* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Identification of chromosome intervals from 129 and C57BL/6 mouse
Gene desert wikipedia , lookup
Medical genetics wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Minimal genome wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Population genetics wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Gene expression profiling wikipedia , lookup
Biology and sexual orientation wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Genome evolution wikipedia , lookup
Pathogenomics wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Gene expression programming wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genomic imprinting wikipedia , lookup
Microevolution wikipedia , lookup
Designer baby wikipedia , lookup
History of genetic engineering wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Y chromosome wikipedia , lookup
Public health genomics wikipedia , lookup
Neocentromere wikipedia , lookup
X-inactivation wikipedia , lookup
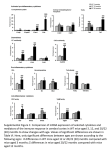
Genes and Immunity (2006) 7, 592–599 & 2006 Nature Publishing Group All rights reserved 1466-4879/06 $30.00 www.nature.com/gene ORIGINAL ARTICLE Identification of chromosome intervals from 129 and C57BL/6 mouse strains linked to the development of systemic lupus erythematosus Y Heidari1,3, AE Bygrave1,3, RJ Rigby1, KL Rose1, MJ Walport1,4, HT Cook2, TJ Vyse1 and M Botto1 Molecular Genetics and Rheumatology Section, Faculty of Medicine, Imperial College, Hammersmith Campus, London, UK and Department of Histopathology, Faculty of Medicine, Imperial College, Hammersmith Campus, London, UK 1 2 Systemic lupus erythematosus is an autoimmune disease in which complex interactions between genes and environmental factors determine the disease phenotype. We have shown that genes from the non-autoimmune strains 129 and C57BL/6 (B6), commonly used for generating gene-targeted animals, can induce a lupus-like disease. Here, we conducted a genome-wide scan analysis of a cohort of (129 B6)F2 C1q-deficient mice to identify loci outside the C1qa locus contributing to the autoimmune phenotype described in these mice. The results were then confirmed in a larger dataset obtained by combining the data from the C1q-deficient mice with data from previously reported wild-type mice. Both analyses showed that a 129-derived interval on distal chromosome 1 is strongly linked to autoantibody production. The B6 genome contributed to anti-nuclear autoantibody production with an interval on chromosome 3. Two regions were linked to glomerulonephritis: a 129 interval on proximal chromosome 7 and a B6 interval on chromosome 13. These findings demonstrate that interacting loci between 129 and B6 mice can cause the expression of an autoimmune phenotype in gene-targeted animals in the absence of any disrupted gene. They also indicate that some susceptibility genes can be inherited from the genome of non-autoimmune parental strains. Genes and Immunity (2006) 7, 592–599. doi:10.1038/sj.gene.6364335; published online 31 August 2006 Keywords: systemic lupus erythematosus; autoantibodies; rodent; gene-targeting Introduction Systemic lupus erythematosus (SLE) is a systemic autoimmune disease characterized by the production of autoantibodies against a variety of nuclear and cell surface antigens, resulting in immune-complex mediated damage to vascular, dermatological, renal, neurological and rheumatological tissues. The aetiology of SLE is complex, with a combination of multiple genes and environmental factors determining both susceptibility and disease phenotype. Spontaneous murine models of SLE, such as the New Zealand, BXSB and MRL mouse strains, have been widely used to dissect the complex genetic component of SLE. Comparison of the many linkage studies performed in these lupus-prone strains illustrates the complexity of this disease. Numerous disease susceptibility loci have been identified, with some loci, such as distal chromosome 1, mid to distal chromosome 4, proximal chromosome 7 and proximal chromosome 17 (the H2 complex) being linked to disease in different murine models.1 However, it is also Correspondence: Professor M Botto, Molecular Genetics and Rheumatology Section, Faculty of Medicine, Imperial College, Hammersmith Campus, Du Cane Road, London W12 0NN, UK. E-mail: [email protected] 3 These authors equally contributed to the work. 4 Current address: The Wellcome Trust, London, UK. Received 5 July 2006; accepted 31 July 2006; published online 31 August 2006 clear that several intervals are strain-specific, confirming the genetic complexity of the disease and indicating the presence of extensive heterogeneity in the genes contributing to SLE pathogenesis. More recently, gene-targeting technology has allowed researchers to investigate the impact of a single gene on murine physiology. There is, however, accumulating evidence that genetic factors other than the actual disrupted gene can influence the resulting phenotype of the knockout mouse. In this regard, it is of note that the majority of the gene-targeted strains are initially developed on a hybrid genetic background between 129 and C57BL/6 (B6) mice, which has been shown to be spontaneously predisposed to development of humoral autoimmunity with low levels of glomerulonephritis.2–5 The initial knockout strain is then usually backcrossed onto B6 in order to remove as much 129 genome as possible. However, despite 10 or more generations of backcrossing, a considerable 129-derived genome interval, flanking the targeted gene, will remain. We have previously shown that a number of genetic loci, derived from both 129 and B6, are linked to the development of disease in the (129 B6) mice, with the most statistically significant of these being a 129-derived interval on distal chromosome 1 linked to increased autoantibody production.6 This region has been consistently linked to autoimmune traits in a number of lupus-prone strains, including NZB (Nba2),7,8 NZM2410 (Sle1)9 and BXSB (Bxs3).10,11 A congenic strain, comprising the 129-derived locus on distal chromosome 1 on a B6 background, a Autoimmunity in 129 and C57BL/6 mice Y Heidari et al combination commonly created by backcrossing onto B6 a knockout strain in which the gene located in that region has been inactivated in 129 embryonic stem cells, also developed an autoimmune phenotype. The humoral autoimmunity in this congenic strain was indistinguishable to that observed in a mouse carrying a deletion of the Apcs gene, located within the lupus-linked genomic region on distal chromosome 1 and considered as a candidate gene for murine SLE.6 Therefore, backgroundderived genes can significantly contribute to the phenotype observed in knockout strains even when the mice have been extensively backcrossed onto the B6 strain, greatly complicating the interpretation of the phenotypic analysis of gene-targeted animals. The influence of background genes on the development or modification of spontaneous autoimmune disease is well known, especially with respect to the lpr and Yaa disease-susceptibility genes.12–14 Not surprisingly important effects of the genetic background on the expression of autoimmunity have also been reported in gene-targeted mice.5,15,16 For example, C1q deficiency, a condition that in humans is strongly associated with the development of a lupus-like disease,17 in mice appears to have a disease-accelerating effect only in lupus-prone strains, including the (129 B6) genetic background.3,16 Likewise, inactivation of the FcgRIIB gene results in the development of a lupus-like phenotype in the context of the B6 genomic background, but not the BALB/c genomic background.15 In this example, it is of note that two recessive B6-derived loci outside the targeted region were linked to the development of the disease phenotype.18 Thus, SLE exists as a complex-trait disorder in which specific combinations of susceptibility alleles are required for the expression of the full phenotype. In order to identify loci outside the C1qa locus that modify the autoimmunity observed in (129 B6).C1qa/ mice we carried out a linkage analysis to both autoantibody production and nephritis in a new cohort of (129 B6)F2.C1qa/ mice. In addition, we combined the data from this cohort with data from a previously published cohort of (129 B6)F2 wild-type mice,6 in order to expand the sample size and confirm our observations. We show here that a number of genetic loci outside the C1q locus on distal chromosome 4 are linked to autoimmunity, both confirming previously described loci and identifying novel regions in this strain combination. Results Mapping of loci predisposing to lupus in the (129 B6).C1qa/ mice The observation that 129.C1qa/ and B6.C1qa/ mice did not develop any autoimmune traits16 while the (129 B6).C1qa/ mice developed a lupus-like disease3 suggest that the disease-modifying loci may arise as a result of interaction between specific combinations of alleles inherited from both the 129 and B6 parental strains. In order to investigate the genetic contribution of 129 and B6 genes to the lupus-like disease observed in the (129 B6).C1qa/ mice, we generated a new cohort of (129 B6)F2.C1qa/ animals and monitored these for a year. In this cohort of C1q-deficient mice, anti-nuclear antibodies (ANA) were detected in 38% (median 0, range 0–10240), anti-chromatin antibodies (Abs) in 58% (median: 2.23, range 0–4.0), anti-double-stranded DNA (anti-dsDNA) Abs in 18% (median 0; range 0–2560) and anti-single-stranded DNA (anti-ssDNA) Abs in 85% of the mice. Histological evidence of glomerulonephritis (above grade I) was found in 32% of the mice. Interval mapping demonstrated significant linkage to ANA (LOD ¼ 4.0, P ¼ 9.9 105, Figure 1a), anti-dsDNA Abs (LOD ¼ 5.3, P ¼ 4.9 106, Figure 1b) and antissDNA Abs (LOD ¼ 5.5, P ¼ 3.1 106, Figure 1c) at a locus approximately 95 cM from the centromere of chromosome 1 in the (129 B6)F2.C1qa/ cohort. Antichromatin Abs were also linked to this chromosome 1 region, but at a more distal locus and with a lower level of significance (LOD ¼ 2.2, P ¼ 6.3 104, Figure 1d). All these loci were derived from 129. In addition to the linkage observed on distal chromosome 1, ANA titres were linked to a locus on mid-distal chromosome 3, albeit with a reduced degree of significance (LOD ¼ 2.3, P ¼ 5 104, Figure 2). This locus was derived from the B6 background. In this context, it is of note that QTL analysis of the mid-distal region of chromosome 4 could not be applied to the (129 B6)F2.C1qa/ mice as this region was of fixed 129 origin. Owing to the reduced variability in the glomerulonephritis (GN) data compared to the autoantibody titre data, the linkage to GN was determined using two methods – as a regular quantitative trait, and in an analysis of extremes, where only mice that were clearly negative or positive were included. Both these analyses showed that GN was linked to a 129-derived region of proximal chromosome 7 in the (129 B6)F2.C1qa/ cohort. The QTL analysis demonstrated suggestive linkage (LOD ¼ 2.2, P ¼ 6.3 104) to a 22 cM region between D7Mit246 (15 cM) and D7Mit30 (37 cM), and the analysis of extremes showed linkage to D7Mit230 at 26.3 cM, with a w2 of 8.84. 593 Confirmation of linkage analysis data in a combined cohort of (129 B6)F2.C1qa/ and (129 B6)F2 mice Overall the study of the (129 B6)F2.C1qa/ cohort confirmed a number of the 129 and B6 loci previously described as being associated with SLE traits, but failed to support others.6 In order to increase the power of our study, we repeated the QTL analysis in a larger data set, obtained by combining data from the (129 B6)F2.C1qa/ mice presented in this study with data from a previously reported (129 B6)F2 intercross,6 resulting in 297 female mice. In this analysis, the medial and distal regions of chromosome 4 were not included. The results from the IgG anti-ssDNA ELISA and IgG anti-chromatin ELISA assays were ranked, as described in the Materials and methods section, in order to compare the data from the two cohorts. The ranking method was verified by comparing the patterns of genome-wide linkage to both ranked and unranked ELISA data in a single cohort, and confirming that the linkage patterns were comparable in both position and magnitude of linkage. The combined cohort of (129 B6)F2.C1qa/ and (129 B6)F2 mice confirmed the linkage of ANA (LOD ¼ 6.3, P ¼ 5 107, Figure 3a), IgG anti-dsDNA Abs (LOD ¼ 7.6, P ¼ 2.5 108, Figure 3b), IgG antissDNA Abs (LOD ¼ 8.3, P ¼ 4.9 109, Figure 3c) and IgG anti-chromatin Abs (LOD ¼ 4.7, P ¼ 2 105, Genes and Immunity Autoimmunity in 129 and C57BL/6 mice Y Heidari et al 594 Figure 1 Interval maps showing QTLs on chromosome 1 with ANA (a), anti-dsDNA Abs (b), anti-ssDNA Abs (c) and anti-chromatin Abs (d) in (129 B6)F2.C1qa/ mice. Centimorgan positions were deduced by interval mapping, anchoring marker locations according to data from www.informatics.jax.org. Intermittent dashed lines indicate threshold over which linkage is considered suggestive; dashed lines indicate threshold over which linkage is considered significant; dotted lines indicate threshold over which linkage is considered highly significant (see Materials and methods). LOD scores were generated with Map Manager QTb29. Figure 2 Linkage of ANA to a B6-derived region of Chromosome 3 in (129 B6)F2.C1qa/ mice. Threshold for suggestive linkage (intermittent dashed line), as determined by 1000 cross- and traitspecific permutation tests, is indicated (LOD ¼ 2.3, P ¼ 5 104). Figure 3d) to a locus on distal chromosome 1 with a peak of around 95 cM. All of the above disease traits were linked to the distal chromosome 1 region with a higher degree of significance in the combined than the Genes and Immunity individual cohorts. It is of note that the linkage to IgG anti-chromatin Abs mapped to a more distal region of chromosome 1 than the other autoantibodies in the (129 B6)F2.C1qa/ cohort, whereas in the combined cohort analysis it mapped to the same locus as the other autoantibodies (B95 cM from the centromere). Akin to the linkage on chromosome 1, the ANA linkage to middistal chromosome 3 was observed in the combined cohort of (129 B6)F2.C1qa/ and (129 B6)F2 mice. ANA titre was linked to a locus of around 50 cM from the centromere, again with an increased degree of significance in the combined cohort (LOD ¼ 8.0, P ¼ 9 109, Figure 4). To investigate any potential interactions between the B6-derived gene(s) on chromosome 3 and the 129derived gene(s) on chromosome 1, we grouped the mice accordingly to their genotype at these two loci and compared ANA titres. We selected a marker at the peak of the linkage to ANA ((D1Mit206 (95.8 cM) on chromosome 1 and D3Mit103 (51.1 cM) on chromosome 3) to define the genotype of the mice. As illustrated in Figure 5, ANA levels were significantly higher in the mice carrying two B6 alleles on chromosome 3 in combination with one or two 129 alleles on chromosome 1 compared to all the other genetic combinations. Of note the ANA Autoimmunity in 129 and C57BL/6 mice Y Heidari et al 595 Figure 3 Interval maps showing QTLs on chromosome 1 with ANA (a), anti-dsDNA Abs (b), anti-ssDNA Abs (c) and anti-chromatin Abs (d) in a combined analyses of (129 B6)F2.C1qa/ and wild-type (129 B6)F2 mice. Centimorgan positions were deduced by interval mapping, anchoring marker locations according to data from www.informatics.jax.org. Intermittent dashed lines indicate threshold over which linkage is considered suggestive; dashed lines indicate threshold over which linkage is considered significant; dotted lines indicate threshold over which linkage is considered highly significant. Thresholds were determined by 1000 cross- and trait-specific permutation tests. Figure 4 Linkage of ANA to a B6-derived region of Chromosome 3 in a combined analyses of (129 B6)F2.C1qa/ and wild-type (129 B6)F2 mice. Centimorgan positions were deduced by interval mapping, anchoring marker locations according to data from www.informatics.jax.org. Intermittent dashed lines indicate threshold over which linkage is considered suggestive; dashed lines indicate threshold over which linkage is considered significant; dotted lines indicate threshold over which linkage is considered highly significant (LOD ¼ 8.0, P ¼ 9 109). titres were not significantly different between the animals homozygous or heterozygous for the 129derived segment on chromosome 1, when in combination with B6 homozygosity on chromosome 3 suggesting that the presence of a single 129-derived allele on chromosome 1 was sufficient to drive loss of tolerance to nuclear antigens. This analysis provided further support to the hypothesis that the B6-derived loci on chromosome 3 tend to operate in a recessive manner while the 129-derived loci on chromosome 1 operate in a dominant fashion. As in the (129 B6)F2.C1qa/ cohort, the linkage analysis of GN in the combined cohort was carried out using both a quantitative trait analysis and analysis of extremes. Using the quantitative trait analysis method, linkage of GN to a 129-derived locus on proximal chromosome 7 was confirmed with an increased significance (LOD ¼ 3.8, P ¼ 1.6 105, Figure 6a). This was confirmed by the analysis of extremes which showed a linkage to D7Mit230 (26.3 cM), with a w2 of 11.72, P ¼ 0.0028 (Table 1). Interestingly, a further linkage to GN, this time derived from a B6 locus, was observed in the combined cohort. Located on proximal chromosome 13, this locus reached the cutoff for suggestive linkage Genes and Immunity Autoimmunity in 129 and C57BL/6 mice Y Heidari et al 596 Figure 5 ANA titres of various chromosome 1 and chromosome 3 genotype combinations in a pooled cohort of (129 B6)F2.C1qa/ and wild-type (129 B6)F2 mice. Small symbols represent one mouse; large symbols a variable number of animals as indicated in parentheses. The genotype was established using a single marker at the peak linkages on chromosome 1 (D1Mit206) and chromosome 3 (D3Mit103). The mice carrying two B6-derived alleles on chromosome 3 and one or two 129-derived alleles on chromosome 1 had significantly higher levels of ANA compared to all the other genetic combinations (Po0.001). The most relevant comparisons are shown. One way ANOVA with Bonferonni’s multiple comparison tests were applied. (LOD ¼ 2.4, P ¼ 4 104, Figure 6b) in the combined cohort, but not in the individual (129 B6)F2.C1qa/ or (129 B6)F2 cohorts. Like the locus on chromosome 7, this linkage was also observed in an analysis of extremes, with GN linked to D13Mit117 at B19 cM, with a w2 of 15.3, P ¼ 4.77 104 (Table 1). Discussion Multiple genetic loci are known to contribute to the development and pathogenesis of SLE in mice and in humans. In this study, we have provided further evidence that epistatic interactions between 129 and B6 mice, even though autoimmunity has not been reported in either of the strains, can lead to a spontaneous lupuslike phenotype. In particular, we confirmed that a 129derived region of chromosome 1, when expressed in context of the B6 genome, is strongly linked to autoantibody production. Consistent with this, the B6 genome contributed to the autoimmune phenotype with an interval on chromosome 3, displaying a highly significant linkage to anti-nuclear autoantibodies. Interestingly glomerulonephritis was linked to different chromosomes: a 129 region on proximal chromosome 7 and a B6 interval on chromosome 13 (Figure 7). A genome-wide scan of the (129 B6)F2.C1qa/ mice demonstrated linkage of lupus serological markers (ANA, anti-dsDNA Abs, anti-ssDNA Abs and antichromatin Abs) to a 129-derived locus on distal chromosome 1 – recently named Sle 16 (http://www.informatics. jax.org). This is in agreement with our previous study,6 in which SLE traits were mapped in (129 C57BL/6)F2 wild-type and (129 C57BL/6)F2.Apcs/ mice. The linkages were markedly increased when we enlarged the sample size by combining data from the Genes and Immunity Figure 6 Interval maps showing QTLs on chromosome 7 (a) and chromosome 13 (b) with GN in a combined analysis of (129 B6)F2.C1qa/ and wild-type (129 B6)F2 mice. Centimorgan positions were deduced by interval mapping, anchoring marker locations according to data from www.informatics.jax.org. Intermittent dashed lines indicate threshold over which linkage is considered suggestive; dashed lines indicate threshold over which linkage is considered significant. (LOD ¼ 3.8, P ¼ 1.6 105 and LOD ¼ 2.4, P ¼ 4 104, respectively). Table 1 Linkage of chromosomes 7 and 13 to GN in the combined cohort of (129 B6)F2.C1qa/ and wild-type (129 B6)F2 mice Marker Position (cM) Origin w2 P D7Mit178 D7Mit246 D7Mit158 D7Mit230 D7Mit30 D7Mit253 0.5 15.0 23.0 26.3 37.0 52.8 129 129 129 129 129 129 4.14 11.04 10.05 11.72 7.57 6.13 1.26E-01 4.01E-03 6.58E-03 2.86E-03 2.27E-02 4.66E-02 D13Mit135 D13Mit117 D13Mit64 D13Mit248 D13Mit193 D13Mit30 D13Mit262 10.0 19.0 30.0 34.0 43.0 52.0 68.0 B6 B6 B6 B6 B6 B6 B6 12.53 15.29 5.81 3.54 2.51 0.63 1.61 1.90E-03 4.77E-04 5.47E-02 1.70E-01 2.85E-01 7.31E-01 4.48E-01 Centimorgan positions were deduced by interval mapping, anchoring marker locations according to data from www.informatics.jax.org. w2-values are calculated with a standard (2 2) contingency table with 2 degree of freedom. Suggestive linkages, defined as described in the material and method section, are underlined. Autoimmunity in 129 and C57BL/6 mice Y Heidari et al 597 1 3 7 Sles3* Bxs5 1,2 1 Sle1 1,2,4 Nba2 1,2 Bxs3 1 Lbw7 ANA ANA anti-ssDNA anti-dsDNA anti-chromatin 13 1,2 1,2 Sle3 3 Lbw5 2 Nba3 GN GN 1,2 Yaa1 Figure 7 A summary of the lupus susceptibility loci mapped in this study on chromosomes 1, 3, 7 and 13. Regions previously linked to SLE in the other lupus models are also indicated. Linkage to SLE traits from this study are shown as open boxes ¼ 129, filled boxes ¼ B6. 1ANA (any), 2GN, 3mortality, 4gp70/gp70 immune complexes, *disease suppressor locus. (129 B6)F2.C1qa/ mice with data from a previously reported (129 B6)F2 intercross, indicating that this 129 region is the main locus capable of initiating the humoral autoimmune response to nuclear antigens upon interaction with the B6 genome. The distal chromosome 1 region is associated with a number of potent SLE susceptibility loci including Sle1,9 Nba219 and Bxs3,10 all of which are strongly associated with the production of autoantibodies to nuclear antigens. The significance of this chromosome 1 locus has also been confirmed by several congenic dissection analyses.1,20 Furthermore, recent genomic characterization of the Sle1b locus (located between 171.8 and 173.1 Mbp) has identified a highly polymorphic cluster of Slam/Cd2 family genes, encoding key regulators of lymphocyte function, as the strongest candidate genes for mediating the Sle1b autoimmune phenotype.21 The autoimmune-associated haplotype of the lupus-prone NZM2410 (NZW) strain, named the Slam/Cd2 haplotype 2, is also present in the 129/SvJ mice,21 indicating that these two strains may share the same pathways leading to loss of peripheral tolerance when in combination with one or more polymorphic genes in the B6 genome. These observations give compelling evidence for the presence of a 129 locus influencing systemic autoimmunity on telomeric chromosome 1 as revealed in the context of B6 genome. Whether this 129 lupus locus can contribute to the development of autoimmunity on other genetic backgrounds is still unknown. In common with the distal chromosome 1 region, numerous studies have demonstrated linkage of SLE traits to proximal chromosome 7. In this study, we confirm that there is a 129-derived locus on proximal chromosome 7 linked to GN.6 Furthermore, this locus colocalizes with a number of loci from various lupusprone mouse strains, including Sle3 (NZM2401 locus linked to GN and ANA);20,22 Lbw5 (NZW locus linked to mortality),23 Lmb3 (MRL locus linked to splenomegaly, lymphadenopathy, anti-dsDNA Abs),24 Nba3 (NZB locus linked to GN)25 and Nba5.26 As this is an analogous situation to that on distal chromosome 1, a similar haplotype-based candidate gene identification strategy could be applied, especially as murine single nucleotide polymorphism (SNP) databases continue to increase in both SNP density and the number of typed strains. The presence of a 129-derived lupus locus on proximal chromosome 7 also has impact on the phenotype of any targeted gene in this region – it is conceivable that the 129 gene(s) that predispose to GN in the (129 B6)F2 strain could modify or augment the GN observed in a knockout strain. Recently a knockout of the Bcl-2 associated protein (Bax) gene, located on proximal chromosome 7, was reported to have GN as one of its resulting phenotypes.27 In light of the data presented herein, one cannot exclude the possibility that the observed GN phenotype may have been due to the surrounding 129 genome, and not the inactive Bax gene. A previously described locus of B6 origin on chromosome 3,6 was here confirmed to be linked to ANA production in both the (129 B6)F2.C1qa/ mice and combined cohorts. Unlike the loci on chromosomes 1 and 7, this locus has not been consistently linked to lupus disease traits, the SLE-associated loci Sles3 and Bxs5 being proximal (between 20 and 40 cM) to our observed linkage peak. However, they intriguingly both involve non-autoimmune strains; in the case of Sles3, NZWxC57BL/6 heterozygosity,1 and in the case of Bxs5, C57BL/10 homozygosity.11 The confirmation of a B6-derived locus on chromosome 3 that modifies ANA titres (in combination with 129-derived genes) highlights the likelihood of epistatic interactions influencing the phenotype of a knockout strain, in spite of extensive backcrossing to B6. The possibility of B6 genes influencing the phenotype of a knockout mouse was further underlined by the observation of B6-derived linkage to GN on proximal chromosome 13. This linkage was only observed in the combined analysis of (129 B6)F2.C1qa/ and (129 B6)F2 mice, and had thus a less penetrant effect than the loci on chromosomes 1, 3 and 7. Nonetheless, such minor loci may together have a significant cumulative influence on the phenotype of a knockout strain bred onto the B6 background. Hence, it is important to identify both the genomic position and the resulting phenotypes of these loci. In this context, it is of note that we had previously reported a B6-derived suggestive linkage to anti-dsDNA Abs on the mid-distal region of chromosome 4.6 However, this observation could not be validated in the current study as this region was of fixed 129-origin in the (129 B6)F2.C1qa/ mice. Hence, we cannot exclude the potential effect of a minor B6-derived locus on distal chromosome 4. In conclusion, we have identified linkage of lupus disease traits to loci present on chromosomes 1, 3, 13 and 7 in the 129 B6 mice (Figure 7). As this is the strain combination of choice for gene-targeted mice, knowledge Genes and Immunity Autoimmunity in 129 and C57BL/6 mice Y Heidari et al 598 of the influence of parental genetic loci on the disease phenotype is critical if we are to avoid attributing false positive phenotypes to knockout strains. Materials and methods Mice The C1q-deficient mice, C1qa/ were generated as previously reported3 and the (129 B6)F1.C1qa/ mice were generated by crossing 129.C1qa/ mice with B6.C1qa/ mice that had been backcrossed onto B6 for 10 generations. (129 B6)F2.C1qa/ were obtained by intercrossing the (129 B6)F1.C1qa/ mice. A total of 156 (129 B6)F2.C1qa/ female mice were produced and monitored for 1 year. The strain- and sex-matched wildtype (129 B6)F2 cohort was as previously described.6 Mice were maintained in specific pathogen-free conditions, and all procedures were in accordance with institutional guidelines. Serological analyses Serum was collected at 12 months of age and assayed for autoantibodies. IgG ANA and IgG anti-dsDNA Abs were measured by indirect immunofluorescence using Hep-2 cell and Crithidia luciliae slides (The Binding Site, Birmingham, UK) respectively.16 Serum samples were screened at a 1:80 (ANA) or 1:20 (anti-dsDNA Abs) dilution and the positive samples titrated to end point. IgG anti-ssDNA Abs and anti-chromatin Abs were measured by ELISA as described previously.28 For determination of anti-chromatin antibodies ELISA plates (Nunc-Immuno MaxiSorp, NUNC, Denmark) were coated with 50 ml of PBS Thimerosal (0.1 g/l) containing 0.5 mg/ml nucleohistones from calf thymus (Lorne laboratories Ltd, Reading, UK) at 41C overnight, then blocked with PBS 5% milk powder for 1 h at 371C. Sera diluted 1/100 in PBS 2% BSA 0.05% Tween20 were incubated for 1 h at 371C. For measuring anti-ssDNA Abs plates were coated with 50 ml of 10 mg/ml ssDNA (Sigma Chemical Co., Poole, UK) in sodium carbonate buffer pH 9.6 at 41C overnight and then blocked with 100 ml PBS 0.5% BSA. Samples were screened at a 1/100 dilution in PBS 2% BSA 0.05% Tween20 for 1 h at 371C with 50 ml/ well. Bound antibodies were detected with alkaline phosphatase (AP)-conjugated goat anti-mouse IgG (g-chain specific) (Sigma-Aldrich, Dorset, UK). The plates were developed using the substrate p-nitrophenyl phosphate (Sigma Chemical Co., Poole, UK). The OD of the reaction mixture at 405 nm wavelength was measured using an ELISA plate reader (Titertek Labsystems, Basingstoke, UK). Samples were tested in duplicate with a non-specific binding control and the results were expressed in arbitrary ELISA units (AEU) relative to serial dilutions of a standard positive sample derived from pooled autoimmune MRL/Mp.lpr/lpr serum. Serum samples were considered positive if above the mean 72s.d. of the blank. The intra-assay coefficient of variation was between 5 and 8%. Histological analysis All the mice were killed at 1 year of age. Kidney tissue was fixed in Bouin’s solution for at least 2 h, transferred into 70% ethanol, and then processed into paraffin. Sections were cut, mounted, stained with periodic acidGenes and Immunity Schiff reagent and scored for GN. Glomerular histology was graded as follows: grade 0 – normal, grade I – focal hypercellularity in 10–25% of the glomeruli, II – hypercellularity involving 450% of the glomerular tuft in 25–50% of glomeruli, grade III – hypercellularity involving 450% of the glomerular tuft in 50–75% of glomeruli, grade IV – glomerular hypercellularity in 475% or crescents in 425% of glomeruli. Histological analysis was performed in a blinded fashion and 50 glomeruli per section were analysed. Genotypic analysis Genotyping of the (129 B6)F2.C1qa/ cohort was carried out using polymorphic microsatellite markers, a standard polymerase chain reaction and either 4% MetaPhor agarose (Cambrex Bioscience Rockland, Rockland, ME, USA) or 16% polyacrylamide gels stained with ethidium bromide. The average marker density was one marker per 10 centimorgans (cM) across the autosomes, the positions and sequences of which were determined from the Mouse Genome Informatics (MGI) database (http://www.informatics.jax.org). The list of markers used is available on request. Statistical analyses All linkage analyses and interval mapping were conducted using MapManager QTb29 (ftp://mcbio.med.buffalo.edu/pub/MapMgr/).29 Marker maps were generated to determine the accuracy of genotyping, with all markers mapping to within 3 cM of the marker position in the MGI database. All centimorgans positions of markers referred to in this study are from the MGI database. Anti-ssDNA and anti-chromatin ELISA data were log transformed prior to linkage analysis as this resulted in a more normalized distribution. As the ELISA assays for these two autoantibodies in the two cohorts of mice had been carried on separate occasions and using a different MRL/Mp.lpr/lpr standard positive sample, for the combined analysis of the two sets of data the samples in each group were ranked using uncategorized (continuous) arbitrary values and the ranked values were subsequently used for the quantitative trait locus (QTL) analysis. Thresholds for suggestive, significant and highly significant linkages were determined using cohort- and trait-specific permutation tests, based on 1000 permutations of the data, in Map Manager QTb29.29 A logarithm of odds ratio (LOD) of X2.0 (Pp9.9 104), of X3.6 (Pp2.5 105) and of X5.7 (Pp2.0 106) was indicative of suggestive, significant and highly significant linkage, respectively, in the (129 B6)F2.C1qa/ cohort of mice. In the combined cohort of C1q-deficient and wild-type F2 mice, the threshold for suggestive, significant and highly significant linkages were LOD X2.1 (Pp7.9 104), X3.5 (Pp3.1 105) and X5.3 (Pp4.9 106), respectively. The calculated thresholds for suggestive, significant and highly significant linkages were similar across the different traits. Glomerulonephritis score was analysed as both a quantitative trait in Map Manager and in an analysis of extremes, in which mice with a GN score of II or above were considered as positive for glomerulonephritis, and mice with a GN score of 0 were considered negative. All mice with grade I GN were excluded. Linkage to Autoimmunity in 129 and C57BL/6 mice Y Heidari et al microsatellite markers was determined using a Chi square (w2) test in an Excel (2003) spreadsheet. w2-values of over 12.9 (P ¼ 0.0016) was considered to be indicative of suggestive linkage.30 Non-parametric data are presented as median, with range of values in parentheses unless otherwise stated. Statistics were calculated using GraphPad Prism version 3.0 (GraphPad Software, San Diego, CA, USA). One way ANOVA with Bonferonni;s multiple comparison tests were applied for analysis of multiple groups. Abbreviations Abs, antibodies; AEU, arbitrary ELISA units; ANA, antinuclear antibody; AP, alkaline phosphatase; antidsDNA, anti-double stranded DNA; anti-ssDNA, anti-single stranded DNA; B6, C57BL/6; GN, glomerulonephritis; QTL, quantitative trait locus; SLE, Systemic lupus erythematosus. Acknowledgements We thank Mrs Margarita Lewis for technical assistance with the processing of tissue for histological studies and the staff of the Biological Services Unit at our institution for the care of the animals involved in this study. This work was supported by the Wellcome Trust (Grant number 071467). References 1 Wakeland EK, Liu K, Graham RR, Behrens TW. Delineating the genetic basis of systemic lupus erythematosus. Immunity 2001; 15: 397–408. 2 Obata Y, Tanaka T, Stockert E, Good RA. Autoimmune and lymphoproliferative disease in (B6-GIX+ X 129)F1 mice: relation to naturally occurring antibodies against murine leukemia virus-related cell surface antigens. Proc Natl Acad Sci USA 1979; 76: 5289–5293. 3 Botto M, Dell’Agnola C, Bygrave AE, Thompson EM, Cook HT, Petry F et al. Homozygous C1q deficiency causes glomerulonephritis associated with multiple apoptotic bodies. Nat Genet 1998; 19: 56–59. 4 Bickerstaff MC, Botto M, Hutchinson WL, Herbert J, Tennent GA, Bybee A et al. Serum amyloid P component controls chromatin degradation and prevents antinuclear autoimmunity. Nat Med 1999; 5: 694–697. 5 Santiago-Raber ML, Lawson BR, Dummer W, Barnhouse M, Koundouris S, Wilson CB et al. Role of cyclin kinase inhibitor p21 in systemic autoimmunity. J Immunol 2001; 167: 4067–4074. 6 Bygrave AE, Rose KL, Cortes-Hernandez J, Warren J, Rigby RJ, Cook HT et al. Spontaneous autoimmunity in 129 and C57BL/ 6 mice-implications for autoimmunity described in genetargeted mice. PLoS Biol 2004; 2: E243. 7 Drake CG, Rozzo SJ, Hirschfeld HF, Smarnworawong NP, Palmer E, Kotzin BL. Analysis of the New Zealand Black contribution to lupus-like renal disease. Multiple genes that operate in a threshold manner. J Immunol 1995; 154: 2441–2447. 8 Vyse TJ, Rozzo SJ, Drake CG, Izui S, Kotzin BL. Control of multiple autoantibodies linked with a lupus nephritis susceptibility locus in New Zealand black mice. J Immunol 1997; 158: 5566–5574. 9 Morel L, Blenman KR, Croker BP, Wakeland EK. The major murine systemic lupus erythematosus susceptibility locus, Sle1, is a cluster of functionally related genes. Proc Natl Acad Sci USA 2001; 98: 1787–1792. 599 10 Hogarth MB, Slingsby JH, Allen PJ, Thompson EM, Chandler P, Davies KA et al. Multiple lupus susceptibility loci map to chromosome 1 in BXSB mice. J Immunol 1998; 161: 2753–2761. 11 Haywood ME, Hogarth MB, Slingsby JH, Rose SJ, Allen PJ, Thompson EM et al. Identification of intervals on chromosomes 1, 3, and 13 linked to the development of lupus in BXSB mice. Arthritis Rheum 2000; 43: 349–355. 12 Izui S, Kelley VE, Masuda K, Yoshida H, Roths JB, Murphy ED. Induction of various autoantibodies by mutant gene lpr in several strains of mice. J Immunol 1984; 133: 227–233. 13 Izui S, Higaki M, Morrow D, Merino R. The Y chromosome from autoimmune BXSB/MpJ mice induces a lupus-like syndrome in (NZW C57BL/6)F1 male mice, but not in C57BL/6 male mice. Eur J Immunol 1988; 18: 911–915. 14 Merino R, Shibata T, De Kossodo S, Izui S. Differential effect of the autoimmune Yaa and lpr genes on the acceleration of lupus-like syndrome in MRL/MpJ mice. Eur J Immunol 1989; 19: 2131–2137. 15 Bolland S, Ravetch JV. Spontaneous autoimmune disease in Fc(gamma)RIIB-deficient mice results from strain-specific epistasis. Immunity 2000; 13: 277–285. 16 Mitchell DA, Pickering MC, Warren J, Fossati-Jimack L, Cortes-Hernandez J, Cook HT et al. C1q deficiency and autoimmunity: the effects of genetic background on disease expression. J Immunol 2002; 168: 2538–2543. 17 Pickering MC, Botto M, Taylor PR, Lachmann PJ, Walport MJ. Systemic lupus erythematosus, complement deficiency, and apoptosis. Adv Immunol 2000; 76: 227–324. 18 Bolland S, Yim Y-S, Tus K, Wakeland EK, Ravetch JV. Genetic modifiers of systemic lupus erythematosus in FcgRIIB/ mice. J Exp Med 2002; 195: 1167–1174. 19 Rozzo SJ, Vyse TJ, Drake CG, Kotzin BL. Effect of genetic background on the contribution of New Zealand black loci to autoimmune lupus nephritis. Proc Natl Acad Sci USA 1996; 93: 15164–15168. 20 Morel L, Tian XH, Croker BP, Wakeland EK. Epistatic modifiers of autoimmunity in a murine model of lupus nephritis. Immunity 1999; 11: 131–139. 21 Wandstrat AE, Nguyen C, Limaye N, Chan AY, Subramanian S, Tian XH et al. Association of extensive polymorphisms in the SLAM/CD2 gene cluster with murine lupus. Immunity 2004; 21: 769–780. 22 Morel L, Rudofsky UH, Longmate JA, Schiffenbauer J, Wakeland EK. Polygenic control of susceptibility to murine systemic lupus erythematosus. Immunity 1994; 1: 219–229. 23 Kono DH, Burlingame RW, Owens DG, Kuramochi A, Balderas RS, Balomenos D et al. Lupus susceptibility loci in New Zealand mice. Proc Natl Acad Sci USA 1994; 91: 10168–10172. 24 Vidal S, Kono DH, Theofilopoulos AN. Loci predisposing to autoimmunity in MRL-Fas lpr and C57BL/6-Faslpr mice. J Clin Invest 1998; 101: 696–702. 25 Xie S, Chang SH, Sedrak P, Kaliyaperumal A, Datta SK, Mohan C. Dominant NZB contributions to lupus in the (SWR NZB)F1 model. Genes Immun 2002; 3 (Suppl 1): S13–S20. 26 Kikuchi S, Fossati-Jimack L, Moll T, Amano H, Amano E, Ida A et al. Differential role of three major New Zealand Blackderived loci linked with Yaa-induced murine lupus nephritis. J Immunol 2005; 174: 1111–1117. 27 Takeuchi O, Fisher J, Suh H, Harada H, Malynn BA, Korsmeyer SJ. Essential role of Bax, Bak in B cell homeostasis and prevention of autoimmune disease. Proc Natl Acad Sci USA 2005; 102: 11272–11277. 28 Burlingame RW, Rubin RL. Subnucleosome structures as substrates in enzyme-linked immunosorbent assays. J Immunol Methods 1990; 134: 187–199. 29 Manly KF, Olson JM. Overview of QTL mapping software and introduction to map manager QT. Mammlian Genome 1999; 10: 327–334. 30 Lander E, Kruglyak L. Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet 1995; 11: 241–247. Genes and Immunity