* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Gene expression Profiling of Duodenal Biopsies
Gene nomenclature wikipedia , lookup
Public health genomics wikipedia , lookup
Genome evolution wikipedia , lookup
Genome (book) wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene therapy wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microevolution wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Nutriepigenomics wikipedia , lookup
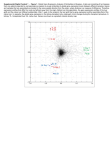
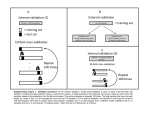
Gene Expression Profiling vs. histopathologic assessment comparative analysis using a simplified classification scheme – a Hanna Bragde, Ulf Jansson, Ingvar Jarlsfelt, Jan Söderman Division of Medical Diagnostics [H.B., I.J., J.S.], Department of Pediatrics [U.J.], Ryhov County Hospital, Jönköping, SE-551 85, Sweden. Objectives: Celiac disease (CD) is identified by histopathologic changes in the small intestine which normalise during a gluten-free diet. The histopathologic assessment (HA) is usually routine, but can be problematic. We have recently described discriminant analysis (DA) of gene expression in duodenal biopsies, for diagnosis and monitoring of CD (accepted for publication). Here we extend the results by building the algorithms using a larger cohort of patients, and examine their performance in relation to routine, clinical HA. Methods: DA was based on gene expression of eight genes selected to reflect crypt hyperplasia (MAD2L1, MKI67), villous atrophy (APOC3, CYP3A4), inflammatory response (CXCL11, IL17A, CTLA4), and intestinal permeability (OCLN). The classification algorithms were derived by fitting gene expression values to a simplified histopathologic classification (Corazza et al. (2005, 2007)) of biopsies (assessed by a single pathologist) from 53 paediatric cases. Performance of DA was investigated in relation to a set of cases assessed during routine, clinical practice. Results: DA correctly classified all cases (except grade A) present in the model using leave-one-out cross-validation. There were too few grade A cases for crossvalidation, but the model resulted in distinct posterior probabilities for these biopsies. Among 10 cases with primary, diagnostic biopsies there were four cases with a normal mucosa and six cases with a lesioned mucosa according to routine, clinical HA. For one of the cases with a histologically lesioned mucosa, DA disagrees. However, for this case, DA presented a rather high posterior probability for grade B1 and reduced expression of villi genes was observed. Among five cases with histologically normalised mucosa, assessed after gluten-free diet, one was classified as grade A according to DA. Conclusion: Using more cases per grade, it seems possible to establish DA as a useful tool for assessing biopsies from patients investigated for CD.