* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download p53
Epitranscriptome wikipedia , lookup
Non-coding RNA wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Histone acetylation and deacetylation wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Community fingerprinting wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
Gene expression profiling wikipedia , lookup
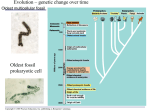
Genome evolution wikipedia , lookup
List of types of proteins wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Non-coding DNA wikipedia , lookup
Gene regulatory network wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Point mutation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Molecular evolution wikipedia , lookup
Gene expression wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Chapter 19 Eukaryotic Genomes: Organization, Regulation, and Evolution Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Overview: How Eukaryotic Genomes Work and Evolve • In eukaryotes, the DNA-protein complex, called chromatin – Is ordered into higher structural levels than the DNA-protein complex in prokaryotes. How can this structure be ordered in this intricate manner? Figure 19.1 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Both prokaryotes and eukaryotes – Must alter their patterns of gene expression in response to changes in environmental conditions Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Concept 19.1: Chromatin structure is based on successive levels of DNA packing • Eukaryotic DNA – Is precisely combined with a large amount of protein with the resulting chromatin undergoes striking changes during the cell cycle – When the cell prepare to mitosis, its chromatin coils and folds to form the chromosomes • Eukaryotic chromosomes – Contain an enormous amount of DNA contain a single linear DNA double helix that averages 200 million base pairs in humans Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Nucleosomes, or “Beads on a String” • Proteins called histones – Are responsible for the first level of DNA packing in chromatin – Bind tightly to DNA because they have a high proportion of positively charged a.a that binds to the negatively charged DNA. • The association of DNA and histones – Seems to remain intact throughout the cell cycle Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings In electron micrographs – Unfolded chromatin has the appearance of beads on a string • Each “bead” is a nucleosome; The basic unit of DNA packing – Histones leave DNA only transiently during DNA replication but stay with DNA during transcription. 2 nm DNA double helix Histones Histone tails Histone H1 Linker DNA (“string”) Nucleosome (“bad”) (a) Nucleosomes (10-nm fiber) Figure 19.2 a Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 10 nm Higher Levels of DNA Packing • With the aid of the histone H1 (one of 5 histones involved in the coiling process), the beaded string coils to from a fiber roughly 30 nm in thickness known as the 30-nm chromatin fiber. 30 nm Figure 19.2 b Nucleosome (b) 30-nm fiber Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The 30-nm fiber, in turn – Forms looped domains, making up a 300-nm fiber which are attached to a chromosome scaffold (platform) made of non-histone proteins. Protein scaffold Loops 300 nm (c) Looped domains (300-nm fiber) Figure 19.2 c Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Scaffold • In a mitotic chromosome – The looped domains themselves coil and fold further compacting the chromatin forming the characteristic metaphase chromosome 700 nm 1,400 nm (d) Metaphase chromosome Figure 19.2 d Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • In interphase – Some portions of certain chromosomes exist in a very condensed state that can be seen under light microscope. This is called heterochromatin and is distinguished from the less compact chromatin that is called euchromatin. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Concept 19.2: Gene expression can be regulated at any stage, but the key step is transcription • All organisms – Must regulate which genes are expressed at any given time. i.e not every gene will be active at all times. • During development of a multicellular organism – Its cells undergo a process of specialization in form and function called cell differentiation. This results in several cell types reaching up to 200 different cells types in adult humans. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Differential Gene Expression • Each cell of a multicellular eukaryote – Expresses only a fraction of its genes (probably 20% ) at any given time. • In each type of differentiated cell – A unique subset of genes is expressed with the highly differentiated cells such as muscle cells expressing a lower number of genes at any given time. – Differences between cell types are NOT due to different genes but to different gene expression by cells with the same genome. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The genome of eukaryotes may contain tens of thousands of genes but only a small amount of DNA (1.5% in humans ) codes for proteins. • What determine which genes to be expressed? – Environmental signals, certain genes turn on while others turn off. – Cell differentiation during organism development. • The enzymes that transcribe DNA must locate the target genes at the right time. This task is like finding a needle in haystack. • When gene expression goes wrong, certain disease such as cancer can arise. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Many key stages of gene expression Signal Stages in Gene Expression that can be regulated in eukaryotic cells; The colored boxes indicates these steps. NUCLEUS Chromatin Chromatin modification: DNA unpacking involving histone acetylation and DNA demethlation DNA Gene available for transcription Gene Transcription RNA Exon Primary transcript Intron RNA processing Tail Cap mRNA in nucleus Transport to cytoplasm CYTOPLASM mRNA in cytoplasm Degradation of mRNA Translation Polypetide Cleavage Chemical modification Transport to cellular destination Active protein Degradation of protein Figure 19.3 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Degraded protein Regulation of Chromatin Structure • Genes within highly packed heterochromatin – Are usually not expressed Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Histone Modification • Chemical modification of histone tails – Can affect the configuration of chromatin and thus gene expression Chromatin changes Transcription RNA processing mRNA degradation Translation Protein processing and degradation Histone tails DNA double helix Figure 19.4a Amino acids available for chemical modification (a) Histone tails protrude outward from a nucleosome Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Histone acetylation • is the attachment of acetyl groups (--COCH3) to certain amino acids of histone proteins. – Seems to loosen chromatin structure and thereby lose their grip on DNA thus enhance transcription Unacetylated histones Figure 19.4 b Acetylated histones (b) Acetylation of histone tails promotes loose chromatin structure that permits transcription Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings DNA Methylation • Addition of methyl groups (…CH3) to certain bases in DNA – Genes are usually more highly methylated when they are not expressed. – De-methylation to certain inactive genes turns them on. – In some species DNA methylation guarantees long term inactivation of certain genes – Once methylated, genes stay as such through successive cell division. This property accounts for genomic imprinting in mammals where methylation permanently turns off either maternal or paternal allele of a certain gene. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Epigenetic Inheritance – Is the inheritance of traits transmitted by mechanisms not directly involving the nucleotide sequence. – Chromatin modifying enzymes are integral part of this process. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Regulation of Transcription Initiation • Chromatin-modifying enzymes provide initial control of gene expression – By making a region of DNA either more or less able to bind the transcription machinery. – Once a gene is optimally modified for expression, the initiation of transcription process is the most important. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Organization of a Typical Eukaryotic Gene • Associated with most eukaryotic genes are multiple control elements which are; – Segments of noncoding DNA that help regulate transcription by binding certain proteins Proximal Enhancer control elements (distal control elements) Poly-A signal Termination sequence region Exon DNA Upstream Intron Transcription Primary RNA transcript 5 (pre-mRNA) Exon Intron Intron RNA RNA processing Intron Exon Cleared 3 end of primary transport Coding segment Translation Protein processing and degradation Exon Poly-A signal RNA processing: Cap and tail added; introns excised and exons spliced together Transcription Figure 19.5 Intron Exon Downstream Promoter Chromatin changes mRNA degradation Exon mRNA G P P P 5 Cap 5 UTR (untranslated region) Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Start Stop 3 UTR Poly-A codon codon (untranslated tail region) The Roles of Transcription Factors • To initiate transcription – Eukaryotic RNA polymerase requires the assistance of proteins called transcription factors. These factors are required for all types of protein coding genes thus called general transcription factors. – One of these transcription factors recognize a DNA sequence called the TATA box within the promoter, while the others recognize other proteins including the RNA polymerase. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Enhancers and Specific Transcription Factors – The interaction of these transcription factors and RNA polymerase II with a promoter usually initiates transcription but inefficiently producing few RNA transcripts. – Control elements, proximal control elements (near the promoter) greatly improve the efficiency of promoters by binding additional transcription factors. – The more distant “ distal control elements” are called the enhancers which might be thousands of nucleotides away from the promoter or may be located in an intron. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • An activator – Is a protein that binds to an enhancer and stimulates transcription of a gene Distal control element Activators Enhancer 1 Activator proteins bind to distal control elements grouped as an enhancer in the DNA. This enhancer has three binding sites. 2A DNA-bending protein brings the bound activators closer to the promoter. Other transcription factors, mediator proteins, and RNA polymerase are nearby. Promoter TATA box Gene General transcription factors DNA-bending protein Group of Mediator proteins RNA Polymerase II Chromatin changes 3The activators bind to certain general transcription factors and mediator proteins, helping them form an active transcription initiation complex on the promoter. Transcription RNA processing mRNA degradation Figure 19.6 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Translation Protein processing and degradation RNA Polymerase II Transcription RNA synthesis Initiation complex • Some specific transcription factors function as repressors – To inhibit expression of a particular gene • Some activators and repressors – Act indirectly by influencing chromatin structure – A gene present in the region of chromatin with high levels of histone acetylation is able to bind to the transcription machinery while the one with low levels can not bind. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings In yeast and some mammals some activator recruit proteins that acetylate histones near the promorter of a gene thus enhance transcription. Repressors recruit proteins that deacetylate histones leading to reduced transcription. This phenomenon is called gene silencing. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Coordinately Controlled Genes • Unlike the genes of a prokaryotic operon which are regulated by one promoter – Coordinately controlled eukaryotic genes each have a promoter and control elements even if they are located close to each other on the same chromosome. • The same regulatory sequences – Are common to all the genes of a group, enabling recognition by the same specific transcription factors – An example of such coordinate control is the activation of a variety of genes by steroid home (sex hormones). – Genes with the same control elements are activated by the same chemical signals. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Mechanisms of Post-Transcriptional Regulation • The expression of a protein-coding gene is ultimately measured in terms of amount of functional protein the cell makes. • In theory a gene expression may be blocked or stimulated at any post-transcriptional step. • A cell can therefore, use its regulatory mechanisms that operate after the transcription to fine tune gene expression in response to environmental changes without altering its transcription patterns. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings RNA Processing • In alternative RNA splicing – Different mRNA molecules are produced from the same primary transcript, depending on which RNA segments are treated as exons and which as introns. Chromatin changes Transcription RNA processing mRNA degradation Translation Protein processing and degradation Exons DNA Primary RNA transcript RNA splicing Figure 19.8 mRNA Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings or mRNA Degradation • The life span of mRNA molecules in the cytoplasm – Is an important factor in determining the protein synthesis in a cell – Prokaryotic mRNA have a very short life span i.e they are degraded enzymatically after few minutes. – In contrast eukaryotic mRNA life span is typically hours, but could be days or even weeks such mRNA for hemoglobin polypeptides – It is believed that the removal of the 5’ cap provides a site for the nuclease enzymes to chew up the mRNA. – Nucleotide sequences that affect mRNA stability are normally found in the trailer region (un-translated region, UTR) at the 3’ end of the molecule. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Recent discoveries • RNA interference by single-stranded microRNAs (miRNAs) – Can lead to degradation of an mRNA or block its translation – This happen due to a small interfering RNAs (siRNAs) which the same size as miRNAs and does the same function 1 The microRNA (miRNA) precursor folds back on itself, held together by hydrogen bonds. 2 An enzyme called Dicer moves along the doublestranded RNA, cutting it into shorter segments. 5 4 3 One strand of each short doublestranded RNA is degraded; the other strand (miRNA) then associates with a complex of proteins. The bound miRNA can base pair with any target mRNA that contains the complementary sequence. 5 The miRNA-protein complex prevents gene expression either by degrading the target mRNA or by blocking its translation. Chromatin changes Transcription RNA processing Protein complex mRNA degradation Translation Protein processing and degradation Dicer Degradation of mRNA OR miRNA Target mRNA Figure 19.9 Hydrogen bond Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Blockage of translation Initiation of Translation • The initiation of translation of selected mRNAs – Can be blocked by regulatory proteins that bind to specific sequences or structures of the mRNA. This adds another opportunity for regulating gene expression • Alternatively, translation of all the mRNAs in a cell – May be regulated simultaneously. This global control usually involves activation or inactivation of one or more of the protein factors required to initiate translation. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Protein Processing and Degradation • The final opportunities for controlling gene expression occurs after translation – Various types of protein processing, including cleavage and the addition of chemical groups, or the length of time each protein functions in the cell are subject to control. Examples; – Cleaving pro-insulin to form the active form, insulin. – Addition of phosphate groups or sugars – Cyclins which are proteins involved in regulating the cell cylce must be short-lived if the cell to function properly. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Proteasomes – The protein molecules to be degraded must be bind to another protein called ubiquitin before being degraded by a giant protein complexes called proteasome. 1 3 2 Multiple ubiquitin molecules are attached to a protein by enzymes in the cytosol. Chromatin changes The ubiquitin-tagged protein is recognized by a proteasome, which unfolds the protein and sequesters it within a central cavity Enzymatic components of the proteasome cut the protein into small peptides, which can be further degraded by other enzymes in the cytosol. Transcription RNA processing mRNA degradation Translation Proteasome and ubiquitin to be recycled Ubiquitin Proteasome Protein processing and degradation Protein to be degraded Ubiquinated protein Protein entering a proteasome Figure 19.10 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Protein fragments (peptides) • Concept 19.3: Cancer results from genetic changes that affect cell cycle control • The gene regulation systems that go wrong during cancer turn out to be the very same systems that play important roles in embryonic development Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Types of Genes Associated with Cancer • The genes that normally regulate cell growth and division during the cell cycle – Include genes for growth factors, their receptors, and the intracellular molecules of signaling pathways. • Its is believed that many causing-cancer mutations results from environmental influences such as chemical carcinogens, UV light, X-Ray or certain viruses. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Examples of viruses that cause cancer are; • EBV that cause infectious mononucleosis and is associated with Burkitt’s lymphoma • Papiloma virus associated with cervix cancer • HTLV-1 virus associated with some adults leukemia All tumor viruses transform cells into cancer cells by integrating their nucleic acids with host DNA. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Oncogenes and Proto-Oncogenes • Oncogenes – Are cancer-causing genes • Proto-oncogenes – Are normal cellular genes that code for proteins that stimulate normal cell growth and division Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings How a proto-oncogene become an oncogene? • An oncogene arises from a genetic change that leads to an increase in either; • the amount of the proto-oncogene’s protein product • or the intrinsic activity of each protein molecule. • Now these genetic changes fall into three catergories; (Figure 19-11). Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Gentic changes that change proto-onco to oncogene • movement of DNA within the genome, where it might land close to an especially active promoter that increase transcription of the gene making it an oncogene. • amplification of proto-oncogene which increase the number of copies of oncogene inside the cell • point mutation in a proto-oncogene that change the gene’s protein product to a more active or more resistant to degradation than the normal protein. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • A DNA change that makes a proto-oncogene excessively active – Converts it to an oncogene, which may promote excessive cell division and cancer Proto-oncogene DNA Translocation or transposition: gene moved to new locus, under new controls Gene amplification: multiple copies of the gene New promoter Normal growth-stimulating protein in excess Point mutation within a control element Oncogene Normal growth-stimulating protein in excess Figure 19.11 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Point mutation within the gene Oncogene Normal growth-stimulating Hyperactive or degradationprotein in excess resistant protein Tumor-Suppressor Genes • Tumor-suppressor genes – Encode proteins that inhibit abnormal cell division. • Protein products of tumor suppressor genes could be; – proteins normally repair damaged DNA – some proteins control adhesion of cells to each other or to an extracellular matrix – other proteins are component of signaling pathways that inhibit the cell cycle. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Interference with Normal Cell-Signaling Pathways • Many proto-oncogenes and tumor suppressor genes – Encode components of growth-stimulating and growth-inhibiting pathways, respectively. • What protein products of cancer genes do or fail to do? – Let us look at products of two cancer genes; the ras proto-ocogenes and p53 tumor suppressor gene. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Mutations in these genes are very common in human cancers; ras is mutated in about 30% of human cancers while it is 50% for p53. • Both proteins, ras and p53 are parts of the signal transduction pathways that convey external signal to the DNA in the nucleus. Figure 19-12 shows two possibilities of an outcome when either gene goes wrong; – one an over expression of ras protein that ends in the increase in the production of protein that stimulates cell cycle and leads to cancer – An under expression of a protein that inhibits the cell cycle as a result of mutation in the p53 gene and leads to cancer. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The p53 gene is names as it encodes for a protein of 53 kDa molecular weight. It is very important to the point that its name is “the guardian angel of the genome”. How does the p53 works in protecting the cells form cancer? • A damage to the cell’s DNA leads to the expression of p53 • This in turn lead to the activation of a p21gene whose product leads to halting the cell cycle thus allowing time for the cell to repair its damaged DNA • The protein p53 can turn on genes directly involved in DNA repair • When DNA damage is irreparable, p53 protein activates a suicide genes whose protein products cause cell death in a process called apoptosis Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Cell cycle stimulating pathway • This pathway is triggered by a growth factor that binds to its receptor in the plasma membrane. • The signal is relayed to a G protein called Ras. Like all G proteins, Ras is active when GTP is bound to it. • Ras passes the signal to a series of protein kinases. • The last kinase activates a transcription activator that turns on one or more genes. • For proteins that stimulate the cell cycle. If a mutation makes Ras or any other pathway component abnormally active, excessive cell division and cancer may result. See next slide for explanation. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The Ras protein, encoded by the ras gene – Is a G protein that relays a signal from a growth factor receptor on the plasma membrane to a cascade of protein kinases 1 Growth factor Mutation Hyperactive Ras protein (product of oncogene) issues signals on its own 3 protein G 4 2 Receptor Protein kinases (phosphorylation cascade) Nucleus 5 Transcription factor (activator) Gene expression Protein that Stimulates the cell cycle Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Cell cycle inhibiting pathway • In this pathway, DNA damage is an intracellular signal that is passed via protein kinases and leads to activation of p53. • Activated p53 promotes transcription of the gene for a • protein that inhibits the cell cycle. • The resulting suppression of cell division ensures that the damaged DNA is not replicated. • Mutations causing deficiencies in any pathway component can contribute to the development of cancer. • See next slide for explanation. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The p53 gene encodes a tumor-suppressor protein – That is a specific transcription factor that promotes the synthesis of cell cycle–inhibiting proteins 2 Protein kinases MUTATION UV light 3 1 DNA damage in genome Defective or missing Transcription factor, such as p53, cannot activate transcription Active form of p53 DNA Protein that inhibits the cell cycle Figure 19.12b Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Mutations that knock out the p53 gene – Can lead to excessive cell growth and cancer Effects of mutations. (c) Increased cell division, possibly leading to cancer, can result if the cell cycle is overstimulated, as in (a), or not inhibited when it normally would be, as in (b). EFFECTS OF MUTATIONS Protein overexpressed Cell cycle overstimulated Figure 19.12c Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Protein absent Increased cell division Cell cycle not inhibited The Multistep Model of Cancer Development • Normal cells are converted to cancer cells – By the accumulation of multiple mutations affecting proto-oncogenes and tumorsuppressor genes • More than one somatic mutation is generally needed to produce the entire changes characteristic of cancer cell. That is why cancers increase with increase age. • Does this means that mutations accumulate until they cause cancer? Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings A study of colorectal cancer • It affect 135,000 patients every year in US with 60,000 deaths each year. • Like other cancers it develops gradually. • The first sign is often a polyp which is small, benign growth in the colon lining. Cells look normal although they divide unusually frequently • Tumor grows and may eventually become malignant • Malignant tumor develops parallel by a graduate accumulation of mutations that activate oncogenes and knock out tumor suppressor genes. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Colorectal cancer…. Cont. • A bout a half dozen changes must occur at the DNA level for a cell to become fully cancerous including; • Appearance of at least one active oncogenes • Mutation or loss of several tumor suppressor genes • Since mutant tumor suppressor alleles are recessive, mutation must knock out both alleles in a cell’s genome to block tumor suppression • The genes for tolemerase gets activated. This enzyme prevent the erosion of the ends of the chromosomes thus removing the natural limits on the number of times the cell can divide. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • A multistep model for the development of colorectal cancer Colon Loss of tumorsuppressor Colon wall gene APC (or other) 1 Activation of ras oncogene 2 Loss of tumorsuppressor gene DCC Loss of tumor-suppressor gene p53 4 3 Normal colon epithelial cells Small benign growth (polyp) Figure 19.13 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Additional mutations 5 Larger benign growth (adenoma) Malignant tumor (carcinoma) Viruses and cancer • Viruses seem to play a part of about 15% of human cancer cases worldwide. • Viruses contribute to cancer development by; – Integrating their DNA into the host DNA thus they might donate an oncogenes to the host – They might insert a cellular gene that disrupts a tumor suppressor gene or converts a protooncogene to an oncogene. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Inherited Predisposition to Cancer • That fact that multiple genetic changes are required to produce a cancer help explain the predisposition to cancer that run in some families. How does that happen? • Individuals who inherit a mutant oncogene or tumor-suppressor allele Have an increased risk of developing certain types of cancer • About 15% of colorectal cancers involve inhereted mutations that affect a tumor-suppressor gene called adenomatous polyposis coli (APC). This gene involves in cell migration and adhesion. • Breast cancer is another example where mutations of BRCA1 and BRCA2 (tumor suppressor gene) are found in at least 50% if inherited breast cancers. • A woman with inherited one mutant of BRCA1 allele has 60% chance of developing breast cancer before age 50 compared with only 2% probability for an individual homozygous for the normal allele. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Concept 19.4: Eukaryotic genomes can have many noncoding DNA sequences in addition to genes • The bulk of most eukaryotic genomes – Consists of noncoding DNA sequences, often described in the past as “junk DNA” • However, much evidence is accumulating – That noncoding DNA plays important roles in the cell. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings The Relationship Between Genomic Composition and Organismal Complexity • Compared with prokaryotic genomes, the genomes of eukaryotes – Generally are larger – Have longer genes; – Contain a much greater amount of noncoding DNA both associated with genes and between genes. This DNA includes the introns (noncoding DNA stritches that interrupt the coding sequences) and repetitive DNA. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • In prokaryotes most of the DNA in the genome account for proteins, tRNA or rRNA, with the coding sequences proceed from the start to finish without interruption. • Now that the complete sequence of the human genome is available we know what makes up most of the 98.5% that does NOT code for proteins, rRNAs, or tRNAs. Figure 19.14 shows this distribution. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Types of DNA sequences in human genome Exons (regions of genes coding for protein, rRNA, tRNA) (1.5%) Repetitive DNA that includes transposable elements and related sequences (44%) Alu elements (10%) Figure 19.14 Simple sequence DNA (3%) Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Introns and regulatory sequences (24%) Repetitive DNA unrelated to transposable elements (about 15%) Unique noncoding DNA (15%) Large-segment duplications (5-6%) Transposable Elements and Related Sequences • All organisms seem to have stretches of DNA that can move from one location to another. • The first evidence for these wandering DNA segments – Came from geneticist Barbara McClintock’s breeding experiments with Indian corn Figure 19.15 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Movement of Transposons and Retrotransposons • Eukaryotic transposable elements are of two types – Transposons, which move within a genome by means of a DNA intermediate and a copy and paste or cut and paste mechanisms. Figure 19.16 a&b – Retrotransposons, which move by means of an RNA intermediate. Most of transposable elements in eukaryotes are reterotransposons. – These transposons encode for reverse transcriptase (RT) enzyme. Thus RT can be present in cells not infected with virus. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Movement of eukaryotic transposable elements Transposon New copy of transposon DNA of genome Transposon is copied Insertion Mobile transposon (a) Transposon movement (“copy-and-paste” mechanism) Retrotransposon New copy of retrotransposon DNA of genome RNA Reverse transcriptase (b) Retrotransposon movement Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Insertion Figure 19.16a, b Sequences Related to Transposable Elements • Multiple copies of transposable elements and sequences related to them – Are scattered throughout the eukaryotic genome and make about 25-50% of the human genome. • In humans and other primates – A large portion of transposable element–related DNA consists of a family of similar sequences called Alu elements (10% of the human genome). – Alu elements are about 300 nucleotides long and they do not code for any protein Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Other Repetitive DNA, Including Simple Sequence DNA • Repetitive DNA that is not related to transposable elements might have originated by mistake during DNA replication and it accounts for 15% of human genome of which 5% consist of very large segment duplications. • Simple sequence DNA – Contains many copies of tandemly repeated short sequences – Is common in centromeres and telomeres, where it probably plays structural roles in the chromosome Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The simple DNA sequences located at the telomeres at the tip of the chromosome prevents genes from being lost during replication. • In mammals a considerable portion of the genome is tandemly repetitive DNA, which are short repetitive sequences in series such as; • ….. GTTAC GTTAC GTTAC GTTAC GTTAC GTTAC…. • The number of these repeated short series could reach several hundred thousands and the repeated units could be up to 10 base pairs. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The nucleotide composition of this DNA differs from the rest of DNA to the point that it can be isolated by differential ultracentrifugation. • Repetitive DNA sequences originally isolated in this way was called satellite DNA because it appears as a satellite band in the centrifuge tube. • The term satellite DNA is now used interchangeably with the term simple sequence DNA. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Genes and Multigene Families • Most eukaryotic genes – Are present in one copy per haploid set of chromosomes • The rest of the genome (particularly in humans) – Occurs in multigene families, collections of identical or very similar genes Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Some multigene families – Consist of identical DNA sequences, usually clustered tandemly, such as those that code for RNA products RNA transcripts DNA Non-transcribed spacer Transcription unit DNA 18S 5.8S 28S rRNA Figure 19.17a Part of the ribosomal RNA gene family Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 28S 18S 5.8S • The classic examples of multigene families of nonidentical genes – Are two related families of genes that encode globins -Globin Heme Hemoglobin -Globin -Globin gene family -Globin gene family Chromosome 16 Chromosome 11 Figure 19.17b The human -globin and -globin gene families Embryo 1 2 1 2 Fetus and adult Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings G A Embryo Fetus Adult • The previous figure shows that human hemoglobin has two subunits located on different chromosomes. • The different forms of each globin is expressed at different stage of development. • Example in humans embryonic and fetal forms of HB have higher affinity to O2 than adult forms to insure efficient transfer of O2 from mother to fetus. • Also found in globin gene family clusters are pseudogenes, which are non-functional sequences similar to the functional ones. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Concept 19.5: Duplications, rearrangements, and mutations of DNA contribute to genome evolution • The basis of change at the genomic level is mutation which underlies much of genome evolution. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Duplication of Chromosome Sets • Accidents in meiosis can result in one or more extra sets of chromosomes (polyploidy). • In such case one complete set of gene can provide normal functions while the other can diverge by accumulating mutations. • This divergence might later lead to new phenotypes if the organism survives. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Duplication and Divergence of DNA Segments • Unequal crossing over during prophase I of meiosis – Can result in one chromosome with a deletion and another with a duplication of a particular gene Transposable element Gene Nonsister chromatids Crossover Incorrect pairing of two homologues during meiosis and Figure 19.18 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Evolution of Genes with Related Functions: The Human Globin Genes • The genes encoding the various globin proteins – Evolved from one common ancestral globin gene, which was duplicated and diverged • Subsequent duplications of these genes and random mutations – Gave rise to the present globin genes, all of which code for oxygen-binding proteins. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Evolution of Genes with Related Functions: The Human Globin Genes Ancestral globin gene Duplication of ancestral gene Mutation in both copies Transposition to different chromosomes Further duplications and mutations Figure 19.19 2 1 2 -Globin gene family on chromosome 16 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 1 G A -Globin gene family on chromosome 11 • The similarity in the amino acid sequences of the various globin proteins – Supports this model of gene duplication and mutation Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Table 19.1 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Evolution of Genes with Novel Functions • The copies of some duplicated genes – Have diverged so much during evolutionary time that the functions of their encoded proteins are now substantially different – Gene of lysozyme and α-lactoalbumin are good examples as they have a similar a.a seqeunce and 3-D structure yet completely different function. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Rearrangements of Parts of Genes: Exon Duplication and Exon Shuffling • The presence of introns in eukaryotic genes may have promoted the development of new proteins by facilitating duplication and rearrangement of exons. • A particular exon within a gene – Could be duplicated on one chromosome and deleted from the homologous chromosome and this could happen due to unequal crossing over. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings In exon shuffling – Errors in meiotic recombination lead to the occasional mixing and matching of different exons either within a gene or between two nonallelic genes EGF EGF EGF EGF Epidermal growth factor gene with multiple EGF exons (green) Exon shuffling F F F Fibronectin gene with multiple “finger” exons (orange) Exon duplication F F EGF K K K Plasminogen gene with a “kfingle” exon (blue) Portions of ancestral genes Figure 19.20 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Exon shuffling TPA gene as it exists today How Transposable Elements Contribute to Genome Evolution • Movement of transposable elements or recombination between copies of the same element Can contribute to the evolution of the genome in several ways; • Promote recombination • Disrupt cellular genes or control elements, thus interfering with production of proteins. • Carry entire genes or exons to new locations e.g the presence of α and β-globin genes on two different chromosomes. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings End of Chapter Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings