* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Fine scale mapping
Fetal origins hypothesis wikipedia , lookup
Gene desert wikipedia , lookup
Gene therapy wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Neocentromere wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Genetic drift wikipedia , lookup
Frameshift mutation wikipedia , lookup
X-inactivation wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Point mutation wikipedia , lookup
Quantitative comparative linguistics wikipedia , lookup
Population genetics wikipedia , lookup
Public health genomics wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Genome (book) wikipedia , lookup
Designer baby wikipedia , lookup
HLA A1-B8-DR3-DQ2 wikipedia , lookup
Gene expression programming wikipedia , lookup
A30-Cw5-B18-DR3-DQ2 (HLA Haplotype) wikipedia , lookup
FINE SCALE MAPPING
ANDREW MORRIS
Wellcome Trust Centre for Human Genetics
March 7, 2003
Outline
Introduction: fine scale mapping using
high-density SNP haplotype data.
Bayesian framework.
Gene trees and the coalescent process.
Genetic heterogeneity and shattered gene
trees.
Markov chain Monte Carlo (MCMC)
algorithm.
SNP genotype data.
Example: cystic fibrosis.
Introduction
Candidate region of the order of
1Mb in length.
Refine location of putative disease
locus within region.
Make use of high-density maps of
single nucleotide polymorphisms
(SNPs).
Type sample of affected cases and
unaffected controls.
Once upon a time…
Disease predisposition determined
by single locus in candidate region.
Each case chromosome carries a
copy of a disease allele, resulting
from a single recent mutation event
at disease locus.
Each control chromosome carries a
copy of the ancient normal allele at
the disease locus.
In an ideal world…
Excess sharing of SNP haplotypes in
the vicinity of the disease locus,
among cases and not among
controls.
Decreased probability of sharing as
distance from disease locus
increases.
Approximate location of disease
locus inferred.
Problems…
Gene tree and ancestral haplotypes
are unknown.
Marker mutations lead to mismatch
of alleles within preserved regions.
Multiple disease genes, multiple
mutations, and dominance.
Example: Cystic fibrosis (CF)
Fully penetrant recessive disorder, incidence ~1/2500
live births in white populations, less common in other
populations.
Preliminary linkage analysis suggested 1.8Mb
candidate region for a single CF gene on chromosome
7q31.
More recently, a 3bp deletion, ΔF508, has been
identified in the CFTR gene at ~0.88Mb into the
candidate region.
Now known that ΔF508 accounts for ~66% of all
chromosomal mutations in individuals with CF.
Remainder of CF chromosomes carry copies of many
other rare mutations in the same gene.
23 RFLPs used to identify haplotypes in 92 control
chromosomes and 94 case chromosomes, 62 of which
have been confirmed to carry ΔF508.
Challenges…
The ΔF508 locus does not lie at the
centre of the region of high LD.
Non-ΔF508 case chromosomes are
not expected to share the same
founder marker haplotype.
Useful test-data set for fine-scale
mapping methods…
Challenges…
The ΔF508 locus does not lie at the
centre of the region of high LD.
Non-ΔF508 case chromosomes are
not expected to share the same
founder marker haplotype.
Useful test-data set for fine-scale
mapping methods…
Published methods…
Bayesian framework (1)
Assume disease locus exists in
candidate region: aim is then to
estimate its location.
Approximate the posterior
distribution of location.
Allows assignment of probabilities
that disease locus lies in any
particular area of the candidate
region.
Bayesian framework (2)
Aim is to approximate the posterior
density of location of the disease locus,
given SNP haplotypes in cases A and
controls U, denoted f(x|A,U).
Depends on other model parameters M,
including gene tree, population haplotype
frequencies, etc…
Recover marginal posterior density by
integration over these nuisance
parameters,
f(x|A,U) =
∫f(x,M|A,U)dM
Bayesian framework (3)
By Bayes’ Theorem…
f(x,M|A,U) = C f(A,U|x,M) f(x,M)
Normalising constant.
Likelihood of haplotype data given
model parameters M and location x.
Prior density of M and x.
Bayesian framework (3)
By Bayes’ Theorem…
f(x,M|A,U) = C f(A,U|x,M) f(x,M)
Normalising constant.
Likelihood of haplotype data given
model parameters M and location x.
Prior density of M and x.
Bayesian framework (3)
By Bayes’ Theorem…
f(x,M|A,U) = C f(A,U|x,M) f(x,M)
Normalising constant.
Likelihood of haplotype data given
model parameters M and location x.
Prior density of M and x.
Bayesian framework (3)
By Bayes’ Theorem…
f(x,M|A,U) = C f(A,U|x,M) f(x,M)
Normalising constant.
Likelihood of haplotype data given
model parameters M and location x.
Prior density of M and x.
Control chromosomes
Assumed to carry an ancient normal allele
at the disease locus.
Effects of recent shared ancestry of less
importance, so simple model assumed:
f(A,U|x,M) = f(A|x,M) f(U|h)
The likelihood, f(U|h), depends only on
population SNP haplotype frequencies, h.
For many SNPs, the number of possible
haplotypes is large, so frequencies are
parameterised in terms of allele
frequencies and first-order LD between
pairs of adjacent loci.
Gene trees
Representation of the recent shared
ancestry of case chromosomes at the
disease locus.
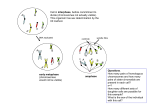
Star shaped tree: each case
chromosome descends independently
from founder. Assumes there is too much
information in sample about ancestral
recombination and mutation events.
Bifurcating tree: shared ancestral
recombination and mutation events
between chromosomes appear only once
in their shared ancestry.
Gene trees
Representation of the recent shared
ancestry of case chromosomes at the
disease locus.
Star shaped tree: each case
chromosome descends independently
from founder. Assumes there is too much
information in sample about ancestral
recombination and mutation events.
Bifurcating tree: shared ancestral
recombination and mutation events
between chromosomes appear only once
in their shared ancestry.
Tree specification
Topology T: the
branching pattern of
the tree.
Branch lengths, τ,
determined by the
waiting times, w,
between merging
events in the gene
tree.
Scaled in units of 2N
generations, where N
is effective population
size.
Root
Leaf nodes
Prior probability model
Uniform prior probability model for
population haplotype frequencies, the
location of disease locus, and the effective
population size.
Each gene tree topology has equal prior
probability.
Prior probability model reduces to:
f(x,M) = C f(w)
Need prior probability model for waiting
times between merging events.
The coalescent process (1)
Time between
merging event
from k to k-1
lineages.
Scaled in units of
2N generations.
Exponential
distribution with
rate k(k-1)/2.
The coalescent process (1)
Time between
merging event
from k to k-1
lineages.
Scaled in units of
2N generations.
Exponential
distribution with
rate k(k-1)/2.
Exponential: rate 8x7/2 = 28
Expected time: 0.0357
The coalescent process (1)
Time between
merging event
from k to k-1
lineages.
Scaled in units of
2N generations.
Exponential
distribution with
rate k(k-1)/2.
Exponential: rate 7x6/2=21
Expected time: 0.0476
The coalescent process (1)
Time between
merging event
from k to k-1
lineages.
Scaled in units of
2N generations.
Exponential
distribution with
rate k(k-1)/2.
Exponential: rate 2x1/2=1
Expected time: 1
The coalescent process (2)
Assumes constant effective population
size, N.
Flexible: can allow for exponential
population growth and population substructure.
Assumes sample is ascertained at random
from the population. Problem: case
chromosomes ascertained because they
carry a copy of the disease mutation.
Assumes sample has single common
ancestor. Problem: genetic
heterogeneity.
The shattered coalescent model
Generalisation of the coalescent process to allow
branches of the gene tree to be removed.
Introduce indicator variable, zb, for each node, b,
taking the value 1 if b has a parent in the gene
tree and 0 otherwise.
Allows for singleton leaf nodes, corresponding to
sporadic case chromosomes, and disconnected
sub-trees, corresponding to independent
mutation events at the same disease locus.
Assume number of branches of gene tree not
removed in the shattered coalescent process
given by binomial distribution, with shattering
parameter ρ.
Ancestral haplotypes
Haplotypes, I, carried by internal nodes of the gene
tree are unknown.
To calculate posterior probability, need to integrate
over distribution of possible ancestral haplotypes,
which depends on gene tree and other model
parameters.
Treated as augmented data in Bayesian framework:
enters posterior probability through likelihood…
f(x|A,U) =
∫∫
f(x,M,I|A,U)dMdI
and…
f(x,M,I|A,U) = C f(A,U,I|x,M) f(x,M)
Likelihood calculations
If node has no parent in
shattered gene tree,
treat as a random
chromosome from the
population (sporadic or
founder for mutation).
If node has parent in
genealogy, depends on
marker haplotype
carried by the parental
node, and the
occurrence of
recombination and
mutation events along
the connecting branch.
Likelihood calculations
If node has no parent in
shattered gene tree,
treat as a random
chromosome from the
population (sporadic or
founder for mutation).
If node has parent in
genealogy, depends on
marker haplotype
carried by the parental
node, and the
occurrence of
recombination and
mutation events along
the connecting branch.
MCMC algorithm (1)
Need to calculate joint posterior distribution
f(x,h,T,w,z,N,ρ,I|A,U).
Parameter space extremely complex, so cannot
be calculated analytically.
Markov chain Monte Carlo (MCMC) algorithm
approximates the posterior distribution by
sampling from f(x,h,T,w,z,N,ρ,I|A,U).
Computationally intensive, but becoming more
practical with improvements in computing power.
Can handle missing SNP data: treat as
augmented data in the same way as ancestral
haplotypes.
MCMC algorithm (2)
Let S denote current set of model parameters
{x,h,T,w,z,N,ρ,I}.
Propose “small” change to model parameters, S*.
Accept S* in place of S with probability
f(S*|A,U)/f(S|A,U).
If S* is not accepted, the current parameter S is
retained.
Initial burn-in to allow convergence of f(S|A,U)
from random starting parameter set.
Subsequent sampling period, parameter set
recorded every rth step of the algorithm: each
recorded output represents a random draw from
f(S|A,U).
MCMC algorithm (3)
Location
101
102
103
104
105
106
107
108
109
110
0.47374
0.40629
0.46534
0.48211
0.43808
0.44607
0.41822
0.40934
0.41032
0.45020
Tree height
ρ
N
2557.62766
2112.19993
1679.71719
2229.24788
2402.10599
2275.33453
3016.70273
2534.50113
3122.91416
3209.14218
4.24189612
4.16846454
4.30423786
4.33740414
4.29011844
4.03331587
4.39000994
4.07270615
4.25386813
4.34316471
10849.19083
8804.63049
7229.90233
9669.14899
10305.31919
9177.14285
13243.35496
10322.27832
13284.46504
13937.83307
0.78104
0.79777
0.75364
0.78009
0.82178
0.82601
0.77768
0.81590
0.82479
0.78422
-1769.51173
-1788.66623
-1854.19049
-1763.70173
-1760.56671
-1775.90300
-1844.20629
-1861.97411
-1814.27448
-1801.44160
Log posterior
probability
MCMC algorithm (3)
Location
101
102
103
104
105
106
107
108
109
110
0.47374
0.40629
0.46534
0.48211
0.43808
0.44607
0.41822
0.40934
0.41032
0.45020
Tree height
ρ
N
2557.62766
2112.19993
1679.71719
2229.24788
2402.10599
2275.33453
3016.70273
2534.50113
3122.91416
3209.14218
4.24189612
4.16846454
4.30423786
4.33740414
4.29011844
4.03331587
4.39000994
4.07270615
4.25386813
4.34316471
10849.19083
8804.63049
7229.90233
9669.14899
10305.31919
9177.14285
13243.35496
10322.27832
13284.46504
13937.83307
0.78104
0.79777
0.75364
0.78009
0.82178
0.82601
0.77768
0.81590
0.82479
0.78422
-1769.51173
-1788.66623
-1854.19049
-1763.70173
-1760.56671
-1775.90300
-1844.20629
-1861.97411
-1814.27448
-1801.44160
Log posterior
probability
Cystic fibrosis: revisited
Assume a fixed recombination rate of
0.5cM per Mb and a marker mutation rate
of 2.5 x 10-5 per locus, per generation.
Each run of MCMC algorithm begins with
20,000 step burn-in period: thrown away.
Subsequent 200,000 step sampling
period, output recorded every 50th step of
the algorithm: 4000 outputs.
Two analyses of CF data performed:
control chromosomes (92) and (i) ΔF508
case chromosomes (62) only; (ii) all case
chromosomes (94).
Cystic fibrosis: summary statistics
Parameter
ΔF508 subset
All cases
Location x
(Mb)
0.864
0.654-1.040
0.851
0.650-1.003
Shattering
parameter ρ
0.935
0.857-0.985
0.829
0.746-0.892
595
183-1877
824
246-3257
Time to MRCA
(generations)
Cystic fibrosis: genetic heterogeneity
Structure of shattered gene tree provides
information about genetic heterogeneity at
disease locus.
For each output of MCMC algorithm, record
shattered gene tree.
For each pair of chromosomes, record whether
they appear in the same sub-tree.
Over all outputs, estimate probability that each
pair of chromosomes carry the same allele at the
disease locus.
Cluster chromosomes according to these
probabilities: cladogram to represent genetic
heterogeneity.
SNP genotype data
SNP haplotype rarely available.
Could infer haplotypes from SNP genotype data:
PHASE, SNPHAP, HAPLOTYPER algorithms.
Better to treat haplotypes as augmented data in
Bayesian framework…
f(x|G) =
∫∫∫∫
f(x,M,I,A,U|G)dMdIdAdU
and…
f(x,M,I,A,U|G) = C f(A,U,I|x,M) f(x,M)
Cystic fibrosis: revisited – again!
Create genotype data from original
CF haplotype data.
Pair together case chromosmes at
random.
Pair together control chromosomes
at random.
Total sample: 46 controls and 47
cases.
Cystic fibrosis: genotypes v haplotypes
Parameter
Genotypes
Haplotypes
Location x
(Mb)
0.855
0.625-1.137
0.851
0.650-1.003
Shattering
parameter ρ
0.842
0.771-0.901
0.829
0.746-0.892
375
107-871
846
367-1657
Effective
population
size N
Limitations
Computationally intensive – limited
to sample sizes ~100 cases and
controls with up to 20 SNPs.
Alternative approach: do not model
gene tree explicitly – estimate
shattered gene tree using standard
clustering methods.
Summary
High density SNP map of the human
genome now available.
Fine scale mapping of disease loci
requires effective modelling of shared
ancestry of sample of case and control
chromosomes.
Methods exist for haplotype and genotype
data: MCMC algorithms are very
computationally intensive and are
currently limited to relatively small
sample sizes.
Further development is necessary…