* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download [PDF]
Epigenetics in stem-cell differentiation wikipedia , lookup
Primary transcript wikipedia , lookup
Epigenetics of cocaine addiction wikipedia , lookup
Genomic imprinting wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Designer baby wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
History of genetic engineering wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Epigenetics of depression wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene expression programming wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
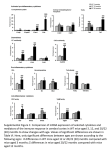
DNA AND CELL BIOLOGY Volume 20, Number 12, 2001 © Mary Ann Liebert, Inc. Pp. 781–789 In Vivo Roles of RORa and Sp4 in the Regulation of Murine Prosaposin Gene PENG JIN, YING SUN, and GREGORY A. GRABOWSKI ABSTRACT Prosaposin has a central role in intracellular glycosphingolipid catabolism and also has extracellular functions. This locus is regulated temporally and spatially. The highest mRNA expression occurs in the central nervous system (CNS) and reproductive system. In vitro, the CNS-expressed proteins Sp4 and RORa bind to Sp1 and RORE sites within a 310-bp fragment directly upstream of the transcription start site. These transcription factors exhibit negative cooperativity in vitro for prosaposin expression. Mice deficient in RORa and Sp4 (Staggerer [Sg2/2 ] and Sp4 knockout [Sp4 KO], respectively) containing selected prosaposin promoter deletion transgenes were used in comparative expression studies to evaluate this negative cooperativity in vivo. Constructs containing the RORE or Sp1/U cluster alone were independently stimulatory. Deletion of the Sp1/U site led to a decrease in reporter activity only in the cerebellum of Sg2/2 mice. The deletion of RORE and Sp1/U sites did alter the increase of reporter activity in the brain and eye, but not in the spinal cord, of Sg2/2 mice. These results indicate that Sp4 and RORa play minor and major roles, respectively, in regional expression of the prosaposin locus in the brain, whereas expression in the spinal cord is independent of RORa. INTRODUCTION P with intracellular and extracellular functions (Hiraiwa et al., 1992; Morales et al., 1995; O’Brien et al., 1994; O’Brien et al., 1988; Rorman and Grabowski, 1989). The mRNA encodes four saposins, termed A, B, C and D, that, together with lysosomal hydrolases, control the complex metabolism of glycosphingolipids (Sandhoff et al., 2001). Intact mature prosaposin also facilitates in vitro glycosphingolipid transfer between liposomal membranes (Hiraiwa et al., 1992). Intriguingly, prosaposin has neuritogenic and nerve regrowth-promoting properties ex vivo and in vivo (Igase et al., 1999; Kotani et al., 1996; O’Brien et al., 1994; Sano et al., 1994). Human prosaposin or individual saposin deficiencies lead to lysosomal storage diseases with massive accumulation of glycosphingolipids in a variety of organs, particularly the neurons of the brain (Bradova et al., 1993; Harzer et al., 1989; Hulkova et al., 2001). Targeted disruption of the murine prosaposin gene produces a similar complex phenotype that includes severe central nervous system (CNS) disease with extensive storage of glycosphingolipid (Fujita et al., 1996). Unlike many other lysosomal “housekeeping” genes, expression ROSAPOSIN IS A MULTIFUNCTIONAL LOCUS of prosaposin is regulated temporally and spatially. In mice, the highest levels of mRNA expression are in specific neurons of the adult cerebrum, the Purkinje cell layer of the cerebellum, and neurons of the lateral horns of spinal cord (Sun et al., 1994). Components of the hindbrain also show higher levels of mRNA expression early in embryogenesis (Sprecher-Levy et al., 1993). To elucidate the basis for this transcriptional control, we characterized the mouse and human promoters (Sun et al., 1997, 1998). In a variety of murine cell lines, functional positive and negative regulatory elements were found within 2400 bp 59 to the transcription start site (TSS) (Sun et al., 1997). A major regulatory fragment was characterized within 310 bp 59 of the TSS. It contains three DNase I protected regions, including a 39 Sp1 binding site, a retinoic acid receptor-related orphan receptor (RORa) binding site (RORE), and a 59 region with three overlapping Sp1 half-binding sites and a site for an unidentified transcription factor, a U region (Fig. 1). This latter segment was termed the Sp1/U cluster. Several members of the Sp family (Sp1, Sp3, and Sp4) and the orphan nuclear receptor (RORa) bind to this region in vitro. Site-directed mutagenesis and in vitro analysis suggest a complex cooperative (positive and negative) interaction among the various transcription fac- The Division of Human Genetics, Children’s Hospital Research Foundation at Children’s Hospital Medical Center, Cincinnati, Ohio. 781 782 JIN ET AL. A B FIG. 1. Schematic representation of the mouse prosaposin promoter region, transgenic constructs, and distribution of luciferase reporter gene in transgenic mice. (A) Different lengths of the prosaposin promoter 59 flanking DNA were fused with luciferase reporter gene. The promoter regions covered by different constructs are indicated, as well as transcription factor binding site identified by in vitro analysis. The segregation of visceral and CNS components of the promoter has been shown previously (Sun et al., 2000). The boundary for initiation of visceral promoter activity is not clearly defined at .310 bp. (B) The distribution and expression level of luciferase reporter gene in each representive of transgenic mouse line (from three to five founders) are compared after normalization by the copy number of each transgenic line and protein concentration. The data represent the average of wildtype transgenic mice at 3 weeks of age. tors that bind to the 310-bp region (Jin et al., 1998). Using transgenic mice, the 310-bp region was found to be responsible for tissue-preferential expression in the CNS. Nearly exclusive expression was observed in the cerebrum, cerebellum, spinal cord, and eyes of adult transgenic mice containing constructs of 234 and 310 bp 59 of the TSS. This CNS-preferential expression was related to RORE and the Sp1/U cluster, as no expression was observed with constructs containing only the 39 Sp1 binding site. Increasing CNS expression and the appearance of substantial expression in visceral tissues (e.g., liver, spleen, lung, thymus and heart) was obtained with transgenic mice bearing longer constructs, up to 2400 bp 59 of the TSS (Sun et al., 2000). The expression of reporter genes from these constructs corresponded closely to the cellular distribution of prosaposin (Sun et al., 2000). The human promoter does not contain the 59 Sp1/U cluster in the comparable region but has AP-1, Oct-1, and two RORa binding sites that are DNase I protected by selected nuclear extracts (Sun et al., 1998). Among the transcription factors that bind to this 310 bp region in the mouse promoter, Sp1 and Sp3 are ubiquitously expressed, but RORa and Sp4 have high-level expression only in CNS tissues (Matsui et al., 1995; Saffer et al., 1991; Supp et al., 1996). The RORa is a retinoic acid receptor-related orphan nuclear receptor that has several isoforms generated by alternative RNA splicing (Giguere et al., 1994). Expression of RORa is confined to the CNS, skin and testis (Steinmayr et al., 1998). In the CNS, RORa is expressed in thalamus, olfactory bulb, and cerebellum, with highest levels in Purkinje cells (Steinmayr et al., 1998). The Staggerer (Sg) mouse, a spontaneous mutant, has a deletion within the RORa gene that prevents translation of the ligand-binding domain (Hamilton et al., 1996). This mouse develops progressive staggering gait, mild tremor and hypotonia (Sidman et al., 1962). Targeted disruption of the RORa gene produced a similar phenotype, confirming the causality of the RORa deletion for the phenotype of Sg2/2 mice (Dussault et al., 1998; Steinmayr et al., 1998). The Sp4 gene belongs to the Sp1 multigene family that share similar DNA consensus binding sequences (Hagen et al., 1992; Kingsley and Winoto, 1992). The gene is highly expressed in mouse brain. Gene targeting of Sp4 resulted in two phenotypes: Two-thirds of the homozygous mutants die within the first few days after birth. The remaining third survive but are much smaller than their wildtype or heterozygous littermates (Gollner et al., 2001; Supp et al., 1996). Whereas the fertility of the TRANSCRIPTIONAL REGULATION OF PROSAPOSIN 783 female mutants appears normal, homozygous males do not breed despite having histologically normal testes containing mature sperm. In this work, Staggerer mice and Sp4 knockout (Sp4 KO) mice were used to analyze the in vivo roles of RORa and Sp4 in driving tissue-preferential expression of prosaposin in the CNS. Several crosses were made between mice containing luciferase driven by various deletion constructs of the prosaposin promoter and the Sg2/2 or the Sp4 KO mice. The distribution and expression of luciferase reporter genes were compared among wildtype, heterozygous, and homozygous animals. These data support an important role for RORa in the regulation of the prosaposin locus in vivo. gerer mice (The Jackson Laboratory, Bar Harbor, Maine). Dr. Steve Potter (Children’s Hospital Research Foundation, Cincinnati) provided the Sp4 gene targeted mouse (Sp4 KO). MATERIALS AND METHODS Generation of transgenic mice within different mutant backgrounds Four transgenic mouse lines with different lengths of prosaposin promoter were used: 234Luc, 310Luc, 2400Luc, and 2400DLuc (Fig. 1) (Sun et al., 2000). All the transgenic mice and Sg mice were B6C3, whereas the Sp4 KO mice were C57Bl/6. Each transgenic mouse line was bred with either Sg or Sp4 KO heterozygous mice to generate Sg or Sp4 KO heterozygotes bearing the various transgenes. The Sg1/2 or Sp41/2 mice bearing each transgene were crossed with Sg1/2 or Sp41/2 mice to obtain Sg2/2 and Sp4 2/2 animals bearing each transgene (Fig. 2A). The F1 offspring were analyzed in this study. Materials The following were from commercial sources: Magna Charge Nylon transfer membrane (Micron Separation Inc, Westbond, MA); PCR 103 buffer (GIBCO, Grand Island, NY); oligonucleotide synthesis and NAP-10 columns (Pharmacia, Piscataway, NJ); [a-32P]-dCTP (DuPont, NEN Research Products, Boston, MA); restriction enzymes and Taq polymerase (New England Biolabs, Beverly, MA); luciferase assay system (Promega, Madison, WI); Monolight 2010 Luminometer (Analytical Luminometer Laboratory, San Diego, CA); Bradford reagent system (Bio-Rad Laboratory, Hercules, CA); and Stag- Genotyping of transgenic mice For each transgenic mouse or Sp4 KO mouse, the genotype was determined by Southern blotting as described (Sun et al., 2000; Supp et al., 1996). For Sg mice, two pairs of primers, located within and outside of the ROR deletion region, were designed for PCR (see below). The PCRs were run at 94°C for 1 min, 55°C for 1 min, 72°C for 2 min for 35 cycles with extension for 7 min at 72°C. Two bands, 315 bp (Sg1S and Sg3R) and 150 bp (SgF2 and SgR3), were obtained for the deleted and normal allele, respectively (Fig. 2B): A B FIG. 2. Breeding strategy and PCR-based genotyping of Staggerer mice. (A) Each mutant heterozygote (Sg1/2 or Sp4 1/2 ) was bred with representatives of each transgenic line (234Luc, 310Luc, 2400Luc, and 2400DLuc). Heterozygotes with different transgenes were generated and bred with mutant heterozygotes. The pups of the next generation were used for analysis. (B) Staggerer mice were genotyped by PCR with two sets of primers located within and outside the deletion region. The 315-bp and 150-bp PCR products were obtained for the deleted and normal allele, respectively. 784 JIN ET AL. Sg1S: 59-GCCTTACCCCAGCACATGGTA-3 9 Sg3R: 59-CTGCTTTTAGAAGCACAATTT-3 9 SgF2: 59-ATCTCCTGTTTTCTCCCCAG-3 9 SgR3: 59-AAGAGAAAGTCCTCCAACTC-3 9 Luciferase activity assay For mice having the 2400Luc and 2400DLuc transgenes, tissues were harvested at 3 weeks of postnatal age and included the cerebrum, cerebellum, eye, spinal cord, thymus, heart, lung, liver, spleen, and kidney. For mice having the 234Luc or 310Luc transgenes, cerebrum, cerebellum, eye, spinal cord and heart were collected. Organs were pulverized under liquid N2 and homogenized in Reporter Lysis Buffer (Promega). The supernatant liquids from tissue homogenates were assayed for luciferase activity using the Promega assay system and a Monolight 2010 Luminometer (Analytical Luminometer Laboratory). The protein concentrations were determined by the Bradford assay (Bio-Rad). The luciferase activity of each tissue (from five or more mice) was normalized for protein concentration and reporter gene copy number. Two-tailed Student’s t-tests were used to evaluate the data. Expression and tissue distribution of luciferase reporter genes in Sg2/2 and Sp4KO 2/2 mice The luciferase expressions of the various transgenes were compared among wildtype, heterozygous, and homozygous Sg and Sp4 KO mice. Compared with wildtype, the luciferase expression of 310Luc/Sg1/2 was unchanged. The expression of 310Luc/Sg2/2 was increased twofold to threefold in the cerebrum, cerebellum, eye, and spinal cord (P ,0.005) (Fig. 3A). Importantly, these comparative analyses were done with littermates from a single founder. For the 310Luc transgenic line in this study, low-level expression of luciferase was also observed in the heart. About twofold to threefold increases in luciferase were present in the hearts of 310Luc/Sg2/2 mice as well (Fig. 3A). Using immunofluorescence with antiluciferase antibody, similar cellular distribution of luciferase was observed in the A RESULTS Generation of Sg2/2 and Sp4 KO mice with transgenes The transgenic mouse lines with mouse prosaposin promoter deletion constructs were 234Luc, 310Luc, 2400Luc, and 2400DLuc (Fig 1A). The tissue specific expression pattern of promoter deletion constructs was reported previously (Sun et al., 2000). The expression pattern for each construct was same among the founder lines. With the 234Luc transgene, low-level expression of luciferase was found only in the CNS, whereas with 310Luc, higher expression was detected. With the 2400Luc transgene, substantial luciferase expression was obtained in visceral tissues in addition to high CNS expression. Deletion of RORE and the Sp1/U cluster (2400DLuc) resulted in greatly decreased expression in whole CNS with little or no change in visceral tissues compared with the expression seen with 2400Luc (Fig. 1). One representative transgenic line for each construct was used in this study. The Sg or Sp4 KO heterozygotes (1/2) with these transgenes were obtained appropriate crosses. The Sg1/2 or Sp4 KO1/2 animals bearing various transgenes were backcrossed to Sg1/2 or Sp4 KO1/2 to obtain homozygous, heterozygous, and wildtype animals with the desired transgene for luciferase analyses (Fig. 2). The Sg2/2 mice and their littermates were sacrificed at 21 days, as the homozygotes do not survive beyond this age owing to the severe phenotype or much smaller size (Sidman et al., 1962; Supp et al., 1996). Age-matched Sp4 KO mice were also obtained. At this age, wildtype mice with the 310Luc or 2400Luc transgene have similar tissue distribution of luciferase gene expression but at higher levels than those in 6-week-old adult mice. This finding is consistent with our results that the expression of these transgenes is developmentally regulated (Sun and Grabowski, unpublished data). B FIG. 3. Comparison of expressions of 310Luc transgene in CNS and heart in wildtype and mutant mice. (A) Expression of 310Luc in Sg mice with wildtype (1) and mutated (2) gene. The P value for each tissue was calculated by comparing wildtype (1/1) or heterozygote (1/2) with homozygote Sg (2/2): cerebrum (P ,0.004), cerebellum (P ,0.001), eye (P ,0.006), heart (P ,0.0001), and spinal cord (P ,0.0007). (B) Expression of 310Luc in Sp4 gene-targeting mice was compared in wildtype and homozygotes. The P value with significant difference was found as above in cerebrum (P ,0.0001), cerebellum (P ,0.015), heart (P ,0.02), and spinal cord (P ,0.01). *P ,0.05; **P ,0.01. Two-tailed Student’s t-test was used to evaluate the data for Figures 3 through 6. TRANSCRIPTIONAL REGULATION OF PROSAPOSIN 785 310Luc and 310Luc/Sg2/2 mice (data not shown). Similarly increased expression (, twofold) of luciferase was present in the CNS and heart of 310Luc/Sp42/2 mice compared with wildtype 310Luc mice (P ,0.02) (Fig. 3B). The exception was in the eye of 310Luc/Sp42/2 animals, in which expression levels were unchanged. This finding correlates with the lack of Sp4 expression in the eye (Supp et al., 1996). Our previous in vitro data suggested a cooperative interaction of RORE and the 59 Sp1/U cluster leading to negative effects on expression (Jin et al., 1998). To separate these two elements and evaluate them individually, a transgenic mouse with 234Luc, which contains only RORE and the 39 Sp1 binding site, was bred to Sg1/2 and Sp4 KO1/2 mice. For 234Luc/Sg2/2 , no change was found in the cerebrum, eye, or spinal cord, but the luciferase activity in the cerebellum was decreased by 75% compared with 234Luc (P ,0.001) (Fig. 4A). This indicates that without the Sp1/U cluster, RORa could bind to the RORE and activate transcription, but deletion of RORa affected only the expression in the cerebellum. Previously, gel supershift assays with anti-Sp4 antibody suggested that Sp4 bound to the 39 Sp1 binding site and not the 59 Sp1/U cluster, yet no change was detected in any of the tissues of 234Luc/Sp4 2/2 (Fig. 4B). The data from 310Luc/Sp42/2 and 234Luc/Sp4 2/2 suggested that Sp4 modulates in vivo expression through the 59 Sp1/U cluster instead of the 39 Sp1 binding site. To further validate the results with 310Luc/Sg2/2 and 310Luc/Sp4 2/2 , the expression of luciferase in the context of 2400Luc was analyzed in Sg2/2 and Sp42/2 mice. For 2400Luc/Sg2/2 , twofold to threefold increases of luciferase activity were present in the cerebrum, eye, and spinal cord (P ,0.05), similar to the increases observed with the 310Luc/Sg2/2 mice. However, the difference in the cerebellum between 2400Luc/Sg1/1 and 2400Luc/Sg2/2 was not significant, with P >0.1 (Fig. 5A). In addition, the low level expression of the reporter gene in visceral tissues, including heart, did not show a significant difference between wildtype and Sg2/2 mice, which suggests that RORa specifically modulates the expression of prosaposin in the CNS (Fig. 5A). Unlike the results with 310Luc/Sp42/2 animals, we did not detect any change in luciferase expression in 2400Luc/Sp42/2 mice (Fig. 5B). As a negative control, we also generated 2400DLuc/Sg2/2 and 2400DLuc/Sp4 2/2 mice. As expected, no change in luciferase activity was detected in the CNS and visceral tissues of either mutant transgenic mouse. The exception was the spinal cord of 2400DLuc/Sg2/2 animals, in which the luciferase activity was A B FIG. 4. Comparison of expression of 234Luc transgene in CNS in wildtype and mutant mice. (A) Expression of 234Luc in Sg mice was examined in CNS. The P value for cerebellum was ,0.001. (B) Expression of 234Luc in Sp4 gene-targeting mice was compared in wildtype and homozygous animals. **P ,0.01. 786 JIN ET AL. A B FIG. 5. Comparison of expression level of 2400Luc transgene in both CNS and visceral tissues in wildtype and mutant mice. (A) Expression of 2400Luc in Sg (Sg2/2 ) mice was examined in both CNS and visceral tissues. The P values for the tissues with significant difference were cerebrum (P ,0.014), eye (P ,0.05), and spinal cord (P ,0.02). The P value for cerebellum was 0.123. (B) Expression of 2400Luc in Sp4 gene-targeting mice was compared in wildtype and homozygous animals. *P ,0.05. still increased relative to the constructs without the deletion of RORE and the Sp1/U cluster (P ,0.01) (Fig. 6). DISCUSSION Prosaposin has a central role in intracellular glycosphingolipid catabolism and other extracellular functions (Sandhoff et al., 2001) and is regulated temporally and spatially (Sun et al., 1994). This locus has highest expression in the CNS and reproductive system. Understanding the basis of prosaposin gene regulation should provide insight into the physiological significance of this intriguing “lysosomal locus.” Previous in vitro and in vivo approaches identified a region within 310 bp 59 of the TSS as responsible for tissue-preferential expression of the prosaposin gene in the CNS (Jin et al., 1998; Sun et al., 2000). Members of the Sp protein family (Sp1, Sp3, and Sp4) and an orphan nuclear receptor (RORa) were shown to be involved in the regulation of this 310-bp region in vitro. Among these transcription factors, RORa and Sp4 have tissue specific expression in the CNS (Matsui et al., 1995; Supp et al., 1996). Using RORa- and Sp4-deficient mice (Sg and Sp4 gene targeted), the in vivo roles of these ligands in driving the tissuepreferential expression in the CNS were evaluated. In Sg mice, the expression level of luciferase reporter gene in the CNS increased twofold to threefold over that in Sg1/2 or Sg1/1 mice having the same 310Luc and 2400Luc transgenes (see Figs. 3A and 5A). These transgenes contain RORE and the Sp1/U cluster. With the shorter construct, 234Luc, which contains only RORE and no 59 upstream elements, the luciferase activity was greatly reduced in the cerebellum of Sg2/2 mice and there was no change in the other CNS tissues. These results confirm the predicted negative cooperativity between RORa and the Sp1/U cluster even though these elements were independently stimulatory. The absence of altered reporter activity in the non-cerebellum brain tissues suggests the presence of other factors in these tissues that may bind to RORE. In the 2400DLuc/Sg2/2 mice, containing no RORE and Sp1/U cluster, the luciferase activity in the spinal cord still increased, thereby indicating that the regulation in spinal cord may be independent of RORa reg- TRANSCRIPTIONAL REGULATION OF PROSAPOSIN A B FIG. 6. Comparison of expression of 2400DLuc transgene in wildtype and mutant mice. (A) Expression of 2400DLuc in Sg mice was examined. The P value for spinal cord was ,0.01. (B) Expression of 2400DLuc in Sp4 gene-targeting mice was compared in wildtype and homozygous animals. **P ,0.01. ulation. In Sp4 KO mice, similarly increased luciferase activity in the CNS was obtained only from 310Luc, not the longer construct, 2400Luc. The RORa gene belongs to subfamilies of nuclear receptors, ROR (retinoic acid receptor-related orphan receptor; also termed RZR), that bind to DNA both as monomers and heterodimers (Carlberg and Wiesenberg, 1995; Giguere et al., 1994). This orphan nuclear receptor group also includes RORb and RORg (Carlberg et al., 1994; Hirose et al., 1994). The RORa gene encodes four isoforms, a1, a2, a3 and a4, which are alternatively spliced products of the gene. These isoforms share common DNA-binding and putative C-terminal ligandbinding domain but differ in their N-terminal domains and display distinct DNA recognition and transactivation properties; e.g., RORa1 and RORa2 have different binding specificities (Giguere et al., 1994). The sequence of RORE in the mouse prosaposin gene is identical to the consensus sequence for RORa1 binding, which suggests that RORa1, rather than other isoforms, is more likely involved in the regulation within this 310-bp region. The spatiotemporal expression of RORa is under isoform-specific regulation. In the thalamus, there is only 787 RORa1 mRNA. The RORa4 transcript is predominant in leukocytes and skin, whereas the RORa2 and RORa3 transcripts are detected exclusively in testis. In the remaining tissues, including the cerebellum, there is a mixture of RORa1 and RORa4 transcripts (Sashihara et al., 1996; Steinmayr et al., 1998). In the brain, RORa mRNA localizes to cerebellar Purkinje cells, various thalamic nuclei, and, during development, other brain areas. Several RORa natural target genes have been identified, and RORa can act as both activator and repressor (Delerive et al., 2001; Dussault and Giguere, 1997; Matsui, 1996; Matsui, 1997; Steinhilber et al., 1995; Vu-Dac et al., 1997; Wiesenberg et al., 1995). They assume their function by interacting with a coactivator or corepressor complex (Jetten et al., 2001). By breeding Staggerer with our transgenic mice, we showed RORa to function negatively in the contexts of the 310bp or 2400-bp region. However, within shorter constructs (234Luc), RORa appears to be a critical activator only in the cerebellum, and other brain regions were not affected by the depletion of RORa expression. In addition, the control of prosaposin expression in spinal cord seems unrelated to RORa. These results are consistent with our previous in vitro mutagenesis data. When we mutated RORE with 310Luc, the promoter activity went up whereas the activity dramatically decreased when RORE was mutated in the context of the 234Luc construct (Jin et al., 1998). A significant question is why only cerebellar promoter activity is decreased in 234Luc/Sg2/2 and not that in other CNS tissues. Interestingly, RORa is highly expressed in cerebrum and cerebellum, but a significant associated phenotype exists only in the cerebellum of Sg mice (Matsui et al., 1995). This result suggests that RORa is very important for the development and maturation of cerebellum but that other redundant factors may be present in the other parts of the brain. Besides RORa, RORE can bind RORb, RORg, RVR, and Rev-erbAa. As we mentioned earlier, RORb and RORg with RORa belong to the same subfamily of nuclear receptors. The RORb regulates genes whose products play a role in the context of sensory input integration as well as in the context of the circadian timing system (Andre et al., 1998). These functions of RORb make it an unattractive candidate to regulate the expression of prosaposin. The RORg is expressed in several tissues but is most highly expressed in skeletal muscle (Hirose et al., 1994). Thus, RORg is unlikely to be involved in driving tissue-preferential expression in CNS. The RVR lacks the activation function normally present at the C-terminal end of the ligand-binding domain and, through direct interaction with transcriptional corepressor N-CoR, represses basal as well as RORa1-mediated promoter activity. The RVR protein is ubiquitously expressed (Dussault and Giguere, 1997). If the phenomena that we observed are attributable to altered expression of RVR instead of absence of RORa in Sg mice, a dramatic decrease in promoter activity should not be restricted to the cerebellum of these mice. The Rev-erbAa is closely related to RVR and represses the basal level of transcription (Harding and Lazar, 1995). Expression of Rev-erbAa was found in muscle, adipocytes, and most regions of the cerebrum and cerebellum. The deletion of this gene alters development of Purkinje cells and delays development of cerebellum (Chomez et al., 2000). It is possible that Rev-erbAa may form a fine turning with RORa to regulate the expression of prosaposin in 234Luc. 788 JIN ET AL. However, no repressor effect was observed in the context of 310Luc in Sg mice. Apparently, the absence of RORa expression in Sg mice is the direct cause of increased promoter activity in the CNS of 310Luc/Sg2/2 animals, and RORa plays a fundamental role in the modulation of prosaposin expression. For Sp4 gene targeted mice, the increased promoter activity in the brain was preserved only with 310Luc. This result suggests that Sp4 modulated prosaposin gene expression in vivo through the 59 Sp1/U cluster in the absence of an upstream region. Although this study cannot exclude involvement of other Sp proteins, the correlation of lack of Sp4 expression and unchanged promoter activity in the eye of 310Luc/Sp42/2 mice indicates that Sp4 participates in the regulation of prosaposin gene in CNS, but in a minor way. In extensive studies with a series of transgenic mice containing deletion constructs, RORa was shown to play an important role in vivo in the regulation of prosaposin, and Sp4 may participate as well. Within shorter constructs (234Luc), RORa interacts with a coactivator to initiate the transcription of prosaposin without interference from more 59 upstream elements in a tissue-specific manner. With longer constructs (310Luc or 2400Luc), there exists potential interaction(s) between RORa and transcription factors that bind to the Sp1/U cluster, possibly Sp4. This interaction may be physically blocking the accessibility of coactivator(s) to RORa or regulator(s) that bind to the Sp1/U cluster. Once this interaction is disrupted with depletion of either RORa or Sp4, coactivators can interact with transcription factors to activate transcription. These results demonstrate the involvement of RORa and Sp4 in the regional expression of prosaposin in the CNS in vivo. ACKNOWLEDGMENTS The authors thank Ms. Maryann Koenig for her expert clerical assistance. This work was supported by grants to G.A.G. (NS 34071 and NS 36681). REFERENCES ANDRE, E., CONQUET, F., STEINMAYR, M., STRATTON, S.C., PORCIATTI, V., and BECKER-ANDRE, M. (1998). Disruption of retinoid-related orphan receptor beta changes circadian behavior, causes retinal degeneration and leads to vacillans phenotype in mice. EMBO J. 17, 3867–3877. BRADOVA, V., SMID, F., ULRICH-BOTT, B., ROGGENDORF, W., PATON, B.C., and HARZER, K. (1993). Prosaposin deficiency: Further characterization of the sphingolipid activator protein-deficien t sibs: Multiple glycolipid elevations (including lactosylceramidosis) , partial enzyme deficiencies and ultrastructure of the skin in this generalized sphingolipid storage disease. Hum. Genet. 92, 143–512. CARLBERG, C., HOOFT VAN HUIJSDUIJNEN, R., STAPLE, J.K., DELAMARTER, J.F., and BECKER-ANDRE, M. (1994). RZRs, a new family of retinoid-related orphan receptors that function as both monomers and homodimers. Mol. Endocrinol. 8, 757–770. CARLBERG, C., and WIESENBERG, I. (1995). The orphan receptor family RZR/ROR, melatonin and 5-lipoxygenase: An unexpected relationship. J. Pineal Res. 18, 171–178. CHOMEZ, P., NEVEU, I., MANSEN, A., KIESLER, E., LARSSON, L., VENNSTROM, B., and ARENAS, E. (2000). Increased cell death and delayed development in the cerebellum of mice lacking the reverbA(alpha) orphan receptor. Development 127, 1489–1498. DELERIVE, P., MONTE, D., DUBOIS, G., TROTTEIN, F., FRUCHART-NAJIB, J., MARIANI, J., FRUCHART, J.C., and STAELS, B. (2001). The orphan nuclear receptor ROR alpha is a negative regulator of the inflammatory response. EMBO Rep. 2, 42–48. DUSSAULT, I., and GIGUERE, V. (1997). Differential regulation of the N-myc proto-oncogene by ROR alpha and RVR, two orphan members of the superfamily of nuclear hormone receptors. Mol. Cell. Biol. 17, 1860–1867. DUSSAULT, I., FAWCETT, D., MATTHYSSEN, A., BADER, J.A., and GIGUERE, V. (1998). Orphan nuclear receptor ROR alpha-deficient mice display the cerebellar defects of staggerer. Mech. Dev. 70, 147–153. FUJITA, N., SUZUKI, K., VANIER, M.T., POPKO, B., MAEDA, N., KLEIN, A., HENSELER, M., SANDHOFF, K., and NAKAYASU, H. (1996). Targeted disruption of the mouse sphingolipid activator protein gene: A complex phenotype, including severe leukodystrophy and wide-spread storage of multiple sphingolipids. Hum. Mol. Genet. 5, 711–725. GIGUERE, V., TINI, M., FLOCK, G., ONG, E., EVANS, R.M., and OTULAKOWSKI, G. (1994). Isoform-specific amino-terminal domains dictate DNA-binding properties of ROR alpha, a novel family of orphan hormone nuclear receptors. Genes Dev. 8, 538–553. GOLLNER, H., BOUWMAN, P., MANGOLD, M., KARIS, A., BRAUN, H., ROHNER, I., DEL REY, A., BESEDOVSKY, H.O., MEINHARDT, A., VAN DEN BROEK, M., CUTFORTH, T., GROSVELD, F., PHILIPSEN, S., and SUSKE, G. (2001). Complex phenotype of mice homozygous for a null mutation in the Sp4 transcription factor gene. Genes Cells 6, 689–697. HAGEN, G., MULLER, S., BEATO, M., and SUSKE, G. (1992). Cloning by recognition site screening of two novel GT box binding proteins: A family of Sp1 related genes. Nucleic Acids Res. 20, 5519–5525. HAMILTON, B.A., FRANKEL, W.N., KERREBROCK, A.W., HAWKINS, T.L., FITZHUGH, W., KUSUMI, K., RUSSELL, L.B., MUELLER, K.L., VAN BERKEL, V., BIRREN, B.W., KRUGLYAK, L., and LANDER, E.S. (1996). Disruption of the nuclear hormone receptor RORalpha in staggerer mice [published erratum appears in Nature 1996;381:346]. Nature 379, 736–739. HARDING, H.P., and LAZAR, M.A. (1995). The monomer-binding orphan receptor Rev-Erb represses transcription as a dimer on a novel direct repeat. Mol. Cell. Biol. 15, 4791–4802. HARZER, K., PATON, B.C., POULOS, A., KUSTERMANN-KUHN, B., ROGGENDORF, W., GRISAR, T., and POPP, M. (1989). Sphingolipid activator protein deficiency in a 16-week-old atypical Gaucher disease patient and his fetal sibling: Biochemical signs of combined sphingolipidoses. Eur. J. Pediatr. 149, 31–39. HIRAIWA, M., SOEDA, S., KISHIMOTO, Y., and O’BRIEN, J.S. (1992). Binding and transport of gangliosides by prosaposin. Proc. Natl. Acad. Sci. USA 89, 11254–11258. HIROSE, T., SMITH, R.J., and JETTEN, A.M. (1994). ROR gamma: The third member of ROR/RZR orphan receptor subfamily that is highly expressed in skeletal muscle. Biochem. Biophys. Res. Commun. 205, 1976–1983. HULKOVA, H., CERVENKOVA, M., LEDVINOVA, J., TOCHACKOVA, M., HREBICEK, M., POUPETOVA, H., BEFEKADU, A., BERNA, L., PATON, B.C., HARZER, K., BOOR, A., SMID, F., and ELLEDER, M. (2001). A novel mutation in the coding region of the prosaposin gene leads to a complete deficiency of prosaposin and saposins, and is associated with a complex sphingolipidosis dominated by lactosylceramide accumulation. Hum. Mol. Genet. 10, 927–940. IGASE, K., TANAKA, J., KUMON, Y., ZHANG, B., SADAMOTO, Y., MAEDA, N., SAKAKI, S., and SAKANAKA, M. (1999). An TRANSCRIPTIONAL REGULATION OF PROSAPOSIN 789 18-mer peptide fragment of prosaposin ameliorates place navigation disability, cortical infarction, and retrograde thalamic degeneration in rats with focal cerebral ischemia. J. Cereb. Blood Flow Metab. 19, 298–306. JETTEN, A.M., KUREBAYASHI, S., and UEDA, E. (2001). The ROR nuclear orphan receptor subfamily: Critical regulators of multiple biological processes. Prog. Nucleic Acid Res. Mol. Biol. 69, 205–247. JIN, P., SUN, Y., and GRABOWSKI, G.A. (1998). Role of Sp proteins and RORalpha in transcription regulation of murine prosaposin. J. Biol. Chem. 273, 13208–13216. KINGSLEY, C., and WINOTO, A. (1992). Cloning of GT box-binding proteins: A novel Sp1 multigene family regulating T-cell receptor gene expression. Mol. Cell. Biol. 12, 4251–4261. KOTANI, Y., MATSUDA, S., WEN, T.C., SAKANAKA, M., TANAKA, J., MAEDA, N., KONDOH, K., UENO, S., and SANO, A. (1996). A hydrophilic peptide comprising 18 amino acid residues of the prosaposin sequence has neurotrophic activity in vitro and in vivo. J. Neurochem. 66, 2197–2200. MATSUI, T. (1996). Differential activation of the murine laminin B1 gene promoter by RAR alpha, ROR alpha, and AP-1. Biochem. Biophys. Res. Commun. 220, 405–410. MATSUI, T. (1997). Transcriptional regulation of a Purkinje cell-specific gene through a functional interaction between ROR alpha and RAR. Genes Cells 2, 263–272. MATSUI, T., SASHIHARA, S., OH, Y., and WAXMAN, S.G. (1995). An orphan nuclear receptor, mROR alpha, and its spatial expression in adult mouse brain. Brain Res. Mol. Brain Res. 33, 217–226. MORALES, C.R., EL-ALFY, M., ZHAO, Q., and IGDOURA, S. (1995). Molecular role of sulfated glycoprotein-1 (SGP-1/prosaposin) in Sertoli cells. Histol. Histopathol. 10, 1023–1034. O’BRIEN, J.S., CARSON, G.S., SEO, H.C., HIRAIWA, M., and KISHIMOTO, Y. (1994). Identification of prosaposin as a neurotrophic factor. Proc. Natl. Acad. Sci. USA 91, 9593–9596. O’BRIEN, J.S., KRETZ, K.A., DEWJI, N., WENGER, D.A., ESCH, F., and FLUHARTY, A.L. (1988). Coding of two sphingolipid activator proteins (SAP-1 and SAP-2) by same genetic locus. Science 241, 1098–1101. RORMAN, E.G., and GRABOWSKI, G.A. (1989). Molecular cloning of a human co-beta-glucosidase cDNA: Evidence that four sphingolipid hydrolase activator proteins are encoded by single genes in humans and rats. Genomics 5, 486–492. SAFFER, J.D., JACKSON, S.P., and ANNARELLA, M.B. (1991). Developmental expression of Sp1 in the mouse. Mol. Cell. Biol. 11, 2189–2199. SANDHOFF, K., KOHLER, T., and HARZER, K. (2001). Sphingolipid activator proteins. In The Metabolic and Molecular Bases of Inherited Diseases, ed 8. C.R. Scriver, A.L. Beaudet, W.S. Sly, and D. Valle, eds. (McGraw-Hill, New York) pp. 3371–3388. SANO, A., MATSUDA, S., WEN, T.C., KOTANI, Y., KONDOH, K., UENO, S., KAKIMOTO, Y., YOSHIMURA, H., and SAKANAKA, M. (1994). Protection by prosaposin against ischemia-induced learning disability and neuronal loss. Biochem. Biophys. Res. Commun. 204, 994–1000. SASHIHARA, S., FELTS, P.A., WAXMAN, S.G. and MATSUI, T. (1996). Orphan nuclear receptor ROR alpha gene: Isoform-specific spatiotemporal expression during postnatal development of brain. Brain Res. Mol. Brain Res. 42, 109–117. SIDMAN R., LANE, P.W., and DICKIE, M.M. (1962). Staggerer, a new mutation in the mouse affecting the cerebellum. Science 137, 610–612. SPRECHER-LEVY, H., ORR-URTREGER, A., LONAI, P., and HOROWITZ, M. (1993). Murine prosaposin: Expression in the reproductive system of a gene implicated in human genetic diseases [published erratum appears in Cell. Mol. Biol. 1994;40:233]. Cell. Mol. Biol. 39, 287–299. STEINHILBER, D., BRUNGS, M., WERZ, O., WIESENBERG, I., DANIELSSON, C., KAHLEN, J.P., NAYERI, S., SCHRADER, M., and CARLBERG, C. (1995). The nuclear receptor for melatonin represses 5-lipoxygenase gene expression in human B lymphocytes. J. Biol. Chem. 270, 7037–7040. STEINMAYR, M., ANDRE, E., CONQUET, F., RONDI-REIG, L., DELHAYE-BOUCHAUD, N., AUCLAIR, N., DANIEL, H., CREPEL, F., MARIANI, J., SOTELO, C., and BECKER-ANDRE, M. (1998). Staggerer phenotype in retinoid-related orphan receptor alpha-deficient mice. Proc. Natl. Acad. Sci. USA 95, 3960–3965. SUN, Y., JIN, P., and GRABOWSKI, G.A. (1997). The mouse prosaposin locus: Promoter organization. DNA Cell Biol. 16, 23–34. SUN, Y., JIN, P., WITTE, D.P., and GRABOWSKI, G.A. (1998). Isolation and characterization of the human prosaposin promoter. Gene 218, 37–47. SUN, Y., JIN, P., WITTE, D.P., and GRABOWSKI, G.A. (2000). Prosaposin: promoter analysis and central-nervous-system-preferential elements for expression in vivo. Biochem. J. 352, 549–56. SUN, Y., WITTE, D.P., and GRABOWSKI, G.A. (1994). Developmental and tissue-specific expression of prosaposin mRNA in murine tissues. Am. J. Pathol. 145, 1390–1398. SUPP, D.M., WITTE, D.P., BRANFORD, W.W., SMITH, E.P., and POTTER, S.S. (1996). Sp4, a member of the Sp1-family of zinc finger transcription factors, is required for normal murine growth, viability, and male fertility. Dev. Biol. 176, 284–299. VU-DAC, N., GERVOIS, P., GROTZINGER, T., DE VOS, P., SCHOONJANS, K., FRUCHART, J.C., AUWERX, J., MARIANI, J., TEDGUI, A., and STAELS, B. (1997). Transcriptional regulation of apolipoprotein A-I gene expression by the nuclear receptor RORalpha. J. Biol. Chem. 272, 22401–22404. WIESENBERG, I., MISSBACH, M., KAHLEN, J.P., SCHRADER, M., and CARLBERG, C. (1995). Transcriptional activation of the nuclear receptor RZR alpha by the pineal gland hormone melatonin and identification of CGP 52608 as a synthetic ligand. Nucleic Acids Res. 23, 327–333. Address reprint requests to: Dr. Gregory A. Grabowski Human Genetics Division Children’s Hospital Medical Center 3333 Burnet Ave Cincinnati, OH 45229-303 9 E-mail: [email protected] Received for publication November 8, 2001; accepted November 21, 2001.