* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download CHAPTER 5: THE INHERITANCE OF SINGLE
Gene therapy wikipedia , lookup
Genetic drift wikipedia , lookup
Genome evolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Point mutation wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene desert wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene nomenclature wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genomic imprinting wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome (book) wikipedia , lookup
Y chromosome wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Gene expression programming wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
X-inactivation wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Neocentromere wikipedia , lookup
Dominance (genetics) wikipedia , lookup
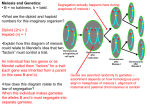
CHAPTER 5: THE INHERITANCE OF SINGLE-GENE DIFFERENCES 1. What determines how alleles of different genes are transmitted from one generation to the next? 2. How are characteristics transmitted from one generation to the next? Terminology (see also Glossary pages 655-681 and web site http://helios.bto.ed.ac.uk/bto/glossary/) Ascus – a sac that encloses a tetrad or octad of sexual spores (ascospores) First division segregation – different alleles go into different nuclei at the first meiotic division producing an MI division pattern of ascospores Second division segregation – different alleles go into different nuclei at the second meiotic division producing an MII division pattern of ascospores homozygous (= true -breeding): an individual having identical alleles of a gene heterozygous: an individual having different alleles of a gene monohybrid: an individual heterozygous at one gene first filial (F1) generation – the first generation resulting from a controlled cross between two known parents (P) second filial (F1) generation – the second generation resulting from a controlled cross between two known parents (P) test cross – a cross between an individual of unknown genotype and an individual homozygous recessive for a particular gene(s) product rule: the probability that two independent events occur together is equal to the product of their individual probabilities sum rule: the probability that either of two mutually exclusive events occurs is equal to the sum of their individual probabilitiessex chromosome – a chromosome whose presence is associated with a particular sex autosome – a chromosome that is not a sex chromosome sex linkage – the location of a gene on a sex chromosome hemizygous – a gene present in only one copy in a diploid organism The segregation of alleles during sexual cell division (Figure 4-20) 1. Consider mitosis in haploid cells, of either genotype A or genotype a: 2. Mitosis in diploid cells, of either genotype AA, aa or Aa Diploid organisms are classified as : o homozygous – meaning that they have identical alleles at a given gene (e.g. aa or AA) o heterozygous – meaning that they have different alleles at a given gene (e.g. Aa) In all cases, mitosis produces daughter cells of identical genotypes to the parent Consider a diploid meiocyte undergoing meiosis: Case 1: If a homozgyous meiocyte goes through meiosis, all resulting gametes are of one type: Genotype of meiocyte genotype of gametes Case 2: If a heterozygous meiocyte goes through meiosis, the resulting gametes are of 2 types (FIGURE 5-2): The segregation of chromosomes results in the segregation of alleles To assess the segregation of alleles, you must determine the genotype of the meiotic products If an allele confers a particular phenotype, the genotype can be inferred from the phenotype In a haploid life cycle, the haploid products of meiosis (non-transient) can be scored phenotypically In a diploid life cycle, the haploid products of meiosis are transient and cannot be scored phentoypically The diploid organism that results from their fusion can be scored phenotypically Based on the phenotype of the diploid, one can infer the genotype of meiotic products i.e. Determining the genotype of meiotic products is simpler and more direct in a haploid life cycle In both diploid and haploid life cycles, the products of multiple meioses are usually pooled Therefore, the segregation of alleles within a single meiosis can only be inferred from the ratio of alleles within the pool Some fungi produce tetrads, in which products of a single meiosis are not pooled, but rather are maintained in an ascus. In other fungi, the tetrads are linear, meaning that they reflect the segregation of chromatids during meiosis The segregation of alleles during meiosis is most directly visible in organisms with a haploid life cycle which produce linear tetrads (see figure 5-3): MEIOSIS I - separation of homologous chromosomes MEIOSIS II - separation of centromere and therefore separation of chromatids (i.e. formation of four meiotic products) MITOSIS - duplication of meiotic products to produce eight spores (FIRST DIVISION SEGREGATION (Figure 5-3) experiment: a diploid, heterozgyous parent undergoes meiosis a a a MI a A A A A a a a a M II a mitosis a A A A A A A results: a 4:4 ratio in spores (in this case, ratio indicates order as well as number), different alleles are separated by the first meiotic division = first division segregation (MI segregation pattern) conclusion: indicates that before meiosis I, different alleles were on different homologues (i.e. in their original conformation) and therefore that no recombination had occurred SECOND DIVISION SEGREGATION (described in lab manual) experiment: a diploid, heterozgyous parent undergoes meiosis a a a a a a MI A M II A A a A A A mitosis A a a A a A A results: - a 2:2:2:2 ratio in spores different alleles are separated in the second meiotic division = second division segregation (MII division pattern) conclusion: - meiosis I does NOT separate different alleles, therefore different alleles must have been in the same homologue (chromatids joined by a centromere) - suggests that crossover occurred between gene and centromere, so that the centromere links 2 non-identical chromatids - seen as 2:2:2:2 ratio in spores CALCULATING DISTANCES BETWEEN TWO POINTS ON A CHROMOSOME - the frequency of crossing over between any two points on a chromosome is directly proportional to the distance between the two points (more about this in Chapter 6) - expressed as an equation: distance between 2 points = % meiotic products showing recombination between the 2 points In linear octads, the frequency of MII octads is a phenotypic indication that a recombination has occurred between the gene and the centromere Therefore, can be used to calculate distance between gene and centromere distance between 2 points = % meiotic products showing recombination between the 2 points Within each MII octad, spores = products of meiosis Within these products of meioses, only 1/2 show recombination between gene and centromere Distance from gene to centromere = 1/2 MII octads x 100 Total tetrads scored Question: in the fungus Sordaria, how far is gene A from the centromere? Experiment: Allow a diploid fungal parent, heterozygous at gene A (Aa) to undergo meiosis Results: 1000 octads (representing 1000 meioses) are scored, and the following types of octad seen (MI = first division segregation pattern, MII = second division segregation pattern) Types of Octads 1 (MI) 2 (MI) 3 (MII) 4 (MII) 5 (MII) 6 (MII) A A A A a a a a a a a a A A A A a a A A a a A A A A a a A A a a A A a a a a A A a a A A A A a a 480 475 13 9 12 11 = 1/2 (MII tetrads)X 100 distance A to centromere 1000 1/2 (13 + 9 + 12 + 11) X 100 1000 = 2.25 map units = conclusion: gene A is 2.25 map units from the centromere Note: in the above data, 4 different, equally frequent MII patterns arise, depending on how the spindle attaches to the centromere INHERITANCE IN ORGANISMS WITH POOLED MEIOTIC PRODUCTS 1. crosses using haploid organisms in which products of meiosis are pooled - in organisms with a haploid life cycle, products of meiosis can be scored for genotype and phenotype in the haploid state - therefore, in the cross above, where one parent carries genotype b+ = orange and one carries genotype b = blue, the products of meiosis will be 1/2 orange and 1/2 blue 2. Crosses using diploid organisms in which products of meiosis are pooled individual 1 individiual 2 gametes offspring - in the diploid life cycle, the gametes are transient - phenotypes can usually only be observed in the diploid stages (in example, parents and offspring) - genotype of the gametes is inferred from the phenotype of the offspring 1) Mendel’s studies on the inheritance of characteristics (box 5-1) Why use pea plants as a study organism? • cheap, small, easy to grow • quick life span, many progeny • different characteristics available • cross- or self-pollinate • diploid Question: How are characteristics transmitted from one generation to the next? Methods: Results: expected results - blended phenotype observed results: purple female x white male = purple F1 white female x purple male = purple F1 - phenotype same as one parent conclusion: purple phenotype is DOMINANT Question: is the white characteristic still present (but masked) in the purple F1? Method: Results: Question: What accounts for the 3:1 ratio? Experiment: Generate F3 progeny from each phenotypic class in the F2. Results: Conclusion: Mendel’s interpretation 1) difference in phenotype (e.g . purple or white) is due to discrete and different hereditary determinants or factors (factors = genes, different factors = alleles) 2) each plant has 2 copies of the factor (2 alleles) for each character 3) for each factor, one copy is dominant to the other 4) parents pass one copy of each factor to offspring (one member of each gene segregates randomly into the gametes, so that each gamete has one allele) 5) union of gametes from each parent is independent of allele the gamete carries Mendel’s First Law The two members of a gene pair (alleles) segregate equally into the gametes, so that 1/2 the gametes carry one allele, and 1/2 the gametes carry the other allele = segregation of homologous chromosomes Consider the genotypes of Mendel’s peas: Parents: White = Purple = What gametes will be produced by each parent? What is the genotype of the F1 (purple) plant? What are the genotypes in the F2 population, and in what ratio are they found? Hypothesis: purple F1 is heterozygous, 2/3 of purple plants in F2 are heterozygous Experiment: Test Cross - an individual of unknown genotype is crossed to a homozygous recessive individual - in a test cross, the tester (pp) contributes only recessive gametes, therefore progeny are a direct indication of the gametes produced by the tested individual e.g. Pp X pp expected gametes from F1= expected gametes from pp = Results: expected progeny: tester observed progeny Conclusions: SEGREGATION OF SEX-LINKED GENES - in many animals and some dioecious plants, sex is determined by the presence of particular chromosomes, the sex chromosomes - remaining, non sex chromosomes are called “autosomes” (Figure 5-6) - e.g. in Drosophila, there are 4 chromosome pairs, - 3 pair = autosomal, 1 pair = sex chromosomes - the two sexes carry a different pair of sex chromosomes - in most species (e.g. Drosophila and humans), males carry an XY pair, females carry XX pair - X and Y chromosomes can pair during meiosis because of the presence of a small area of homology, the pairing region (Figure 5-7) - Remaining regions of the X and Y chromosomes carry different genes, with the Y chromosome carrying only a small number of genes female (homogametic sex) produces only X containing gametes male (heterogametic sex) produces X and Y containing gametes Meiosis in males: Sex Linkage • gene is on one of the sex chromsomes, usually the X • because the gene is on a sex chromsome, the phenotype associated with that gene will be associated with one sex more frequently than the other Sex linkage first recognized in experiments with Drosophila - Thomas Morgan (1910) (Figure 5-8) characteristic = eye colour wild type = red eyed = w+ mutant = white eyed = w Cross 1: parental: red eyed female x white eyed male F1: all red eyed F2: 3 red eyed : 1 white eyed Cross 2: Parental: white eyed female x red eyed male F1: 1 red eyed: 1 white eyed Conclusions Reciprocal crosses give different results Hypothesis: males = females = Key points: -all females get Xw+ from father = all are red - males receive Y from father that does not carry the w gene and therefore does not contribute to eye colour phenotype - phenotype depends on the mother’s contribution -males get either Xw+ or Xw from mother 1/2 red, 1/2 white -male is called hemizygous e.g. Xw Y • has only 1 allele of the w gene, so cannot be homozygous or heterozygous -whatever allele is present on the male’s X chromosome is expressed phenotypically Reciprocal cross parental: white eyed X red eyed female male w+ (Xw X w ) (X Y) gametes: only Xw Xw+ or Y F1: 1 red eyed : 1 white eyed (all female) (all male) X w+X w and Xw Y gametes: X w+ and Xw Xw contributes nothing to eye colour phenotype Xw and Y X w+X w (red female) X w+ Y Xw Xw+Y (red male) X w X w (white female) Xw Y Xw Y (white male) PEDIGREE ANALYSIS -applies the principles of Mendelian Genetics to humans, where controlled crosses are not possible, and existing family trees must be used -random crosses -small numbers, may not fit expected ratios Symbols used in pedigrees (figure 5-9): = male = female = unknown gender = affected individual (female) = mating between two individuals children from mating (in order of birth) RULES Autosomal recessive trait (Figure 5-10) 1) phenotype appears with equal frequency in each sex 2) two unaffected individuals produce 3) two affected individuals produce Assumptions 1) If an allele is rare and recessive, an unaffected individual marrying into a family will most likely be homozygous for the dominant alelele E.g. consider a recessive allele a, with a frequency of 1/100 autosomal dominant trait (Figure 5-14) 1) phenotype appears with equal frequency in each sex 2) affected individuals produce 3) affected individuals in every generation (usually = dominant) X-linked recessive trait (figures 5-16, 5-17) 1) More males than females with the trait 2) No children of an affected male show the trait, but 1/2 of grandsons will Xw Y Xw+Y Xw Y Xw+Y X w+X w Xw Y X w+X w+ X w+X w Xw+Y Xw+Y X-linked dominant trait (figure 5-19) 1) affected males pass the trait on to their daughters but not their sons 2) females married to unaffected males pass the trait on to 1/2 their sons and daughters SEGREGATION OF GENES IN THE ORGANELLAR GENOMES - both mitochondria and chloroplasts contain DNA - during cell fusion in sexual reproduction, male and female cells contribute nuclear DNA equally, but the female contributes most of the cytoplasm - cytoplasmic DNA comes only from the mother - maternal inheritance (Figures 5-21, 5-22)