* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Bacterial Metabolism
Biochemical cascade wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Radical (chemistry) wikipedia , lookup
Proteolysis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Biosynthesis wikipedia , lookup
Phosphorylation wikipedia , lookup
Basal metabolic rate wikipedia , lookup
Photosynthesis wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Electron transport chain wikipedia , lookup
Metalloprotein wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Microbial metabolism wikipedia , lookup
Citric acid cycle wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
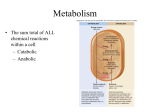
Topic 11 (ch8) Microbial Metabolism Topics – Metabolism – Energy – Pathways – Biosynthesis 1 Metabolism • Catabolism • Anabolism • Enzymes 2 Metabolic balancing act Catabolism Enzymes involved in breakdown of complex organic molecules to extract energy and form simpler end products Anabolism Enzymes involved in the use of energy from catabolism to synthesize macromolecules and cell structures from precursors (simpler products) 3 1 Catabolism and anabolism – simple model 4 Enzymes – what are they? • • • • • • Function Structure Enzyme-substrate interaction Cofactors Action Regulation 5 Function • Catalysts for chemical reactions • Lower the energy of activation 6 2 What Affects Enzymes? 7 Unfolding Kills Enzymes 8 Enzymes Inhibited by Competition (more later…) 9 3 Enzyme are Made of… – Many protein enzymes are complete without additions – Apoenzymes are inactive if not bound to non-protein cofactors (inorganic ions or coenzymes) – Binding of apoenzyme and its cofactor(s) yields holoenzyme – Some are RNA molecules called ribozymes 10 Types of Enzyme (Based on Structure) • Simple enzyme – protein alone • Conjugated enzyme – protein and nonprotein • Three-dimensional features – Enable specificity • Active site or catalytic site 11 A Conjugated Protein Enzyme 12 4 Another Example of Conjugated Enzymes 13 Enzymes in 3D 14 Enzyme-substrate interactions • Substrates specifically bind to the active sites on the enzyme – “lock-and-key” – Induced fit • Once the reaction is complete, the product is released and the enzyme reused 15 5 Substrate Fit 16 Lock-and-key model, and induced fit 17 Cofactors • Bind to and activate the enzyme • Ex. Metallic cofactors – Iron, copper, magnesium • Coenzymes 18 6 Coenzyme • Transient carrier - alter a substrate by removing a chemical group from one substrate and adding it to another substrate • Ex. vitamins 19 Example: coenzyme transfers chemical groups between substrates 20 Action • • • • • Exoenzymes Endoenzymes Constitutive Induction or repression Types of reactions 21 7 Exoenzymes vs. Endoenzymes •Exoenzymes inactive inside the cell, active after release. •Endoenzymes remain in the cell and are active 22 Constitiutive and Regulated enzymes Constitutive enzymespresent in constant amounts Regulated enzymes are either induced or repressed. 23 Reaction types • Condensation (associated with anabolic reactions) • Hydrolysis (associated with catabolic reactions) • Transfer reactions 24 8 enzyme-catalyzed synthesis and hydrolysis reactions 25 Transfer reactions • Transfer of electrons from one substrate to another – Oxidation (add O2 to a compound with a loss of electrons, increase charge) – Reduction (add e- to a compound, decrease charge) • Oxidoreductase • Transfer of functional groups from one molecule to another – Transferases • Aminotransferases 26 How to remember re-dox? • LEO goes GER • Losing Electrons is Oxidation • Gaining Electrons is Reduction Image from http://library.thinkquest.org 27 9 Examples of oxidoreductase, transferase, and hydrolytic enzymes 28 Enzyme Regulation • Metabolic pathways • Direct control • Genetic control 29 Patterns of metabolism Different metabolic pathways are regulated by the enzymes that catalyze the reactions. 30 10 Two common direct control mechanismsCompetitive and Noncompetitive inhibitions 31 Another Control Feedback Inhibition 32 Genetic control • Repression • Induction 33 11 Energy • Cell energetics – Exergonic – Endergonic • Redox reaction (change in oxidation number) • Electron carrier • Adenosine Triphosphate (ATP) 34 Cell energy model (simplified) 35 Redox Reaction (again) • Reduction and oxidation reaction (LEO goes GER): – Oxidation: loss of electrons, or gain of oxygen, gives increase in oxidation number. – Reduction: gain of electrons, or loss of oxygen, gives decrease in oxidation number. • Electron carriers transfer electrons and hydrogens – Electron donor – Electron acceptor • Energy is also transferred and captured by the phosphate in form of ATP 36 12 Electron carriers • Coenzymes – Nicotinamide adenine dinucleotide (NAD) • Respiratory chain carriers – Cytochromes (protein) 37 NAD reduction simple annotated 38 Adenosine Triphosphate (ATP - $$!) • Temporary energy repository - energy storage! • Break phosphates bonds to release free energy • Three part molecule: – Nitrogen base – 5-carbon sugar (ribose) – Chain of phosphates 39 13 AMP grows to ATP, each stage higher in energy 40 Carbohydrates – Most Important • Many organisms oxidize carbohydrates as primary energy source for anabolic reactions • Glucose most common carbohydrate used • Glucose catabolized by two processes: – Cellular respiration – Fermentation 41 Phosphorylation of glucose by ATP ATP can phosphorylate an organic molecule like glucose during catabolism 42 14 ATP can be synthesized by substrate-level phosphorylation. 43 Various Pathways • Catabolism – Embden-Meyerhof-Parnas (EMP) pathway or glycolysis – Tricarboxylic acid cycle (TCA) – Respiratory chain • Aerobic • Anaerobic – Alternate pathways – Fermentation 44 Metabolism Summary of Glucose and Energy 45 15 Aerobic respiration (O2) Three stages: • Glycolysis • Tricarboxylic acid (TCA) • Electron transport 46 Basics of Glycolysis • In cytoplasm for most cells – Divided into three stages involving 10 total steps • Energy-investment stage • Lysis stage • Energy-conserving stage (cont. next slide) 47 Basics of Glycolysis • In cytoplasm for most cells • Oxidation of glucose • Phosphorylation of some intermediates (Uses two ATPs) • Splits a 6 carbon sugar into two 3 carbon molecules • Coenzyme NAD is reduced to NADH • Substrate-level-phosphorylation (Four ATPs are synthesized) 48 (cont. next slide) 16 Basic of Glycolysis (continued) • Water is generated • Net yield of 2 ATPs • Final intermediates are two Pyruvic acid molecules (notice that it require 2 different, parallel pathways to finally generate both pyruvic acid molecules…) 49 Overview 50 Glycolytic Steps: Metabolism of Glucose to 2 Pyruvic Acid molecules (pyruvate) 51 17 TCA cycle • Each pyruvic acid is processed to enter the TCA cycle (two complete cycles) • CO2 is generated • Coenzymes NAD and FAD are reduced to NADH and FADH2 • Net yield of two ATPs • Critical intermediates are synthesized 52 TCA Cycle Steps Each cycle handles one 3 carbon molecule. This means it takes two cycles to burn up one 6 carbon glucose molecule. 53 Electron transport • NADH and FADH2 donate electrons to the electron carriers • Membrane bound carriers transfer electrons (redox reactions) • The final electron acceptor completes the terminal step (ex. Oxygen) 54 18 Electron Transport (continued) Driven by: • Chemiosmosis • Proton motive force (PMF) 55 e- Transport 56 Chemiosmosis and the electron transport system, with oxidative phosphorylation e- transport and formation of a proton gradient 57 19 Electron Transport Chain Locations • Eukaryotes – mitochondria • Prokaryotes – Cytoplasmic membrane 58 ATP Yield: One glucose molecule, aerobic respiration 59 Anaerobic respiration (No O2) • Similar to aerobic respiration, except nitrate or nitrite is the final electron acceptor 60 20 Fermentation • Glycolysis only • NADH from glycolysis is used to reduce the organic products • Organic compounds as the final electron acceptors • ATP yields are small (per glucose molecule), compared to respiration • Must metabolize large amounts of glucose to produce equivalent respiratory ATPs 61 Types of fermenters • Facultative anaerobes – Fermentation in the absence of oxygen – Respiration in the presence of oxygen – Ex. Escherichia coli • Strict fermenters – No respiration – Ex. yeast 62 Why Fermentation? – Sometimes cells cannot completely oxidize glucose by cellular respiration – Cells require constant source of NAD+ • Cannot be obtained simply using glycolysis and Krebs cycle – Fermentation pathways provide cells with alternate source of NAD+ • Partial oxidation of sugar (or other metabolites) to release energy using an organic molecule from within the cell as final electron acceptor 63 21 Products of fermentation • Alcoholic fermentation • Acidic fermentation • Mixed acid fermentation 64 Fermentation of ethyl alcohol and lactic acid 65 Pyruvate mixed acid fermentation diverse products 66 22 Fermentation Products 67 Respiration comparison 68 Biosynthesis • Anabolism – Amphibolic ( ) – Gluconeogenesis – Beta oxidation – Amination – Transamination – Deamination – Macromolecules 69 23 Amphibolic • Integration of the catabolic and anabolic pathways • Intermediates serve multiple purposes 70 Overview of anabolic and catabolic relationships 71 Gluconeogenesis • Pyruvate (intermediate) converts to glucose • Occurs when the glucose supply is low 72 24 Gluconeogenesis 73 Beta oxidation • Metabolism of fatty acids into acetylCoA Krebs (TCA cycle) • 4 steps: – Oxidation by FAD – Hydation of C2=C3 bond – Oxidation by NAD+ – Thiol cleavage at C2-C3 Acetyl-CoA 74 Production and Conversion of Amino Acids: amination, transamination, deamination 75 25 Fitting Cellular Together 76 The Big Picture 77 Macromolecules • Cellular building blocks – Monosaccharides – Amino acids – Fatty acids – Nitrogen bases – Vitamins 78 26 Photosynthesis – free power! 6H2O + 6CO2 C6H12O6+ 6O2 2 independent reactions in the chloroplast 79 Photosynthesis (cont.) 80 27