* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download PPT
History of genetic engineering wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genetic engineering wikipedia , lookup
History of RNA biology wikipedia , lookup
Genomic library wikipedia , lookup
Primary transcript wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Transposable element wikipedia , lookup
Maximum parsimony (phylogenetics) wikipedia , lookup
Genome (book) wikipedia , lookup
Point mutation wikipedia , lookup
Population genetics wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Microevolution wikipedia , lookup
Viral phylodynamics wikipedia , lookup
Pathogenomics wikipedia , lookup
Public health genomics wikipedia , lookup
Helitron (biology) wikipedia , lookup
Metagenomics wikipedia , lookup
Non-coding DNA wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Human genome wikipedia , lookup
Genome evolution wikipedia , lookup
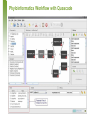
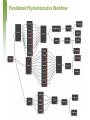
Phyloinformatics of Neuraminidase at Micro and Macro Levels using Grid-enabled HPC Technologies B. Schmidt (UNSW) D.T. Singh (Genvea Biosciences) R. Trehan, T. Bretschneider (NTU, Singapore) March 26, 2007 Contents • • • • • • H5N1 Genetics H5N1 Phyloinformatics Design Principles of Quascade H5N1 Phyloinformatics with Quascade Results Conclusion and Future work March 26, 2007 H5N1 Genetics • • • • Belongs to the Influenza A virus type Segmented RNA genome 8 genes, 11 proteins Classification based on: – Hemagglutinin (HA): 15 subtypes – Neuraminidase (NA): 9 subtypes • Genetic variations in HA/NA • Genetic drift – Point mutations – 1918 Spanish flu • Genetic shift – Reassortment of the segmented genome – 1957, 1968, 1997 pandemics – 2003 Z strain of H5N1 March 26, 2007 H5N1 Phyloinformatics • Essential to monitor new emerging strains – Molecular evolution at gene and genome level – Phylogenetic analysis for determining the origin of new strains • Phylogenetics – How fast do proteins evolve? – What is the best method to measure the evolution? – How to obtain the best phylogenetic tree? • Phylogenetic algorithms – Character based • Maximum Parsimony, Maximum Likelihood (ML) – Distance based • UPGMA, Neighborhood Join (NJ) – Bayesian MCMC based • Mr. Bayes, BEAST March 26, 2007 Quascade – User Interface Example Processing pipeline Communication • A data-flow tool in which each black-box represents Java objects running on different computers! • Assignment of objects to available computers done automatically (manually if required) • Communication between objects done transparently • Configuration of objects done before run-time March 26, 2007 Object Features Java Object Java Object Java Object • • • • • Coding in regular Java/ C/ C++ Persistent – activated whenever all data-inputs present No explicit messaging protocol required No distributed computing concepts need to be understood Objects automatically or manually assigned to computers / CPU-cores March 26, 2007 Phyloinformatics Workflow with Quascade March 26, 2007 Parallelized Phyloinformatics Workflow March 26, 2007 Data and Algorithms • Core Group – 22 H5N1 NA sequences from SwissProt and TREMBL • Medium Set – 581 NA H5N1 sequences from Uniprot • Large Set – 909 NA Influenza A sequences from Uniprot • ProtDist – NJ – UPGMA • ProtPars • ProtML • Mr. Bayes March 26, 2007 Runtime and Scalability (NA Bird Flu Protein) 1 processor 25 processors 400 360 300 200 145 100 16 0 909 sequences 6 581 sequences Processing time [h] Processing time [h] 400 Distance-based workflow MP workflow 360 300 200 140 100 16 5 909 sequences 581 sequences 0 March 26, 2007 P18269Sial Q05JH9H9N2 Q6DTU0swinech03 Mr Bayes – Tree Core Set 0.75 A1EHP1goBav06 0.99 A1EHP3goBa06 Q0A2H3Chsc59 1.00 Q710U6chSc59 1.00 Q0PEF9chIn06 0.99 Q0PEG0chIn06 Q5MD56TiTh04 Q6PUP6HuTh04 0.63 Q307V5catth04 Q5SDA6chTh04 0.71 Q45ZM8wpfth04 Q307U7PigeonTh04 Q6PUP7HuTh04 0.90 Q2L700HuTh05 0.70 0.91 Q2LDC0QuTh06 0.54 Q2LDC8chTh05 Q6B518chTh04 0.86 March 26, 2007 Q4PKD4chTh04 Analysis and Observations • • • Clustering possibilities – Temporal, host-based, geographical Algorithms – Mr. Bayes and ProtML are most consistent in their performance – Too compute-intensive for the larger “macro” sets Observed pattern – All phylograms yielded geographic-based clustering rather than timebased clustering – Host ranges along clustered clades vary – Same strain with identical NA sequences can infect different hosts – NA may not be the sole factor responsible for determining the diverse host range – Glycan site acquisition or loss seems to play a critical role in the molecular evolution of H5N1 NA – Identification of “bridging isolates” may help in rapid monitoring and development of global scale warning system for H5N1 March 26, 2007 Conclusion and Future Work • Quascade – – – – New graphical data-flow tool to design automatically grid-enabled pipelines / workflows Supports implicit high-performance parallelization Supports persistent components Can be used with Java / C/ C++ code or application-binaries • H5N1 Phyloinformatics – – – – Can take advantage of workflow system and HPC Can be easily used and modified by biologists Use H5N1 NA sequences to better understand evolution of H5N1 Analysis of H5N1 NA data with different algorithms indicates spatial clustering based on geographical distribution rather than temporal or host. • Future work – Studies in conjunction with other proteins such as HA, Polymerase etc., and also at gene and genome level March 26, 2007