specifies a chemically dissimilar amino acid Type of mutation Frameshift mutation Reverse mutations Exact reversion Equivalent reversion Examples Any addition or deletion of base pairs that is not a multiple of 3. forward AAA(Lys) Wild-type UCC(Ser) Wild-type CGC(Arg) Wild-type ⇒ GAA(Glu) Mutant forward ⇒ UCG(Cys) Mutant forward ⇒ CCC(Pro) Mutant reverse AAA(Lys) Wild-type ⇒ reverse ⇒ AGC(Ser) Wild-type reverse ⇒ CAC(His) Pseudo wild-type Intragenic suppressor mutation For example, compensation of a frameshift mutation by a second mutation Extragenic suppressor mutation Nonsense suppressors, missense suppressors, frameshift suppressors, physiological suppressors Consequences of point mutations Salvatoria Luria and Max Delbrück's fluctuation test Hypothesis: random mutation or directed physiological change (1943) The frequency of mutations can be enhanced by mutagend Mechanism of point-mutation induction a) Base replacement by base analogs All bases can exist in one of several forms, called tautomers, which are isomers that differ in the positions of their atoms and in the bonds between the atoms. Mutation by tautomeric shifts Alternative pairing possibilities for 5-bromouracil. 5-BU is an analog of thymine that can be incorporated into DNA. 5-BU switches relatively frequently to the ionized form. Mechanisms of mutation induction b) Base alteration Some mutagens alter a base, causing specific mispairing: O C2H5 O S CH3 O Ethyl methanesulfonate Mechanisms of mutation induction c) intercalating agents Intercalating agents inhibit DNA synthesis and cause small deletions or insertions Mechanisms of mutation induction d) Base damage Many mutagens damage bases with the consequence that no specific base pairing is possible. Examples: UV light, Aflatoxin B1. The SOS system DNA polymerase stops at a noncoding lesion, generating ssDNA that attracts SsB and RecA which forms filaments. These filaments stimulate the expression of UmuD. UmuD is cleaved by RecA to yield UmuD' and UmuC which permit the DNA polymerase to proceed with DNA synthesis. Mutations are caused, because the inserted bases have a high error frequency. UV light generates pyrimidine photo dimers. A cyclobutan and a 6-4 photoproduct can be formed. These dimers interfere with normal base pairing. Ames test Cytochrom P450 monooxygenases in the liver convert certain metabolites by hydroxylation into mutagens. Ames test showing the mutagenicity of aflatoxin. Salmonella strain TA100 is sensitive to reversion through base substitution. Strains TA1538, TA1535 are sensitive to reversion by frameshift mutation. Binding of metabolically activated aflatocin Spontaneous Mutation Spontaneous lesions are caused by deamination or depurination. DNA replication errors can also lead to mutations. Deamination of 5-methlyctosin generates a "normal" base Damage products formed attack by oxygen radicals dR: deoxyribose Biological repair mechanisms 1. 2. 3. 4. 5. Photolyase Excision repair Nucleotide exision repair Mismatch repair Post-replication repair Photolyases catalyse the repair of pyrimidine dimers. The enzyme uses energy from visible light to break the carboncarbon bonds that join adjacent pyrimidine residues after UV irradiation. The base excision repair pathway. In this example, a uracil that was formed by deamination of cytosin is removed from the sugar-phosphate backbone by DNA glycosylase. These enzymes create apurinic or apyrimidinic (AP) sites. AP endonuclease recognizes these AP sites and cleaves the DNA strand. The remaining deoxyribose molecule is removed by deoxyribose phosphodiesterase and the gap is filled. Nucleotide excision repair (of a pyrimidine dimer). The DNA replication machinery is used to remove distortions in the DNA double helix. Damaged DNA is first recognized by endonucleases and cleaved. The oligonucleotide is then exised after helicase unwindes the DNA at this site. Eukaryotic nucleotide excision repair in the context of chromatin. Highly conserved among eukaryotes. About 25 proteins are needed for the ordered process of damage recognition, excision of damaged DNA and repair synthesis. For repair, about 100bp of nucleosome free DNA are required. Nucleotide excision repair (NER) proteins are mobile. After DNA damage, NER proteins assemble transiently at the site of DNA lesion. One repair event needs ca. 4 minutes. After repair, the chromation assembly factor CAF-1 restores the chromatin structure. Xeroderma pigmentosum, a human disorder, results from a defect in any one of eight genes involved in nucleotide excision repair. Mismatch repair in E. coli. Mismatch repair corrects errors made during DNA replication. A complex of three proteins (MutH, MutL, MutS) recognizes mismatched bases introduced by DNA polymerase into the newely synthesized, unmethylated DNA strand. MutH cleaves the new strand opposite the methylated site and the damaged DNA is then removed. Post-replication repair. Extensive DNA damage can be repaired by homologous recombination. An unrepaired lesion can be bypassed by DNA polymerase. The resulting gap is then repaired by recombination with the undamaged parental strand. Mutational analysis 1. Somatic versus germinal mutation 2. Mutant sectors 3. Mutant phenotypes Somatic versus germinal mutation Mutant sectors Summary