* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Restriction Enzymes - mvhs
Survey
Document related concepts
Comparative genomic hybridization wikipedia , lookup
Agarose gel electrophoresis wikipedia , lookup
Maurice Wilkins wikipedia , lookup
Molecular evolution wikipedia , lookup
Non-coding DNA wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Biosynthesis wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
DNA vaccination wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Genomic library wikipedia , lookup
DNA supercoil wikipedia , lookup
Community fingerprinting wikipedia , lookup
Molecular cloning wikipedia , lookup
Transcript
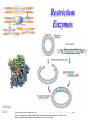
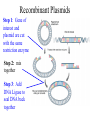
Restriction Enzymes AP Biology Unit 2 Images obtained without permission from http://w3.dwm.ks.edu.tw/bio/activelearner/14/images/ch14summary.gif and http://www.bioteach.ubc.ca/MolecularBiology/RestrictionEndonucleases/endonuclease%202.gif and http://www.symmation.com/gallery/images/restriction-enzyme-ecorV-th.jpg Restriction Enzymes: Molecular Scissors • Restriction enzymes (endonuleases) cut DNA at specific sequences • What kinds of bonds are broken when restriction enzymes cut? – Covalent bonds (within a single strand) – Hydrogen bonds (between Hydrogen strands) as a result of the bond Covalent bond strands coming apart Image taken without permission from http://www.bioteach.ubc.ca/MolecularBiology/RestrictionEndonucleases/endonuclease%202.gif Origins of Restriction Enzymes • Naturally found in different types of bacteria • Bacteria use restriction enzymes to protect themselves from foreign DNA • Bacteria have mechanisms to protect themselves from the actions of their own restriction enzymes • Have been isolated and sold for use in lab work Examples of Restriction Enzymes Enzyme Organism source Recognized Sequence EcoRI Escherichia coli 5' GAATTC 3’ 3' CTTAAG 5’ TaqI Thermus aquaticus 5' TCGA 3’ 3' AGCT 5’ HindIII Haemophilus influenzae BamHI Bacillus amyloliquefaciens 5'AAGCTT 3’ 3'TTCGAA 5’ 5' GGATCC 3’ 3' CCTAGG 5’ AluI Arthrobacter luteus 5' AGCT 3’ 3' TCGA 5’ Sticky Ends vs. Blunt Ends • When restriction enzymes cut, they produce either – Sticky ends (single stranded sections at the ends) – Blunt ends 5’ - - - G A A T T C - - - 3’ III I 3’ - - - C T T A A I I I G - - - 5’ Sticky ends Recombinant DNA • Recombinant DNA is constructed using restriction enzymes • Important: In order to join 2 pieces of DNA together they have to be cut by the same restriction enzyme – Why? – Otherwise, the sticky ends won’t match– DNA can’t bind together Recombinant Plasmids Step 1: Gene of interest and plasmid are cut with the same restriction enzyme Step 2: mix together Step 3: Add DNA Ligase to seal DNA back together Question… • What kinds of bonds does DNA Ligase form? – Covalent bonds (between nucleotides in a single strand of DNA) Putting It All Together Create recombinant plasmid Transform plasmid into E. coli Restriction Analysis • Using restriction enzymes to find out information about a piece of DNA • We can use restriction enzymes to find out – The size of a plasmid – If there are any restriction sites for a particular enzyme on a piece of DNA (ex. EcoRI) – How many restriction sites for a particular enzyme – Where the restriction sites are located